
Prepares IRT analysis for Conquest, TAM, or mirt
defineModel.RdFacilitates data analysis using the software Conquest, TAM, or mirt. It automatically
checks data for IRT consistency, generates Conquest syntax, label, anchor and data files or
corresponding TAM/mirt call for a single model specified by several arguments in R. Finally, an
R object is created which contain the required input for Conquest or TAM. To start the estimation,
call runModel with the argument returned by defineModel.
Usage
defineModel (dat, items, id, splittedModels = NULL,
irtmodel = c("1PL", "2PL", "PCM", "PCM2", "RSM", "GPCM", "GPCM.groups", "2PL.groups", "GPCM.design", "3PL"),
qMatrix=NULL, DIF.var=NULL, HG.var=NULL, group.var=NULL, weight.var=NULL, anchor = NULL,
domainCol=NULL, itemCol=NULL, valueCol=NULL, catCol = NULL, check.for.linking = TRUE, minNperItem = 50, removeMinNperItem = FALSE,
boundary = 6, remove.boundary = FALSE, remove.no.answers = TRUE, remove.no.answersHG = TRUE,
remove.missing.items = TRUE, remove.constant.items = TRUE, remove.failures = FALSE,
remove.vars.DIF.missing = TRUE, remove.vars.DIF.constant = TRUE,
verbose=TRUE, software = c("conquest","tam", "mirt"), dir = NULL, analysis.name,
schooltype.var = NULL, model.statement = "item", compute.fit = TRUE,
pvMethod = c("regular", "bayesian"), fitTamMmlForBayesian = TRUE,
n.plausible=5, seed = NULL, conquest.folder= NULL,
constraints=c("cases","none","items"), std.err=c("quick","full","none"), distribution=c("normal","discrete"),
method=c("gauss", "quadrature", "montecarlo", "quasiMontecarlo"), n.iterations=2000,
nodes=NULL, p.nodes=2000, f.nodes=2000,converge=0.001,deviancechange=0.0001,
equivalence.table=c("wle","mle","NULL"), use.letters=FALSE,
allowAllScoresEverywhere = TRUE, guessMat = NULL, est.slopegroups = NULL,
fixSlopeMat = NULL, slopeMatDomainCol=NULL, slopeMatItemCol=NULL, slopeMatValueCol=NULL,
progress = NULL, Msteps = NULL, increment.factor=1 , fac.oldxsi=0,
export = list(logfile = TRUE, systemfile = FALSE, history = TRUE,
covariance = TRUE, reg_coefficients = TRUE, designmatrix = FALSE))Arguments
- dat
A data frame containing all variables necessary for analysis.
- items
Names or column numbers of variables with item responses. Item response values must be numeric (i.e. 0, 1, 2, 3 ... ). Character values (i.e. A, B, C ... or a, b, c, ...) are not allowed. Class of item columns are expected to be numeric or integer. Columns of class
characterwill be transformed.- id
Name or column number of the identifier (ID) variable.
- splittedModels
Optional: Object returned by
splitModels. Definition for multiple model handling.- irtmodel
If
software = "conquest", the argument is ignored as the IRT model is specified with themodel.statementargument. Ifsoftware = "tam", the argumentirtmodelcorresponds to theirtmodelargument oftam.mml. See the help page oftam.mmlfor further details. Ifsoftware = "mirt",irtmodelis a data.frame with two columns. First column: item identifier. Second column: Type of response model for this item. See theitemtypeargument in the help file ofmirtfor further details. Additionally, see example 10 below for specifying a mixed model with Rasch-, 2pl, partial credit, and generalized partial credit items.- qMatrix
Optional: A named data frame indicating how items should be grouped to dimensions. The first column contains the unique names of all items and should be named “item”. The other columns contain dimension definitions and should be named with the respective dimension names. A positive value (e.g., 1 or 2 or 1.4) indicates the loading weight with which an item loads on the dimension, a value of 0 indicates that the respective item does not load on this dimension. If no q matrix is specified by the user, an unidimensional structure is assumed.
- DIF.var
Name or column number of one grouping variable for which differential item functioning analysis is to be done.
- HG.var
Optional: Names or column numbers of one or more context variables (e.g., sex, school). These variables will be used for latent regression model in Conquest or TAM.
- group.var
Applies only if
software = "tam". Optional: Optional: Names or column numbers of one or more grouping variables. Descriptive statistics for WLEs and Plausible Values will be computed separately for each group in Conquest.- weight.var
Optional: Name or column number of one weighting variable.
- anchor
Optional: A named data frame with anchor parameters. If the data.frame only has two columns, the columns do not need to be named. In this case, the item name should be in the first column. The second column contains anchor parameters. Anchor items can be a subset of the items in the dataset and vice versa. If the data frame contains more than two columns, columns must be named explicitly using the following arguments
domainCol,itemCol,valueCol, andcatCol.- domainCol
Optional: Only necessary if the
anchorargument was used to define anchor parameters. Moreover, specifyingdomainColis only necessary, if the item identifiers inanchorare not unique—for example, if a specific item occurs with two parameters, one domain-specific item parameter and one additional “global” item parameter. The domain column than must specify which parameter belongs to which domain.- itemCol
Optional: Only necessary if the
anchorargument was used to define anchor parameters. Moreover, specifyingitemColis only necessary, if theanchordata frame has more than two columns. TheitemColcolumn then must specify which column contains the item identifier.- valueCol
Optional: Only necessary if the
anchorargument was used to define anchor parameters. Moreover, specifyingvalueColis only necessary, if theanchordata frame has more than two columns. ThevalueColcolumn then must specify which column contains the item parameter values.- catCol
Optional: Only necessary if the
anchorargument was used to define anchor parameters in a partial credit model. ThecatColcolumn then must specify which column contains the item parameter category indices. The values in this column must be named according to the convention"Cat1","Cat2","Cat3", and so on.- check.for.linking
A logical value indicating whether the items in dataset are checked for being connected with each other via design.
- minNperItem
Numerical: A message is printed on console if an item has less valid values than the number defined in
minNperItem.- removeMinNperItem
Logical: Remove items with less valid responses than defined in
minNperItem?- boundary
Numerical: A message is printed on console if a subject has answered less than the number of items defined in boundary.
- remove.boundary
Logical: Remove subjects who have answered less items than defined in the
boundaryargument?- remove.no.answers
Logical: Should persons without any item responses being removed prior to analysis?
- remove.no.answersHG
Logical: Should persons without any responses on any background variable being removed prior to analysis?
- remove.missing.items
Logical: Should items without any item responses being removed prior to analysis?
- remove.constant.items
Logical: Should items without variance being removed prior to analysis?
- remove.failures
Logical: Should persons without any correct item response (i.e., only “0” responses) being removed prior to analysis?
- remove.vars.DIF.missing
Logical: Applies only in DIF analyses. Should items without any responses in at least one DIF group being removed prior to analyses? Note: Conquest may crash if these items remain in the data.
- remove.vars.DIF.constant
Logical: Applies only in DIF analyses. Should items without variance in at least one DIF group being removed prior to analyses? Note: Conquest may crash if these items remain in the data.
- verbose
A logical value indicating whether messages are printed on the R console.
- software
The desired estimation software for the analysis.
- dir
The directory in which the output will be written to. If
software = "conquest",dirmust be specified. Ifsoftware = "tam",diris not mandatory.- analysis.name
A character string specifying the analysis name. If
software = "conquest",analysis.namemust be specified. All Conquest input and output files will namedanalysis.namewith their corresponding extensions. Ifsoftware = "tam",analysis.nameis not mandatory. In the case of multiple models estimation,split.modelsautomatically definesanalysis.namefor each model.- schooltype.var
Optional: Name or column number of the variable indicating the school type (e.g. academic track, non-academic track). Only necessary if p values should be computed for each school type separately.
- model.statement
Optional: Applies only if
software = "conquest". A character string given the model statement in the Conquest syntax. If omitted, the statement is generated automatically with respect to the defined model.- compute.fit
Applies only if
software = "conquest". Compute item fit statistics?- pvMethod
Applies only if
software = "tam": Specifies whether PVs should be drawn regularly or using a Bayesian algorithm.- fitTamMmlForBayesian
Logical, applies only if
software = "tam": If PVs are drawn using a Bayesian algorithm, it is not necessary to fit the model viatam.mmlbefore.fitTamMmlForBayesianspecifies whether the model should be fitted before though. See the help page oftam.pv.mcmcfor further details.- n.plausible
The number of plausible values which are to be drawn from the conditioning model.
- seed
Optional: Set seed value for analysis. The value will be used in Conquest syntax file ('set seed'-statement, see conquest manual, p. 225) or in TAM (control$seed). Note that seed only occurs for stochastic integration.
koennen
- conquest.folder
Applies only if
software = "conquest". A character string with path and name of the Conquest console, for example"c:/programme/conquest/console_Feb2007.exe". In package version 0.7.24 and later, conquest executable file is included in the package, so the user needn't to specify this argument.- constraints
A character string specifying how the scale should be constrained. Possible options are
"cases"(default),"items"and"none". When anchor parameter are specified in anchor, constraints will be set to"none"automatically. InTAMthe option"none"is not allowed. (See the help file oftam.mmlfor further details.)- std.err
Applies only if
software = "conquest". A character string specifying which type of standard error should be estimated. Possible options are"full","quick"(default) and"none". See Conquest manual pp.167 for details on standard error estimation.- distribution
Applies only if
software = "conquest". A character string indicating the a priori trait distribution. Possible options are"normal"(default) and"discrete". See Conquest manual pp.167 for details on population distributions.- method
A character string indicating which method should be used for numerical or stochastic integration. Possible options are
"gauss"(Gauss-Hermite quadrature: default),"quadrature"(Bock/Aitken quadrature) and"montecarlo". See Conquest manual pp.167 for details on these methods. When usingsoftware = "tam","gauss"and"quadrature"essentially leads to numerical integration, i.e TAM is called withcontrol$snodes = 0and withcontrol$nodes = seq(-6,6,len=nn), wherennequals the number of nodes specified in thenodesargument ofdefineModel(see below). Whether or notcontrol$QMCis set to TRUE or FALSE, depends on whether"montecarlo"or"quasiMontecarlo"is chosen."montecarlo"leads to calling TAM withcontrol$QMC = FALSEandsnodes = nn, wherennequals the number of nodes specified in thenodesargument ofdefineModel."quasiMontecarlo"leads to calling TAM withcontrol$QMC = TRUEandsnodes = nn, wherennequals the number of nodes specified in thenodesargument ofdefineModel. To met thesoftware = "tam"default (Quasi Monte Carlo integration withcontrol$QMC = TRUE), usesoftware="tam", nodes = 21, method = "quasiMontecarlo".- n.iterations
An integer value specifying the maximum number of iterations for which estimation will proceed without improvement in the deviance.
- nodes
An integer value specifying the number of nodes to be used in the analysis. The default value is 20. When using
software = "tam", the value specified here leads to calling TAM withnodes = 20ANDsnodes = 0if"gauss"or"quadrature"or"quasiMontecarlo"was used in themethodargument. If"montecarlo"was used in themethodargument, the value specified here leads to calling TAM withcontrol$snodes = 20. For numerical integration, for example,method = "gauss"andnodes = 21(TAM default) may be appropriate. For quasi monte carlo integration,method = "quasiMontecarlo"andnodes = 1000may be appropriate (TAM authors recommend to use at least 1000 nodes).- p.nodes
Applies only if
software = "conquest". An integer value specifying the number of nodes that are used in the approximation of the posterior distributions, which are used in the drawing of plausible values and in the calculation of EAP estimates. The default value is 2000.- f.nodes
Applies only if
software = "conquest". An integer value specifying the number of nodes that are used in the approximation of the posterior distributions in the calculation of fit statistics. The default value is 2000.- converge
An integer value specifying the convergence criterion for parameter estimates. The estimation will terminate when the largest change in any parameter estimate between successive iterations of the EM algorithm is less than converge. The default value is 0.001.
- deviancechange
An integer value specifiying the convergence criterion for the deviance. The estimation will terminate when the change in the deviance between successive iterations of the EM algorithm is less than deviancechange. The default value is 0.0001.
- equivalence.table
Applies only if
software = "conquest". A character string specifying the type of equivalence table to print. To be more precise, the person estimator that is to be used to create the table must be specified here. Possible options are"wle"(default),"mle"and NULL. If NULL, no equivalence table is computed at all.- use.letters
Applies only if
software = "conquest". A logical value indicating whether item response values should be coded as letters. This option can be used in partial credit models comprising items with more than 10 categories to avoid response columns with width 2 in Conquest.- allowAllScoresEverywhere
Applies only if
software = "Conquest". Defines score statement generation in multidimensional polytomous models. Consider two dimensions, `reading' and `listening'. In `reading', values 0, 1, 2, 3 occur. In `listening', values 1, 2, 3, 4 occur. IfTRUE, values 0, 1, 2, 3, 4 are defined for both dimensions. Otherwise, values 0, 1, 2, 3 are defined for `reading', values 1, 2, 3, 4 are defined for `listening'.- guessMat
Applies only if
software = "tam"for 3PL models. A named data frame with two columns indicating for which items a common guessing parameter should be estimated. The first column contains the names of all items in the analysis and should be named"item". The second column is numerical (integer values recommended) and allocates the items to groups. For each group of items, a separate guessing parameter is estimated. If the value in the second columns equals zero, the guessing parameter is fixed to zero.- est.slopegroups
Applies only if
software = "tam"for 2PL models. Optionally, a named data frame with two columns indicating for which items a common discrimination parameter should be estimated. The first column contains the names of all items in the analysis and should be named"item". The second column is numerical (integer values recommended) and allocates the items to groups. For each group of items, a separate discrimination parameter is estimated. Without specifyingest.slopegroups, a discrimination parameter for each item is estimated.- fixSlopeMat
Applies only if
software = "tam"for 2PL models. Optionally, a named data frame with two columns indicating for which items a fixed discrimination should be assumed. The first column contains the names of the items which discrimination should be fixed. Note that item indicators should be unique—if not, use further argumentsslopeMatDomainCol,slopeMatItemColandslopeMatValueCol. The second column is numerical and contains the discrimination value. Note: To date, this works only for between item dimensionality models. Within item dimensionality models must be specified directly in TAM, using theB.fixedargument oftam.mml. Items which discrimation should be estimated should not occur in this data frame.- slopeMatDomainCol
Optional: Only necessary if the
fixSlopeMatargument was used to define fixed slope parameters. Moreover, specifyingslopeMatDomainColis only necessary, if the item identifiers infixSlopeMatare not unique—for example, if a specific item occurs with two slope parameters, one domain-specific item slope parameter and one additional “global” item parameter. The domain column than must specify which parameter belongs to which domain.- slopeMatItemCol
Optional: Only necessary if the
fixSlopeMatargument was used to define fixed slope parameters. Moreover, specifyingitemColis only necessary, if thefixSlopeMatdata frame has more than two columns. TheitemColcolumn then must specify which column contains the item identifier.- slopeMatValueCol
Optional: Only necessary if the
fixSlopeMatargument was used to define slope parameters. Moreover, specifyingvalueColis only necessary, if thefixSlopeMatdata frame has more than two columns. ThevalueColcolumn than must specify which column contains the item parameter values.- progress
Logical: Applies only if
software = "tam"orsoftware = "mirt". Print estimation progress messages on console? IfNULL(the default), progress is printed forsoftware = "mirt", but not forsoftware = "tam".- Msteps
Number of M steps for item parameter estimation. A high value of M steps could be helpful in cases of non-convergence. The default value is 4; the default for 3pl models is set to 10.
- increment.factor
Applies only if
software = "tam". Should only be varied if the model does not converge. See help page oftam.mmlfor further details.- fac.oldxsi
Applies only if
software = "tam". Should only be varied if the model does not converge. See help page oftam.mmlfor further details.- export
Applies only if
software = "conquest". Specifies which additional files should be written on hard disk.
Value
A list which contains information about the desired estimation. The list is intended for
further processing via runModel. Structure of the list varies depending on
whether multiple models were called using splitModels or not. If
splitModels was called, the number of elements in the list equals
the number of models defined via splitModels. Each element in the list is
a list with various elements:
- software
Character string of the software which is intended to use for the further estimation, i.e.
"conquest"or"tam"- qMatrix
The Q matrix allocating items to dimensions.
- all.Names
Named list of all relevant variables of the data set.
- dir
Character string of the directory the results are to be saved.
- analysis.name
Character string of the analysis' name.
- deskRes
Data frame with descriptives (e.g., p values) of the test items.
- discrim
Data frame with item discrimination values.
- perNA
The person identifiers of examinees which are excluded from the analysis due to solely missing values.
- per0
The person identifiers of examinees which have solely false responses. if
remove.failueswas set to be TRUE, these persons are excluded from the data set.- perA
The person identifiers of examinees which have solely correct responses.
- perExHG
The person identifiers of examinees which are excluded from the analysis due to missing values on explicit variables.
- itemsExcluded
Character string of items which were excluded, for example due to zero variance or solely missing values.
If software == "conquest", the output additionally includes the following elements:
- input
Character string of the path with Conquest input (cqc) file.
- conquest.folder
Character string of the path of the conquest executable file.
- model.name
Character string of the model name.
If software == "tam", the output additionally includes the following elements:
- anchor
Optional: data frame of anchor parameters (if anchor parameters were defined).
- daten
The prepared data for TAM analysis.
- irtmodel
Character string of the used IRT model.
- est.slopegroups
Applies for 2pl modeling. Information about which items share a common slope parameter.
- guessMat
Applies for 3pl modeling. Information about which items share a common guessing parameter.
- control
List of control parameters for TAM estimation.
- n.plausible
Desired number of plausible values.
Examples
################################################################################
### Preparation: necessary for all examples ###
################################################################################
# load example data
# data set 'trends' contains item response data for three measurement occasions
# in a single data.frame
data(trends)
# second, create the q matrix from long format data
qMat <- unique(trends[ ,c("item","domain")])
qMat <- data.frame ( qMat[,"item", drop=FALSE], model.matrix(~domain-1, data = qMat))
################################################################################
### Example 1: Unidimensional Rasch Model ###
################################################################################
# Example 1: define and run a unidimensional Rasch model for the 2010 cohort
# with all variables from booklet 02 (this booklet contains reading and listening
# items together). For the estimation, Conquest is used.
# Since the wide-format dataset generated here will be used in later analyses,
# several covariates - such as gender and SES - are generated at the same time.
# These variables are not yet used in this example.
datW <- subset(trends, booklet == "Bo02" & year == 2010) |>
reshape2::dcast(idstud+language+sex+ses+country~item, value.var = "value")
# defining the unidimensional model: specifying q matrix is not necessary
mod1 <- defineModel(dat=datW, items= -c(1:5), id="idstud", analysis.name = "unidim", dir = tempdir())
#> Cannot find conquest 2007 executable file. Please choose manually.
#> Error in file.choose(): file choice cancelled
# run the model
run1 <- runModel(mod1)
#> Error: object 'mod1' not found
# get the results
res1 <- getResults(run1)
#> Error: object 'run1' not found
# extract the item parameters from the results object
item1<- itemFromRes(res1)
#> Error: object 'res1' not found
################################################################################
### Example 1a: Unidimensional Rasch Model with DIF estimation ###
################################################################################
# nearly the same procedure as in example 1. Using 'sex' as DIF variable
# note that 'sex' is a factor variable here. Conquest needs all explicit variables
# to be numeric. Variables will be automatically transformed to numeric by
# 'defineModels'. However, it might be the better idea to transform the variable
# manually.
datW[,"sexNum"] <- car::recode ( datW[,"sex"] , "'male'=0; 'female'=1", as.factor = FALSE)
# as we have defined a new variable ('sexNum') in the data, it is a good idea
# to explicitly specify item columns ... instead of saying 'items= -c(1:5)' which
# means: Everything except column 1 to 5 are item columns
items<- grep("^T[[:digit:]]{2}", colnames(datW))
mod1a<- defineModel(dat=datW, items= items, id="idstud", DIF.var = "sexNum",
analysis.name = "unidimDIF", dir = tempdir())
#> Cannot find conquest 2007 executable file. Please choose manually.
#> Error in file.choose(): file choice cancelled
# run the model
run1a<- runModel(mod1a)
#> Error: object 'mod1a' not found
# get the results
res1a<- getResults(run1a)
#> Error: object 'run1a' not found
################################################################################
### Example 1b: Rasch Model with DIF variable in customized model statement ###
################################################################################
# builing on example 1a, the DIF variable should be used without interaction term
# and without eliminating items which do not vary within DIF groups. We need the
# 'sexNum' variable which was defined previously
# Example based on 'model 6' in 'r:\Nawi\main\10_Studien\Auswertung\Alex Robitzsch R-Skripte\skalierung_conquest_mit_R_04.R'
items<- grep("^T[[:digit:]]{2}", colnames(datW))
mod1b<- defineModel(dat=datW, items= items, id="idstud", model.statement = "item + sexNum",
analysis.name = "unidimCustom", dir = tempdir())
#> Cannot find conquest 2007 executable file. Please choose manually.
#> Error in file.choose(): file choice cancelled
# run the model
run1b<- runModel(mod1b)
#> Error: object 'mod1b' not found
# get the results
res1b<- getResults(run1b)
#> Error: object 'run1b' not found
# item parameter
it1b <- itemFromRes(res1b)
#> Error: object 'res1b' not found
################################################################################
### Example 2a: Multidimensional Rasch Model with anchoring ###
################################################################################
# Example 2a: running a multidimensional Rasch model on a subset of items with latent
# regression. Use item parameter from the first model as anchor parameters
# read in anchor parameters from the results object of the first example
aPar <- itemFromRes(res1)[,c("item", "est")]
#> Error: object 'res1' not found
# defining the model: specifying q matrix now is necessary.
# Please note that all latent regression variables have to be of class numeric.
# If regression variables are factors, dummy variables automatically will be used.
# (This behavior is equivalent as in lm() for example.)
mod2a<- defineModel(dat=datW, items= grep("^T[[:digit:]]{2}", colnames(datW)),
id="idstud", analysis.name = "twodim", HG.var = c("language","sex", "ses"),
qMatrix = qMat, anchor = aPar, n.plausible = 20,dir = tempdir())
#> Error: object 'aPar' not found
# run the model
run2a<- runModel(mod2a)
#> Error: object 'mod2a' not found
# get the results
res2a<- getResults(run2a)
#> Error: object 'run2a' not found
################################################################################
### Example 2b: Multidimensional Rasch Model with equating ###
################################################################################
# Example 2b: running a multidimensional Rasch model on a subset of items
# without anchoring. Defining the model: specifying q matrix now is necessary.
mod2b<- defineModel(dat=datW, items= grep("^T[[:digit:]]{2}", colnames(datW)),
id="idstud", analysis.name = "twodim2", qMatrix = qMat,
n.plausible = 20, dir = tempdir())
#> Cannot find conquest 2007 executable file. Please choose manually.
#> Error in file.choose(): file choice cancelled
# run the model
run2b <- runModel(mod2b)
#> Error: object 'mod2b' not found
# get the results
res2b<- getResults(run2b)
#> Error: object 'run2b' not found
### equating (wenn nicht verankert)
eq2b <- equat1pl( results = res2b, prmNorm = aPar, difBound=.64, iterativ=TRUE)
#> Error: object 'res2b' not found
### transformation to the 'bista' metric: needs reference population definition
### we use some arbitrary values here
ref <- data.frame ( domain = c("domainreading", "domainlistening"),
m = c(0.03890191, 0.03587727), sd= c(1.219, 0.8978912))
cuts <- list ( domainreading = list ( values = 390+0:3*75),
domainlistening = list ( values = 360+0:3*85))
tf2b <- transformToBista ( equatingList = eq2b, refPop = ref, cuts = cuts)
#> Error: object 'eq2b' not found
################################################################################
### Example 3: Multidimensional Rasch Model in TAM ###
################################################################################
# estimate model 2 with latent regression and anchored parameters in TAM
# specification of an output folder (via 'dir' argument) no longer necessary
mod3T<- defineModel(dat=datW, items= grep("^T[[:digit:]]{2}", colnames(datW)),
id="idstud", qMatrix = qMat, HG.var = "sex", anchor = aPar, software = "tam")
#> Error: object 'aPar' not found
# run the model
run3T<- runModel(mod3T)
#> Error: object 'mod3T' not found
# Object 'run2T' is of class 'tam.mml'
class(run3T)
#> Error: object 'run3T' not found
# the class of 'run2T' corresponds to the class defined by the TAM package; all
# functions of the TAM package intended for further processing (e.g. drawing
# plausible values, plotting deviance change etc.) work, for example:
wle <- tam.wle(run3T)
#> Error: object 'run3T' not found
# Finally, the model result are collected in a single data frame
res3T<- getResults(run3T)
#> Error: object 'run3T' not found
# repeat the same model with mirt
items<- grep("^T[[:digit:]]{2}", colnames(datW), value=TRUE)
irtM <- data.frame(item = items, task = substr(items,1,4), stringsAsFactors = FALSE) |> dplyr::mutate(irtmod = "Rasch")
mod3M<- defineModel(dat=datW, items= grep("^T[[:digit:]]{2}", colnames(datW)),id="idstud",
irtmodel = irtM[,c("item", "irtmod")], anchor = aPar, qMatrix = qMat, HG.var = "sex", software = "mirt")
#> Error: object 'aPar' not found
run3M<- runModel(mod3M)
#> Error: object 'mod3M' not found
res3M<- getResults(run3M)
#> Error: object 'run3M' not found
itM <- itemFromRes(res3M)
#> Error: object 'res3M' not found
################################################################################
### Example 4: define und run multiple models defined by 'splitModels' ###
################################################################################
# Example 4: define und run multiple models defined by 'splitModels'
# Model split is possible for different groups of items (i.e. domains) and/or
# different groups of persons (for example, federal states within Germany)
# define person grouping
pers <- data.frame ( idstud = datW[,"idstud"] , group1 = datW[,"sex"],
group2 = datW[,"country"], stringsAsFactors = FALSE )
# define 24 models, splitting according to person groups and item groups separately
# by default, multicore processing is applied
l1 <- splitModels ( qMatrix = qMat, person.groups = pers, nCores = 1)
#> ---------------------------------
#> splitModels: generating 24 models
#> ........................
#> see <returned>$models
#> number of cores: 1
#> ---------------------------------
# In this example, the sample sizes in the individual subgroups or models are far
# too small. The example is therefore intended solely to illustrate how the
# analysis can be broken down by item and respondent group.
table(datW[,c("sex", "country")])
#> country
#> sex countryA countryB countryC
#> female 42 27 20
#> male 32 35 16
# apply 'defineModel' for each of the 24 models in 'l1'
modMul<- defineModel(dat = datW, items = grep("^T[[:digit:]]{2}", colnames(datW)),
id = "idstud", check.for.linking = TRUE, splittedModels = l1, software = "tam")
#>
#> Specification of 'qMatrix' and 'person.groups' results in 24 model(s).
#> Warning: Model split preparation for model No. 1, model name
#> domainlistening__group1.female_group2.countryC: 118 items from 134 items listed
#> the Q matrix not found in data:
#> ℹ 'T12_11', 'T13_15', 'T15_03', 'T13_04', 'T13_07', 'T13_16', 'T13_01',
#> 'T15_02', 'T15_01', 'T15_15', 'T15_16', 'T15_09', 'T13_12', 'T13_13',
#> 'T15_14', 'T13_08', 'T13_06', 'T15_12', 'T13_10', 'T13_17', 'T15_11',
#> 'T13_05', 'T13_09', 'T15_10', 'T15_13', 'T15_17', 'T15_18', 'T13_18',
#> 'T16_08', 'T16_09', 'T16_11', 'T16_13', 'T16_01', 'T16_04', 'T16_10',
#> 'T16_02', 'T16_12', 'T16_06', 'T16_07', 'T19_06', 'T19_08', 'T19_05',
#> 'T19_11', 'T19_01', 'T19_02', 'T19_30', 'T19_03', 'T19_07', 'T19_04',
#> 'T19_09', 'T21_11', 'T18_01', 'T21_01', 'T18_05', 'T21_12', 'T18_13',
#> 'T21_10', 'T18_11', 'T21_05', 'T21_13', 'T18_08', 'T18_07', 'T21_09',
#> 'T18_04', 'T18_02', 'T21_02', 'T21_06', 'T18_09', 'T21_04', 'T18_12',
#> 'T21_14', 'T21_03', 'T18_10', 'T30_06', 'T29_04', 'T29_02', 'T30_02',
#> 'T30_03', 'T30_01', 'T29_05', 'T29_03', 'T28_08', 'T28_09', 'T28_02',
#> 'T28_03', 'T28_04', 'T28_01', 'T28_10', 'T28_06', 'T20_05', 'T20_09',
#> 'T20_30', 'T20_08', 'T20_12', 'T20_06', 'T20_07', 'T20_02', 'T20_13',
#> 'T20_03', 'T20_01', 'T20_04', 'T20_11', 'T20_10', 'T32_04', 'T31_03',
#> 'T31_01', 'T31_04', 'T32_03', 'T32_02', 'T32_01', 'T17_02', 'T17_09',
#> 'T17_10', 'T17_04', 'T17_03', 'T17_08', 'T17_06', 'T17_01'
#> Warning: Model split preparation for model No. 2, model name
#> domainlistening__group1.female_group2.countryA: 118 items from 134 items listed
#> the Q matrix not found in data:
#> ℹ 'T12_11', 'T13_15', 'T15_03', 'T13_04', 'T13_07', 'T13_16', 'T13_01',
#> 'T15_02', 'T15_01', 'T15_15', 'T15_16', 'T15_09', 'T13_12', 'T13_13',
#> 'T15_14', 'T13_08', 'T13_06', 'T15_12', 'T13_10', 'T13_17', 'T15_11',
#> 'T13_05', 'T13_09', 'T15_10', 'T15_13', 'T15_17', 'T15_18', 'T13_18',
#> 'T16_08', 'T16_09', 'T16_11', 'T16_13', 'T16_01', 'T16_04', 'T16_10',
#> 'T16_02', 'T16_12', 'T16_06', 'T16_07', 'T19_06', 'T19_08', 'T19_05',
#> 'T19_11', 'T19_01', 'T19_02', 'T19_30', 'T19_03', 'T19_07', 'T19_04',
#> 'T19_09', 'T21_11', 'T18_01', 'T21_01', 'T18_05', 'T21_12', 'T18_13',
#> 'T21_10', 'T18_11', 'T21_05', 'T21_13', 'T18_08', 'T18_07', 'T21_09',
#> 'T18_04', 'T18_02', 'T21_02', 'T21_06', 'T18_09', 'T21_04', 'T18_12',
#> 'T21_14', 'T21_03', 'T18_10', 'T30_06', 'T29_04', 'T29_02', 'T30_02',
#> 'T30_03', 'T30_01', 'T29_05', 'T29_03', 'T28_08', 'T28_09', 'T28_02',
#> 'T28_03', 'T28_04', 'T28_01', 'T28_10', 'T28_06', 'T20_05', 'T20_09',
#> 'T20_30', 'T20_08', 'T20_12', 'T20_06', 'T20_07', 'T20_02', 'T20_13',
#> 'T20_03', 'T20_01', 'T20_04', 'T20_11', 'T20_10', 'T32_04', 'T31_03',
#> 'T31_01', 'T31_04', 'T32_03', 'T32_02', 'T32_01', 'T17_02', 'T17_09',
#> 'T17_10', 'T17_04', 'T17_03', 'T17_08', 'T17_06', 'T17_01'
#> Warning: Model split preparation for model No. 3, model name
#> domainlistening__group1.female_group2.countryB: 118 items from 134 items listed
#> the Q matrix not found in data:
#> ℹ 'T12_11', 'T13_15', 'T15_03', 'T13_04', 'T13_07', 'T13_16', 'T13_01',
#> 'T15_02', 'T15_01', 'T15_15', 'T15_16', 'T15_09', 'T13_12', 'T13_13',
#> 'T15_14', 'T13_08', 'T13_06', 'T15_12', 'T13_10', 'T13_17', 'T15_11',
#> 'T13_05', 'T13_09', 'T15_10', 'T15_13', 'T15_17', 'T15_18', 'T13_18',
#> 'T16_08', 'T16_09', 'T16_11', 'T16_13', 'T16_01', 'T16_04', 'T16_10',
#> 'T16_02', 'T16_12', 'T16_06', 'T16_07', 'T19_06', 'T19_08', 'T19_05',
#> 'T19_11', 'T19_01', 'T19_02', 'T19_30', 'T19_03', 'T19_07', 'T19_04',
#> 'T19_09', 'T21_11', 'T18_01', 'T21_01', 'T18_05', 'T21_12', 'T18_13',
#> 'T21_10', 'T18_11', 'T21_05', 'T21_13', 'T18_08', 'T18_07', 'T21_09',
#> 'T18_04', 'T18_02', 'T21_02', 'T21_06', 'T18_09', 'T21_04', 'T18_12',
#> 'T21_14', 'T21_03', 'T18_10', 'T30_06', 'T29_04', 'T29_02', 'T30_02',
#> 'T30_03', 'T30_01', 'T29_05', 'T29_03', 'T28_08', 'T28_09', 'T28_02',
#> 'T28_03', 'T28_04', 'T28_01', 'T28_10', 'T28_06', 'T20_05', 'T20_09',
#> 'T20_30', 'T20_08', 'T20_12', 'T20_06', 'T20_07', 'T20_02', 'T20_13',
#> 'T20_03', 'T20_01', 'T20_04', 'T20_11', 'T20_10', 'T32_04', 'T31_03',
#> 'T31_01', 'T31_04', 'T32_03', 'T32_02', 'T32_01', 'T17_02', 'T17_09',
#> 'T17_10', 'T17_04', 'T17_03', 'T17_08', 'T17_06', 'T17_01'
#> Warning: Model split preparation for model No. 4, model name
#> domainlistening__group1.female_group2.all: 118 items from 134 items listed the
#> Q matrix not found in data:
#> ℹ 'T12_11', 'T13_15', 'T15_03', 'T13_04', 'T13_07', 'T13_16', 'T13_01',
#> 'T15_02', 'T15_01', 'T15_15', 'T15_16', 'T15_09', 'T13_12', 'T13_13',
#> 'T15_14', 'T13_08', 'T13_06', 'T15_12', 'T13_10', 'T13_17', 'T15_11',
#> 'T13_05', 'T13_09', 'T15_10', 'T15_13', 'T15_17', 'T15_18', 'T13_18',
#> 'T16_08', 'T16_09', 'T16_11', 'T16_13', 'T16_01', 'T16_04', 'T16_10',
#> 'T16_02', 'T16_12', 'T16_06', 'T16_07', 'T19_06', 'T19_08', 'T19_05',
#> 'T19_11', 'T19_01', 'T19_02', 'T19_30', 'T19_03', 'T19_07', 'T19_04',
#> 'T19_09', 'T21_11', 'T18_01', 'T21_01', 'T18_05', 'T21_12', 'T18_13',
#> 'T21_10', 'T18_11', 'T21_05', 'T21_13', 'T18_08', 'T18_07', 'T21_09',
#> 'T18_04', 'T18_02', 'T21_02', 'T21_06', 'T18_09', 'T21_04', 'T18_12',
#> 'T21_14', 'T21_03', 'T18_10', 'T30_06', 'T29_04', 'T29_02', 'T30_02',
#> 'T30_03', 'T30_01', 'T29_05', 'T29_03', 'T28_08', 'T28_09', 'T28_02',
#> 'T28_03', 'T28_04', 'T28_01', 'T28_10', 'T28_06', 'T20_05', 'T20_09',
#> 'T20_30', 'T20_08', 'T20_12', 'T20_06', 'T20_07', 'T20_02', 'T20_13',
#> 'T20_03', 'T20_01', 'T20_04', 'T20_11', 'T20_10', 'T32_04', 'T31_03',
#> 'T31_01', 'T31_04', 'T32_03', 'T32_02', 'T32_01', 'T17_02', 'T17_09',
#> 'T17_10', 'T17_04', 'T17_03', 'T17_08', 'T17_06', 'T17_01'
#> Warning: Model split preparation for model No. 5, model name
#> domainlistening__group1.male_group2.countryC: 118 items from 134 items listed
#> the Q matrix not found in data:
#> ℹ 'T12_11', 'T13_15', 'T15_03', 'T13_04', 'T13_07', 'T13_16', 'T13_01',
#> 'T15_02', 'T15_01', 'T15_15', 'T15_16', 'T15_09', 'T13_12', 'T13_13',
#> 'T15_14', 'T13_08', 'T13_06', 'T15_12', 'T13_10', 'T13_17', 'T15_11',
#> 'T13_05', 'T13_09', 'T15_10', 'T15_13', 'T15_17', 'T15_18', 'T13_18',
#> 'T16_08', 'T16_09', 'T16_11', 'T16_13', 'T16_01', 'T16_04', 'T16_10',
#> 'T16_02', 'T16_12', 'T16_06', 'T16_07', 'T19_06', 'T19_08', 'T19_05',
#> 'T19_11', 'T19_01', 'T19_02', 'T19_30', 'T19_03', 'T19_07', 'T19_04',
#> 'T19_09', 'T21_11', 'T18_01', 'T21_01', 'T18_05', 'T21_12', 'T18_13',
#> 'T21_10', 'T18_11', 'T21_05', 'T21_13', 'T18_08', 'T18_07', 'T21_09',
#> 'T18_04', 'T18_02', 'T21_02', 'T21_06', 'T18_09', 'T21_04', 'T18_12',
#> 'T21_14', 'T21_03', 'T18_10', 'T30_06', 'T29_04', 'T29_02', 'T30_02',
#> 'T30_03', 'T30_01', 'T29_05', 'T29_03', 'T28_08', 'T28_09', 'T28_02',
#> 'T28_03', 'T28_04', 'T28_01', 'T28_10', 'T28_06', 'T20_05', 'T20_09',
#> 'T20_30', 'T20_08', 'T20_12', 'T20_06', 'T20_07', 'T20_02', 'T20_13',
#> 'T20_03', 'T20_01', 'T20_04', 'T20_11', 'T20_10', 'T32_04', 'T31_03',
#> 'T31_01', 'T31_04', 'T32_03', 'T32_02', 'T32_01', 'T17_02', 'T17_09',
#> 'T17_10', 'T17_04', 'T17_03', 'T17_08', 'T17_06', 'T17_01'
#> Warning: Model split preparation for model No. 6, model name
#> domainlistening__group1.male_group2.countryA: 118 items from 134 items listed
#> the Q matrix not found in data:
#> ℹ 'T12_11', 'T13_15', 'T15_03', 'T13_04', 'T13_07', 'T13_16', 'T13_01',
#> 'T15_02', 'T15_01', 'T15_15', 'T15_16', 'T15_09', 'T13_12', 'T13_13',
#> 'T15_14', 'T13_08', 'T13_06', 'T15_12', 'T13_10', 'T13_17', 'T15_11',
#> 'T13_05', 'T13_09', 'T15_10', 'T15_13', 'T15_17', 'T15_18', 'T13_18',
#> 'T16_08', 'T16_09', 'T16_11', 'T16_13', 'T16_01', 'T16_04', 'T16_10',
#> 'T16_02', 'T16_12', 'T16_06', 'T16_07', 'T19_06', 'T19_08', 'T19_05',
#> 'T19_11', 'T19_01', 'T19_02', 'T19_30', 'T19_03', 'T19_07', 'T19_04',
#> 'T19_09', 'T21_11', 'T18_01', 'T21_01', 'T18_05', 'T21_12', 'T18_13',
#> 'T21_10', 'T18_11', 'T21_05', 'T21_13', 'T18_08', 'T18_07', 'T21_09',
#> 'T18_04', 'T18_02', 'T21_02', 'T21_06', 'T18_09', 'T21_04', 'T18_12',
#> 'T21_14', 'T21_03', 'T18_10', 'T30_06', 'T29_04', 'T29_02', 'T30_02',
#> 'T30_03', 'T30_01', 'T29_05', 'T29_03', 'T28_08', 'T28_09', 'T28_02',
#> 'T28_03', 'T28_04', 'T28_01', 'T28_10', 'T28_06', 'T20_05', 'T20_09',
#> 'T20_30', 'T20_08', 'T20_12', 'T20_06', 'T20_07', 'T20_02', 'T20_13',
#> 'T20_03', 'T20_01', 'T20_04', 'T20_11', 'T20_10', 'T32_04', 'T31_03',
#> 'T31_01', 'T31_04', 'T32_03', 'T32_02', 'T32_01', 'T17_02', 'T17_09',
#> 'T17_10', 'T17_04', 'T17_03', 'T17_08', 'T17_06', 'T17_01'
#> Warning: Model split preparation for model No. 7, model name
#> domainlistening__group1.male_group2.countryB: 118 items from 134 items listed
#> the Q matrix not found in data:
#> ℹ 'T12_11', 'T13_15', 'T15_03', 'T13_04', 'T13_07', 'T13_16', 'T13_01',
#> 'T15_02', 'T15_01', 'T15_15', 'T15_16', 'T15_09', 'T13_12', 'T13_13',
#> 'T15_14', 'T13_08', 'T13_06', 'T15_12', 'T13_10', 'T13_17', 'T15_11',
#> 'T13_05', 'T13_09', 'T15_10', 'T15_13', 'T15_17', 'T15_18', 'T13_18',
#> 'T16_08', 'T16_09', 'T16_11', 'T16_13', 'T16_01', 'T16_04', 'T16_10',
#> 'T16_02', 'T16_12', 'T16_06', 'T16_07', 'T19_06', 'T19_08', 'T19_05',
#> 'T19_11', 'T19_01', 'T19_02', 'T19_30', 'T19_03', 'T19_07', 'T19_04',
#> 'T19_09', 'T21_11', 'T18_01', 'T21_01', 'T18_05', 'T21_12', 'T18_13',
#> 'T21_10', 'T18_11', 'T21_05', 'T21_13', 'T18_08', 'T18_07', 'T21_09',
#> 'T18_04', 'T18_02', 'T21_02', 'T21_06', 'T18_09', 'T21_04', 'T18_12',
#> 'T21_14', 'T21_03', 'T18_10', 'T30_06', 'T29_04', 'T29_02', 'T30_02',
#> 'T30_03', 'T30_01', 'T29_05', 'T29_03', 'T28_08', 'T28_09', 'T28_02',
#> 'T28_03', 'T28_04', 'T28_01', 'T28_10', 'T28_06', 'T20_05', 'T20_09',
#> 'T20_30', 'T20_08', 'T20_12', 'T20_06', 'T20_07', 'T20_02', 'T20_13',
#> 'T20_03', 'T20_01', 'T20_04', 'T20_11', 'T20_10', 'T32_04', 'T31_03',
#> 'T31_01', 'T31_04', 'T32_03', 'T32_02', 'T32_01', 'T17_02', 'T17_09',
#> 'T17_10', 'T17_04', 'T17_03', 'T17_08', 'T17_06', 'T17_01'
#> Warning: Model split preparation for model No. 8, model name
#> domainlistening__group1.male_group2.all: 118 items from 134 items listed the Q
#> matrix not found in data:
#> ℹ 'T12_11', 'T13_15', 'T15_03', 'T13_04', 'T13_07', 'T13_16', 'T13_01',
#> 'T15_02', 'T15_01', 'T15_15', 'T15_16', 'T15_09', 'T13_12', 'T13_13',
#> 'T15_14', 'T13_08', 'T13_06', 'T15_12', 'T13_10', 'T13_17', 'T15_11',
#> 'T13_05', 'T13_09', 'T15_10', 'T15_13', 'T15_17', 'T15_18', 'T13_18',
#> 'T16_08', 'T16_09', 'T16_11', 'T16_13', 'T16_01', 'T16_04', 'T16_10',
#> 'T16_02', 'T16_12', 'T16_06', 'T16_07', 'T19_06', 'T19_08', 'T19_05',
#> 'T19_11', 'T19_01', 'T19_02', 'T19_30', 'T19_03', 'T19_07', 'T19_04',
#> 'T19_09', 'T21_11', 'T18_01', 'T21_01', 'T18_05', 'T21_12', 'T18_13',
#> 'T21_10', 'T18_11', 'T21_05', 'T21_13', 'T18_08', 'T18_07', 'T21_09',
#> 'T18_04', 'T18_02', 'T21_02', 'T21_06', 'T18_09', 'T21_04', 'T18_12',
#> 'T21_14', 'T21_03', 'T18_10', 'T30_06', 'T29_04', 'T29_02', 'T30_02',
#> 'T30_03', 'T30_01', 'T29_05', 'T29_03', 'T28_08', 'T28_09', 'T28_02',
#> 'T28_03', 'T28_04', 'T28_01', 'T28_10', 'T28_06', 'T20_05', 'T20_09',
#> 'T20_30', 'T20_08', 'T20_12', 'T20_06', 'T20_07', 'T20_02', 'T20_13',
#> 'T20_03', 'T20_01', 'T20_04', 'T20_11', 'T20_10', 'T32_04', 'T31_03',
#> 'T31_01', 'T31_04', 'T32_03', 'T32_02', 'T32_01', 'T17_02', 'T17_09',
#> 'T17_10', 'T17_04', 'T17_03', 'T17_08', 'T17_06', 'T17_01'
#> Warning: Model split preparation for model No. 9, model name
#> domainlistening__group1.all_group2.countryC: 118 items from 134 items listed
#> the Q matrix not found in data:
#> ℹ 'T12_11', 'T13_15', 'T15_03', 'T13_04', 'T13_07', 'T13_16', 'T13_01',
#> 'T15_02', 'T15_01', 'T15_15', 'T15_16', 'T15_09', 'T13_12', 'T13_13',
#> 'T15_14', 'T13_08', 'T13_06', 'T15_12', 'T13_10', 'T13_17', 'T15_11',
#> 'T13_05', 'T13_09', 'T15_10', 'T15_13', 'T15_17', 'T15_18', 'T13_18',
#> 'T16_08', 'T16_09', 'T16_11', 'T16_13', 'T16_01', 'T16_04', 'T16_10',
#> 'T16_02', 'T16_12', 'T16_06', 'T16_07', 'T19_06', 'T19_08', 'T19_05',
#> 'T19_11', 'T19_01', 'T19_02', 'T19_30', 'T19_03', 'T19_07', 'T19_04',
#> 'T19_09', 'T21_11', 'T18_01', 'T21_01', 'T18_05', 'T21_12', 'T18_13',
#> 'T21_10', 'T18_11', 'T21_05', 'T21_13', 'T18_08', 'T18_07', 'T21_09',
#> 'T18_04', 'T18_02', 'T21_02', 'T21_06', 'T18_09', 'T21_04', 'T18_12',
#> 'T21_14', 'T21_03', 'T18_10', 'T30_06', 'T29_04', 'T29_02', 'T30_02',
#> 'T30_03', 'T30_01', 'T29_05', 'T29_03', 'T28_08', 'T28_09', 'T28_02',
#> 'T28_03', 'T28_04', 'T28_01', 'T28_10', 'T28_06', 'T20_05', 'T20_09',
#> 'T20_30', 'T20_08', 'T20_12', 'T20_06', 'T20_07', 'T20_02', 'T20_13',
#> 'T20_03', 'T20_01', 'T20_04', 'T20_11', 'T20_10', 'T32_04', 'T31_03',
#> 'T31_01', 'T31_04', 'T32_03', 'T32_02', 'T32_01', 'T17_02', 'T17_09',
#> 'T17_10', 'T17_04', 'T17_03', 'T17_08', 'T17_06', 'T17_01'
#> Warning: Model split preparation for model No. 10, model name
#> domainlistening__group1.all_group2.countryA: 118 items from 134 items listed
#> the Q matrix not found in data:
#> ℹ 'T12_11', 'T13_15', 'T15_03', 'T13_04', 'T13_07', 'T13_16', 'T13_01',
#> 'T15_02', 'T15_01', 'T15_15', 'T15_16', 'T15_09', 'T13_12', 'T13_13',
#> 'T15_14', 'T13_08', 'T13_06', 'T15_12', 'T13_10', 'T13_17', 'T15_11',
#> 'T13_05', 'T13_09', 'T15_10', 'T15_13', 'T15_17', 'T15_18', 'T13_18',
#> 'T16_08', 'T16_09', 'T16_11', 'T16_13', 'T16_01', 'T16_04', 'T16_10',
#> 'T16_02', 'T16_12', 'T16_06', 'T16_07', 'T19_06', 'T19_08', 'T19_05',
#> 'T19_11', 'T19_01', 'T19_02', 'T19_30', 'T19_03', 'T19_07', 'T19_04',
#> 'T19_09', 'T21_11', 'T18_01', 'T21_01', 'T18_05', 'T21_12', 'T18_13',
#> 'T21_10', 'T18_11', 'T21_05', 'T21_13', 'T18_08', 'T18_07', 'T21_09',
#> 'T18_04', 'T18_02', 'T21_02', 'T21_06', 'T18_09', 'T21_04', 'T18_12',
#> 'T21_14', 'T21_03', 'T18_10', 'T30_06', 'T29_04', 'T29_02', 'T30_02',
#> 'T30_03', 'T30_01', 'T29_05', 'T29_03', 'T28_08', 'T28_09', 'T28_02',
#> 'T28_03', 'T28_04', 'T28_01', 'T28_10', 'T28_06', 'T20_05', 'T20_09',
#> 'T20_30', 'T20_08', 'T20_12', 'T20_06', 'T20_07', 'T20_02', 'T20_13',
#> 'T20_03', 'T20_01', 'T20_04', 'T20_11', 'T20_10', 'T32_04', 'T31_03',
#> 'T31_01', 'T31_04', 'T32_03', 'T32_02', 'T32_01', 'T17_02', 'T17_09',
#> 'T17_10', 'T17_04', 'T17_03', 'T17_08', 'T17_06', 'T17_01'
#> Warning: Model split preparation for model No. 11, model name
#> domainlistening__group1.all_group2.countryB: 118 items from 134 items listed
#> the Q matrix not found in data:
#> ℹ 'T12_11', 'T13_15', 'T15_03', 'T13_04', 'T13_07', 'T13_16', 'T13_01',
#> 'T15_02', 'T15_01', 'T15_15', 'T15_16', 'T15_09', 'T13_12', 'T13_13',
#> 'T15_14', 'T13_08', 'T13_06', 'T15_12', 'T13_10', 'T13_17', 'T15_11',
#> 'T13_05', 'T13_09', 'T15_10', 'T15_13', 'T15_17', 'T15_18', 'T13_18',
#> 'T16_08', 'T16_09', 'T16_11', 'T16_13', 'T16_01', 'T16_04', 'T16_10',
#> 'T16_02', 'T16_12', 'T16_06', 'T16_07', 'T19_06', 'T19_08', 'T19_05',
#> 'T19_11', 'T19_01', 'T19_02', 'T19_30', 'T19_03', 'T19_07', 'T19_04',
#> 'T19_09', 'T21_11', 'T18_01', 'T21_01', 'T18_05', 'T21_12', 'T18_13',
#> 'T21_10', 'T18_11', 'T21_05', 'T21_13', 'T18_08', 'T18_07', 'T21_09',
#> 'T18_04', 'T18_02', 'T21_02', 'T21_06', 'T18_09', 'T21_04', 'T18_12',
#> 'T21_14', 'T21_03', 'T18_10', 'T30_06', 'T29_04', 'T29_02', 'T30_02',
#> 'T30_03', 'T30_01', 'T29_05', 'T29_03', 'T28_08', 'T28_09', 'T28_02',
#> 'T28_03', 'T28_04', 'T28_01', 'T28_10', 'T28_06', 'T20_05', 'T20_09',
#> 'T20_30', 'T20_08', 'T20_12', 'T20_06', 'T20_07', 'T20_02', 'T20_13',
#> 'T20_03', 'T20_01', 'T20_04', 'T20_11', 'T20_10', 'T32_04', 'T31_03',
#> 'T31_01', 'T31_04', 'T32_03', 'T32_02', 'T32_01', 'T17_02', 'T17_09',
#> 'T17_10', 'T17_04', 'T17_03', 'T17_08', 'T17_06', 'T17_01'
#> Warning: Model split preparation for model No. 12, model name
#> domainlistening__group1.all_group2.all: 118 items from 134 items listed the Q
#> matrix not found in data:
#> ℹ 'T12_11', 'T13_15', 'T15_03', 'T13_04', 'T13_07', 'T13_16', 'T13_01',
#> 'T15_02', 'T15_01', 'T15_15', 'T15_16', 'T15_09', 'T13_12', 'T13_13',
#> 'T15_14', 'T13_08', 'T13_06', 'T15_12', 'T13_10', 'T13_17', 'T15_11',
#> 'T13_05', 'T13_09', 'T15_10', 'T15_13', 'T15_17', 'T15_18', 'T13_18',
#> 'T16_08', 'T16_09', 'T16_11', 'T16_13', 'T16_01', 'T16_04', 'T16_10',
#> 'T16_02', 'T16_12', 'T16_06', 'T16_07', 'T19_06', 'T19_08', 'T19_05',
#> 'T19_11', 'T19_01', 'T19_02', 'T19_30', 'T19_03', 'T19_07', 'T19_04',
#> 'T19_09', 'T21_11', 'T18_01', 'T21_01', 'T18_05', 'T21_12', 'T18_13',
#> 'T21_10', 'T18_11', 'T21_05', 'T21_13', 'T18_08', 'T18_07', 'T21_09',
#> 'T18_04', 'T18_02', 'T21_02', 'T21_06', 'T18_09', 'T21_04', 'T18_12',
#> 'T21_14', 'T21_03', 'T18_10', 'T30_06', 'T29_04', 'T29_02', 'T30_02',
#> 'T30_03', 'T30_01', 'T29_05', 'T29_03', 'T28_08', 'T28_09', 'T28_02',
#> 'T28_03', 'T28_04', 'T28_01', 'T28_10', 'T28_06', 'T20_05', 'T20_09',
#> 'T20_30', 'T20_08', 'T20_12', 'T20_06', 'T20_07', 'T20_02', 'T20_13',
#> 'T20_03', 'T20_01', 'T20_04', 'T20_11', 'T20_10', 'T32_04', 'T31_03',
#> 'T31_01', 'T31_04', 'T32_03', 'T32_02', 'T32_01', 'T17_02', 'T17_09',
#> 'T17_10', 'T17_04', 'T17_03', 'T17_08', 'T17_06', 'T17_01'
#> Warning: Model split preparation for model No. 13, model name
#> domainreading__group1.female_group2.countryC: 132 items from 149 items listed
#> the Q matrix not found in data:
#> ℹ 'T07_08', 'T02_02', 'T07_01', 'T02_06', 'T07_05', 'T07_06', 'T02_07',
#> 'T07_02', 'T07_04', 'T02_03', 'T07_09', 'T02_01', 'T02_04', 'T07_03',
#> 'T02_05', 'T07_07', 'T07_10', 'T09_12', 'T01_08', 'T03_07', 'T03_01',
#> 'T03_08', 'T03_05', 'T03_06', 'T03_03', 'T10_08', 'T11_06', 'T11_03',
#> 'T11_01', 'T04_02', 'T04_01', 'T04_06', 'T04_04', 'T11_04', 'T10_01',
#> 'T11_08', 'T10_02', 'T10_09', 'T10_07', 'T11_02', 'T04_05', 'T04_03',
#> 'T11_05', 'T10_06', 'T11_10', 'T11_09', 'T04_07', 'T10_10', 'T04_09',
#> 'T04_08', 'T11_0X', 'T06_05', 'T05_03', 'T05_06', 'T05_05', 'T06_02',
#> 'T06_04', 'T06_03', 'T05_04', 'T05_02', 'T05_01', 'T06_01', 'T08_04',
#> 'T08_06', 'T08_03', 'T08_07', 'T08_02', 'T08_05', 'T08_01', 'T08_09',
#> 'T08_08', 'T03_09', 'T06_1X', 'T26_03', 'T26_05', 'T26_07', 'T26_09',
#> 'T26_06', 'T26_10', 'T26_08', 'T26_02', 'T26_04', 'T27_01', 'T27_05',
#> 'T27_07', 'T27_06', 'T27_04', 'T27_02', 'T27_03', 'T24_05', 'T24_08',
#> 'T24_02', 'T24_10', 'T24_06', 'T24_24', 'T24_25', 'T24_09', 'T24_21',
#> 'T24_23', 'T23_10', 'T23_13', 'T23_09', 'T23_15', 'T23_03', 'T23_04',
#> 'T23_14', 'T23_07', 'T23_08', 'T23_02', 'T23_06', 'T23_05', 'T25_05',
#> 'T25_22', 'T22_03', 'T25_06', 'T22_12', 'T25_09', 'T25_10', 'T25_08',
#> 'T22_11', 'T22_02', 'T25_11', 'T25_07', 'T25_14', 'T25_12', 'T25_04',
#> 'T22_08', 'T25_23', 'T22_10', 'T22_06', 'T25_13', 'T22_01'
#> Warning: Model split preparation for model No. 14, model name
#> domainreading__group1.female_group2.countryA: 132 items from 149 items listed
#> the Q matrix not found in data:
#> ℹ 'T07_08', 'T02_02', 'T07_01', 'T02_06', 'T07_05', 'T07_06', 'T02_07',
#> 'T07_02', 'T07_04', 'T02_03', 'T07_09', 'T02_01', 'T02_04', 'T07_03',
#> 'T02_05', 'T07_07', 'T07_10', 'T09_12', 'T01_08', 'T03_07', 'T03_01',
#> 'T03_08', 'T03_05', 'T03_06', 'T03_03', 'T10_08', 'T11_06', 'T11_03',
#> 'T11_01', 'T04_02', 'T04_01', 'T04_06', 'T04_04', 'T11_04', 'T10_01',
#> 'T11_08', 'T10_02', 'T10_09', 'T10_07', 'T11_02', 'T04_05', 'T04_03',
#> 'T11_05', 'T10_06', 'T11_10', 'T11_09', 'T04_07', 'T10_10', 'T04_09',
#> 'T04_08', 'T11_0X', 'T06_05', 'T05_03', 'T05_06', 'T05_05', 'T06_02',
#> 'T06_04', 'T06_03', 'T05_04', 'T05_02', 'T05_01', 'T06_01', 'T08_04',
#> 'T08_06', 'T08_03', 'T08_07', 'T08_02', 'T08_05', 'T08_01', 'T08_09',
#> 'T08_08', 'T03_09', 'T06_1X', 'T26_03', 'T26_05', 'T26_07', 'T26_09',
#> 'T26_06', 'T26_10', 'T26_08', 'T26_02', 'T26_04', 'T27_01', 'T27_05',
#> 'T27_07', 'T27_06', 'T27_04', 'T27_02', 'T27_03', 'T24_05', 'T24_08',
#> 'T24_02', 'T24_10', 'T24_06', 'T24_24', 'T24_25', 'T24_09', 'T24_21',
#> 'T24_23', 'T23_10', 'T23_13', 'T23_09', 'T23_15', 'T23_03', 'T23_04',
#> 'T23_14', 'T23_07', 'T23_08', 'T23_02', 'T23_06', 'T23_05', 'T25_05',
#> 'T25_22', 'T22_03', 'T25_06', 'T22_12', 'T25_09', 'T25_10', 'T25_08',
#> 'T22_11', 'T22_02', 'T25_11', 'T25_07', 'T25_14', 'T25_12', 'T25_04',
#> 'T22_08', 'T25_23', 'T22_10', 'T22_06', 'T25_13', 'T22_01'
#> Warning: Model split preparation for model No. 15, model name
#> domainreading__group1.female_group2.countryB: 132 items from 149 items listed
#> the Q matrix not found in data:
#> ℹ 'T07_08', 'T02_02', 'T07_01', 'T02_06', 'T07_05', 'T07_06', 'T02_07',
#> 'T07_02', 'T07_04', 'T02_03', 'T07_09', 'T02_01', 'T02_04', 'T07_03',
#> 'T02_05', 'T07_07', 'T07_10', 'T09_12', 'T01_08', 'T03_07', 'T03_01',
#> 'T03_08', 'T03_05', 'T03_06', 'T03_03', 'T10_08', 'T11_06', 'T11_03',
#> 'T11_01', 'T04_02', 'T04_01', 'T04_06', 'T04_04', 'T11_04', 'T10_01',
#> 'T11_08', 'T10_02', 'T10_09', 'T10_07', 'T11_02', 'T04_05', 'T04_03',
#> 'T11_05', 'T10_06', 'T11_10', 'T11_09', 'T04_07', 'T10_10', 'T04_09',
#> 'T04_08', 'T11_0X', 'T06_05', 'T05_03', 'T05_06', 'T05_05', 'T06_02',
#> 'T06_04', 'T06_03', 'T05_04', 'T05_02', 'T05_01', 'T06_01', 'T08_04',
#> 'T08_06', 'T08_03', 'T08_07', 'T08_02', 'T08_05', 'T08_01', 'T08_09',
#> 'T08_08', 'T03_09', 'T06_1X', 'T26_03', 'T26_05', 'T26_07', 'T26_09',
#> 'T26_06', 'T26_10', 'T26_08', 'T26_02', 'T26_04', 'T27_01', 'T27_05',
#> 'T27_07', 'T27_06', 'T27_04', 'T27_02', 'T27_03', 'T24_05', 'T24_08',
#> 'T24_02', 'T24_10', 'T24_06', 'T24_24', 'T24_25', 'T24_09', 'T24_21',
#> 'T24_23', 'T23_10', 'T23_13', 'T23_09', 'T23_15', 'T23_03', 'T23_04',
#> 'T23_14', 'T23_07', 'T23_08', 'T23_02', 'T23_06', 'T23_05', 'T25_05',
#> 'T25_22', 'T22_03', 'T25_06', 'T22_12', 'T25_09', 'T25_10', 'T25_08',
#> 'T22_11', 'T22_02', 'T25_11', 'T25_07', 'T25_14', 'T25_12', 'T25_04',
#> 'T22_08', 'T25_23', 'T22_10', 'T22_06', 'T25_13', 'T22_01'
#> Warning: Model split preparation for model No. 16, model name
#> domainreading__group1.female_group2.all: 132 items from 149 items listed the Q
#> matrix not found in data:
#> ℹ 'T07_08', 'T02_02', 'T07_01', 'T02_06', 'T07_05', 'T07_06', 'T02_07',
#> 'T07_02', 'T07_04', 'T02_03', 'T07_09', 'T02_01', 'T02_04', 'T07_03',
#> 'T02_05', 'T07_07', 'T07_10', 'T09_12', 'T01_08', 'T03_07', 'T03_01',
#> 'T03_08', 'T03_05', 'T03_06', 'T03_03', 'T10_08', 'T11_06', 'T11_03',
#> 'T11_01', 'T04_02', 'T04_01', 'T04_06', 'T04_04', 'T11_04', 'T10_01',
#> 'T11_08', 'T10_02', 'T10_09', 'T10_07', 'T11_02', 'T04_05', 'T04_03',
#> 'T11_05', 'T10_06', 'T11_10', 'T11_09', 'T04_07', 'T10_10', 'T04_09',
#> 'T04_08', 'T11_0X', 'T06_05', 'T05_03', 'T05_06', 'T05_05', 'T06_02',
#> 'T06_04', 'T06_03', 'T05_04', 'T05_02', 'T05_01', 'T06_01', 'T08_04',
#> 'T08_06', 'T08_03', 'T08_07', 'T08_02', 'T08_05', 'T08_01', 'T08_09',
#> 'T08_08', 'T03_09', 'T06_1X', 'T26_03', 'T26_05', 'T26_07', 'T26_09',
#> 'T26_06', 'T26_10', 'T26_08', 'T26_02', 'T26_04', 'T27_01', 'T27_05',
#> 'T27_07', 'T27_06', 'T27_04', 'T27_02', 'T27_03', 'T24_05', 'T24_08',
#> 'T24_02', 'T24_10', 'T24_06', 'T24_24', 'T24_25', 'T24_09', 'T24_21',
#> 'T24_23', 'T23_10', 'T23_13', 'T23_09', 'T23_15', 'T23_03', 'T23_04',
#> 'T23_14', 'T23_07', 'T23_08', 'T23_02', 'T23_06', 'T23_05', 'T25_05',
#> 'T25_22', 'T22_03', 'T25_06', 'T22_12', 'T25_09', 'T25_10', 'T25_08',
#> 'T22_11', 'T22_02', 'T25_11', 'T25_07', 'T25_14', 'T25_12', 'T25_04',
#> 'T22_08', 'T25_23', 'T22_10', 'T22_06', 'T25_13', 'T22_01'
#> Warning: Model split preparation for model No. 17, model name
#> domainreading__group1.male_group2.countryC: 132 items from 149 items listed the
#> Q matrix not found in data:
#> ℹ 'T07_08', 'T02_02', 'T07_01', 'T02_06', 'T07_05', 'T07_06', 'T02_07',
#> 'T07_02', 'T07_04', 'T02_03', 'T07_09', 'T02_01', 'T02_04', 'T07_03',
#> 'T02_05', 'T07_07', 'T07_10', 'T09_12', 'T01_08', 'T03_07', 'T03_01',
#> 'T03_08', 'T03_05', 'T03_06', 'T03_03', 'T10_08', 'T11_06', 'T11_03',
#> 'T11_01', 'T04_02', 'T04_01', 'T04_06', 'T04_04', 'T11_04', 'T10_01',
#> 'T11_08', 'T10_02', 'T10_09', 'T10_07', 'T11_02', 'T04_05', 'T04_03',
#> 'T11_05', 'T10_06', 'T11_10', 'T11_09', 'T04_07', 'T10_10', 'T04_09',
#> 'T04_08', 'T11_0X', 'T06_05', 'T05_03', 'T05_06', 'T05_05', 'T06_02',
#> 'T06_04', 'T06_03', 'T05_04', 'T05_02', 'T05_01', 'T06_01', 'T08_04',
#> 'T08_06', 'T08_03', 'T08_07', 'T08_02', 'T08_05', 'T08_01', 'T08_09',
#> 'T08_08', 'T03_09', 'T06_1X', 'T26_03', 'T26_05', 'T26_07', 'T26_09',
#> 'T26_06', 'T26_10', 'T26_08', 'T26_02', 'T26_04', 'T27_01', 'T27_05',
#> 'T27_07', 'T27_06', 'T27_04', 'T27_02', 'T27_03', 'T24_05', 'T24_08',
#> 'T24_02', 'T24_10', 'T24_06', 'T24_24', 'T24_25', 'T24_09', 'T24_21',
#> 'T24_23', 'T23_10', 'T23_13', 'T23_09', 'T23_15', 'T23_03', 'T23_04',
#> 'T23_14', 'T23_07', 'T23_08', 'T23_02', 'T23_06', 'T23_05', 'T25_05',
#> 'T25_22', 'T22_03', 'T25_06', 'T22_12', 'T25_09', 'T25_10', 'T25_08',
#> 'T22_11', 'T22_02', 'T25_11', 'T25_07', 'T25_14', 'T25_12', 'T25_04',
#> 'T22_08', 'T25_23', 'T22_10', 'T22_06', 'T25_13', 'T22_01'
#> Warning: Model split preparation for model No. 18, model name
#> domainreading__group1.male_group2.countryA: 132 items from 149 items listed the
#> Q matrix not found in data:
#> ℹ 'T07_08', 'T02_02', 'T07_01', 'T02_06', 'T07_05', 'T07_06', 'T02_07',
#> 'T07_02', 'T07_04', 'T02_03', 'T07_09', 'T02_01', 'T02_04', 'T07_03',
#> 'T02_05', 'T07_07', 'T07_10', 'T09_12', 'T01_08', 'T03_07', 'T03_01',
#> 'T03_08', 'T03_05', 'T03_06', 'T03_03', 'T10_08', 'T11_06', 'T11_03',
#> 'T11_01', 'T04_02', 'T04_01', 'T04_06', 'T04_04', 'T11_04', 'T10_01',
#> 'T11_08', 'T10_02', 'T10_09', 'T10_07', 'T11_02', 'T04_05', 'T04_03',
#> 'T11_05', 'T10_06', 'T11_10', 'T11_09', 'T04_07', 'T10_10', 'T04_09',
#> 'T04_08', 'T11_0X', 'T06_05', 'T05_03', 'T05_06', 'T05_05', 'T06_02',
#> 'T06_04', 'T06_03', 'T05_04', 'T05_02', 'T05_01', 'T06_01', 'T08_04',
#> 'T08_06', 'T08_03', 'T08_07', 'T08_02', 'T08_05', 'T08_01', 'T08_09',
#> 'T08_08', 'T03_09', 'T06_1X', 'T26_03', 'T26_05', 'T26_07', 'T26_09',
#> 'T26_06', 'T26_10', 'T26_08', 'T26_02', 'T26_04', 'T27_01', 'T27_05',
#> 'T27_07', 'T27_06', 'T27_04', 'T27_02', 'T27_03', 'T24_05', 'T24_08',
#> 'T24_02', 'T24_10', 'T24_06', 'T24_24', 'T24_25', 'T24_09', 'T24_21',
#> 'T24_23', 'T23_10', 'T23_13', 'T23_09', 'T23_15', 'T23_03', 'T23_04',
#> 'T23_14', 'T23_07', 'T23_08', 'T23_02', 'T23_06', 'T23_05', 'T25_05',
#> 'T25_22', 'T22_03', 'T25_06', 'T22_12', 'T25_09', 'T25_10', 'T25_08',
#> 'T22_11', 'T22_02', 'T25_11', 'T25_07', 'T25_14', 'T25_12', 'T25_04',
#> 'T22_08', 'T25_23', 'T22_10', 'T22_06', 'T25_13', 'T22_01'
#> Warning: Model split preparation for model No. 19, model name
#> domainreading__group1.male_group2.countryB: 132 items from 149 items listed the
#> Q matrix not found in data:
#> ℹ 'T07_08', 'T02_02', 'T07_01', 'T02_06', 'T07_05', 'T07_06', 'T02_07',
#> 'T07_02', 'T07_04', 'T02_03', 'T07_09', 'T02_01', 'T02_04', 'T07_03',
#> 'T02_05', 'T07_07', 'T07_10', 'T09_12', 'T01_08', 'T03_07', 'T03_01',
#> 'T03_08', 'T03_05', 'T03_06', 'T03_03', 'T10_08', 'T11_06', 'T11_03',
#> 'T11_01', 'T04_02', 'T04_01', 'T04_06', 'T04_04', 'T11_04', 'T10_01',
#> 'T11_08', 'T10_02', 'T10_09', 'T10_07', 'T11_02', 'T04_05', 'T04_03',
#> 'T11_05', 'T10_06', 'T11_10', 'T11_09', 'T04_07', 'T10_10', 'T04_09',
#> 'T04_08', 'T11_0X', 'T06_05', 'T05_03', 'T05_06', 'T05_05', 'T06_02',
#> 'T06_04', 'T06_03', 'T05_04', 'T05_02', 'T05_01', 'T06_01', 'T08_04',
#> 'T08_06', 'T08_03', 'T08_07', 'T08_02', 'T08_05', 'T08_01', 'T08_09',
#> 'T08_08', 'T03_09', 'T06_1X', 'T26_03', 'T26_05', 'T26_07', 'T26_09',
#> 'T26_06', 'T26_10', 'T26_08', 'T26_02', 'T26_04', 'T27_01', 'T27_05',
#> 'T27_07', 'T27_06', 'T27_04', 'T27_02', 'T27_03', 'T24_05', 'T24_08',
#> 'T24_02', 'T24_10', 'T24_06', 'T24_24', 'T24_25', 'T24_09', 'T24_21',
#> 'T24_23', 'T23_10', 'T23_13', 'T23_09', 'T23_15', 'T23_03', 'T23_04',
#> 'T23_14', 'T23_07', 'T23_08', 'T23_02', 'T23_06', 'T23_05', 'T25_05',
#> 'T25_22', 'T22_03', 'T25_06', 'T22_12', 'T25_09', 'T25_10', 'T25_08',
#> 'T22_11', 'T22_02', 'T25_11', 'T25_07', 'T25_14', 'T25_12', 'T25_04',
#> 'T22_08', 'T25_23', 'T22_10', 'T22_06', 'T25_13', 'T22_01'
#> Warning: Model split preparation for model No. 20, model name
#> domainreading__group1.male_group2.all: 132 items from 149 items listed the Q
#> matrix not found in data:
#> ℹ 'T07_08', 'T02_02', 'T07_01', 'T02_06', 'T07_05', 'T07_06', 'T02_07',
#> 'T07_02', 'T07_04', 'T02_03', 'T07_09', 'T02_01', 'T02_04', 'T07_03',
#> 'T02_05', 'T07_07', 'T07_10', 'T09_12', 'T01_08', 'T03_07', 'T03_01',
#> 'T03_08', 'T03_05', 'T03_06', 'T03_03', 'T10_08', 'T11_06', 'T11_03',
#> 'T11_01', 'T04_02', 'T04_01', 'T04_06', 'T04_04', 'T11_04', 'T10_01',
#> 'T11_08', 'T10_02', 'T10_09', 'T10_07', 'T11_02', 'T04_05', 'T04_03',
#> 'T11_05', 'T10_06', 'T11_10', 'T11_09', 'T04_07', 'T10_10', 'T04_09',
#> 'T04_08', 'T11_0X', 'T06_05', 'T05_03', 'T05_06', 'T05_05', 'T06_02',
#> 'T06_04', 'T06_03', 'T05_04', 'T05_02', 'T05_01', 'T06_01', 'T08_04',
#> 'T08_06', 'T08_03', 'T08_07', 'T08_02', 'T08_05', 'T08_01', 'T08_09',
#> 'T08_08', 'T03_09', 'T06_1X', 'T26_03', 'T26_05', 'T26_07', 'T26_09',
#> 'T26_06', 'T26_10', 'T26_08', 'T26_02', 'T26_04', 'T27_01', 'T27_05',
#> 'T27_07', 'T27_06', 'T27_04', 'T27_02', 'T27_03', 'T24_05', 'T24_08',
#> 'T24_02', 'T24_10', 'T24_06', 'T24_24', 'T24_25', 'T24_09', 'T24_21',
#> 'T24_23', 'T23_10', 'T23_13', 'T23_09', 'T23_15', 'T23_03', 'T23_04',
#> 'T23_14', 'T23_07', 'T23_08', 'T23_02', 'T23_06', 'T23_05', 'T25_05',
#> 'T25_22', 'T22_03', 'T25_06', 'T22_12', 'T25_09', 'T25_10', 'T25_08',
#> 'T22_11', 'T22_02', 'T25_11', 'T25_07', 'T25_14', 'T25_12', 'T25_04',
#> 'T22_08', 'T25_23', 'T22_10', 'T22_06', 'T25_13', 'T22_01'
#> Warning: Model split preparation for model No. 21, model name
#> domainreading__group1.all_group2.countryC: 132 items from 149 items listed the
#> Q matrix not found in data:
#> ℹ 'T07_08', 'T02_02', 'T07_01', 'T02_06', 'T07_05', 'T07_06', 'T02_07',
#> 'T07_02', 'T07_04', 'T02_03', 'T07_09', 'T02_01', 'T02_04', 'T07_03',
#> 'T02_05', 'T07_07', 'T07_10', 'T09_12', 'T01_08', 'T03_07', 'T03_01',
#> 'T03_08', 'T03_05', 'T03_06', 'T03_03', 'T10_08', 'T11_06', 'T11_03',
#> 'T11_01', 'T04_02', 'T04_01', 'T04_06', 'T04_04', 'T11_04', 'T10_01',
#> 'T11_08', 'T10_02', 'T10_09', 'T10_07', 'T11_02', 'T04_05', 'T04_03',
#> 'T11_05', 'T10_06', 'T11_10', 'T11_09', 'T04_07', 'T10_10', 'T04_09',
#> 'T04_08', 'T11_0X', 'T06_05', 'T05_03', 'T05_06', 'T05_05', 'T06_02',
#> 'T06_04', 'T06_03', 'T05_04', 'T05_02', 'T05_01', 'T06_01', 'T08_04',
#> 'T08_06', 'T08_03', 'T08_07', 'T08_02', 'T08_05', 'T08_01', 'T08_09',
#> 'T08_08', 'T03_09', 'T06_1X', 'T26_03', 'T26_05', 'T26_07', 'T26_09',
#> 'T26_06', 'T26_10', 'T26_08', 'T26_02', 'T26_04', 'T27_01', 'T27_05',
#> 'T27_07', 'T27_06', 'T27_04', 'T27_02', 'T27_03', 'T24_05', 'T24_08',
#> 'T24_02', 'T24_10', 'T24_06', 'T24_24', 'T24_25', 'T24_09', 'T24_21',
#> 'T24_23', 'T23_10', 'T23_13', 'T23_09', 'T23_15', 'T23_03', 'T23_04',
#> 'T23_14', 'T23_07', 'T23_08', 'T23_02', 'T23_06', 'T23_05', 'T25_05',
#> 'T25_22', 'T22_03', 'T25_06', 'T22_12', 'T25_09', 'T25_10', 'T25_08',
#> 'T22_11', 'T22_02', 'T25_11', 'T25_07', 'T25_14', 'T25_12', 'T25_04',
#> 'T22_08', 'T25_23', 'T22_10', 'T22_06', 'T25_13', 'T22_01'
#> Warning: Model split preparation for model No. 22, model name
#> domainreading__group1.all_group2.countryA: 132 items from 149 items listed the
#> Q matrix not found in data:
#> ℹ 'T07_08', 'T02_02', 'T07_01', 'T02_06', 'T07_05', 'T07_06', 'T02_07',
#> 'T07_02', 'T07_04', 'T02_03', 'T07_09', 'T02_01', 'T02_04', 'T07_03',
#> 'T02_05', 'T07_07', 'T07_10', 'T09_12', 'T01_08', 'T03_07', 'T03_01',
#> 'T03_08', 'T03_05', 'T03_06', 'T03_03', 'T10_08', 'T11_06', 'T11_03',
#> 'T11_01', 'T04_02', 'T04_01', 'T04_06', 'T04_04', 'T11_04', 'T10_01',
#> 'T11_08', 'T10_02', 'T10_09', 'T10_07', 'T11_02', 'T04_05', 'T04_03',
#> 'T11_05', 'T10_06', 'T11_10', 'T11_09', 'T04_07', 'T10_10', 'T04_09',
#> 'T04_08', 'T11_0X', 'T06_05', 'T05_03', 'T05_06', 'T05_05', 'T06_02',
#> 'T06_04', 'T06_03', 'T05_04', 'T05_02', 'T05_01', 'T06_01', 'T08_04',
#> 'T08_06', 'T08_03', 'T08_07', 'T08_02', 'T08_05', 'T08_01', 'T08_09',
#> 'T08_08', 'T03_09', 'T06_1X', 'T26_03', 'T26_05', 'T26_07', 'T26_09',
#> 'T26_06', 'T26_10', 'T26_08', 'T26_02', 'T26_04', 'T27_01', 'T27_05',
#> 'T27_07', 'T27_06', 'T27_04', 'T27_02', 'T27_03', 'T24_05', 'T24_08',
#> 'T24_02', 'T24_10', 'T24_06', 'T24_24', 'T24_25', 'T24_09', 'T24_21',
#> 'T24_23', 'T23_10', 'T23_13', 'T23_09', 'T23_15', 'T23_03', 'T23_04',
#> 'T23_14', 'T23_07', 'T23_08', 'T23_02', 'T23_06', 'T23_05', 'T25_05',
#> 'T25_22', 'T22_03', 'T25_06', 'T22_12', 'T25_09', 'T25_10', 'T25_08',
#> 'T22_11', 'T22_02', 'T25_11', 'T25_07', 'T25_14', 'T25_12', 'T25_04',
#> 'T22_08', 'T25_23', 'T22_10', 'T22_06', 'T25_13', 'T22_01'
#> Warning: Model split preparation for model No. 23, model name
#> domainreading__group1.all_group2.countryB: 132 items from 149 items listed the
#> Q matrix not found in data:
#> ℹ 'T07_08', 'T02_02', 'T07_01', 'T02_06', 'T07_05', 'T07_06', 'T02_07',
#> 'T07_02', 'T07_04', 'T02_03', 'T07_09', 'T02_01', 'T02_04', 'T07_03',
#> 'T02_05', 'T07_07', 'T07_10', 'T09_12', 'T01_08', 'T03_07', 'T03_01',
#> 'T03_08', 'T03_05', 'T03_06', 'T03_03', 'T10_08', 'T11_06', 'T11_03',
#> 'T11_01', 'T04_02', 'T04_01', 'T04_06', 'T04_04', 'T11_04', 'T10_01',
#> 'T11_08', 'T10_02', 'T10_09', 'T10_07', 'T11_02', 'T04_05', 'T04_03',
#> 'T11_05', 'T10_06', 'T11_10', 'T11_09', 'T04_07', 'T10_10', 'T04_09',
#> 'T04_08', 'T11_0X', 'T06_05', 'T05_03', 'T05_06', 'T05_05', 'T06_02',
#> 'T06_04', 'T06_03', 'T05_04', 'T05_02', 'T05_01', 'T06_01', 'T08_04',
#> 'T08_06', 'T08_03', 'T08_07', 'T08_02', 'T08_05', 'T08_01', 'T08_09',
#> 'T08_08', 'T03_09', 'T06_1X', 'T26_03', 'T26_05', 'T26_07', 'T26_09',
#> 'T26_06', 'T26_10', 'T26_08', 'T26_02', 'T26_04', 'T27_01', 'T27_05',
#> 'T27_07', 'T27_06', 'T27_04', 'T27_02', 'T27_03', 'T24_05', 'T24_08',
#> 'T24_02', 'T24_10', 'T24_06', 'T24_24', 'T24_25', 'T24_09', 'T24_21',
#> 'T24_23', 'T23_10', 'T23_13', 'T23_09', 'T23_15', 'T23_03', 'T23_04',
#> 'T23_14', 'T23_07', 'T23_08', 'T23_02', 'T23_06', 'T23_05', 'T25_05',
#> 'T25_22', 'T22_03', 'T25_06', 'T22_12', 'T25_09', 'T25_10', 'T25_08',
#> 'T22_11', 'T22_02', 'T25_11', 'T25_07', 'T25_14', 'T25_12', 'T25_04',
#> 'T22_08', 'T25_23', 'T22_10', 'T22_06', 'T25_13', 'T22_01'
#> Warning: Model split preparation for model No. 24, model name
#> domainreading__group1.all_group2.all: 132 items from 149 items listed the Q
#> matrix not found in data:
#> ℹ 'T07_08', 'T02_02', 'T07_01', 'T02_06', 'T07_05', 'T07_06', 'T02_07',
#> 'T07_02', 'T07_04', 'T02_03', 'T07_09', 'T02_01', 'T02_04', 'T07_03',
#> 'T02_05', 'T07_07', 'T07_10', 'T09_12', 'T01_08', 'T03_07', 'T03_01',
#> 'T03_08', 'T03_05', 'T03_06', 'T03_03', 'T10_08', 'T11_06', 'T11_03',
#> 'T11_01', 'T04_02', 'T04_01', 'T04_06', 'T04_04', 'T11_04', 'T10_01',
#> 'T11_08', 'T10_02', 'T10_09', 'T10_07', 'T11_02', 'T04_05', 'T04_03',
#> 'T11_05', 'T10_06', 'T11_10', 'T11_09', 'T04_07', 'T10_10', 'T04_09',
#> 'T04_08', 'T11_0X', 'T06_05', 'T05_03', 'T05_06', 'T05_05', 'T06_02',
#> 'T06_04', 'T06_03', 'T05_04', 'T05_02', 'T05_01', 'T06_01', 'T08_04',
#> 'T08_06', 'T08_03', 'T08_07', 'T08_02', 'T08_05', 'T08_01', 'T08_09',
#> 'T08_08', 'T03_09', 'T06_1X', 'T26_03', 'T26_05', 'T26_07', 'T26_09',
#> 'T26_06', 'T26_10', 'T26_08', 'T26_02', 'T26_04', 'T27_01', 'T27_05',
#> 'T27_07', 'T27_06', 'T27_04', 'T27_02', 'T27_03', 'T24_05', 'T24_08',
#> 'T24_02', 'T24_10', 'T24_06', 'T24_24', 'T24_25', 'T24_09', 'T24_21',
#> 'T24_23', 'T23_10', 'T23_13', 'T23_09', 'T23_15', 'T23_03', 'T23_04',
#> 'T23_14', 'T23_07', 'T23_08', 'T23_02', 'T23_06', 'T23_05', 'T25_05',
#> 'T25_22', 'T22_03', 'T25_06', 'T22_12', 'T25_09', 'T25_10', 'T25_08',
#> 'T22_11', 'T22_02', 'T25_11', 'T25_07', 'T25_14', 'T25_12', 'T25_04',
#> 'T22_08', 'T25_23', 'T22_10', 'T22_06', 'T25_13', 'T22_01'
#>
#>
#> ========================================================================
#> Model No. 1
#> Model name: domainlistening__group1.female_group2.countryC
#> Number of items: 16
#> Number of persons: 20
#> Number of dimensions: 1
#> ========================================================================
#>
#> Following 118 item(s) missed in data frame will be removed from Q matrix:
#> T12_11, T13_15, T15_03, T13_04, T13_07, T13_16, T13_01, T15_02, T15_01, T15_15, T15_16, T15_09, T13_12, T13_13, T15_14, T13_08, T13_06, T15_12, T13_10, T13_17, T15_11, T13_05, T13_09, T15_10, T15_13, T15_17, T15_18, T13_18, T16_08, T16_09, T16_11, T16_13, T16_01, T16_04, T16_10, T16_02, T16_12, T16_06, T16_07, T19_06, T19_08, T19_05, T19_11, T19_01, T19_02, T19_30, T19_03, T19_07, T19_04, T19_09, T21_11, T18_01, T21_01, T18_05, T21_12, T18_13, T21_10, T18_11, T21_05, T21_13, T18_08, T18_07, T21_09, T18_04, T18_02, T21_02, T21_06, T18_09, T21_04, T18_12, T21_14, T21_03, T18_10, T30_06, T29_04, T29_02, T30_02, T30_03, T30_01, T29_05, T29_03, T28_08, T28_09, T28_02, T28_03, T28_04, T28_01, T28_10, T28_06, T20_05, T20_09, T20_30, T20_08, T20_12, T20_06, T20_07, T20_02, T20_13, T20_03, T20_01, T20_04, T20_11, T20_10, T32_04, T31_03, T31_01, T31_04, T32_03, T32_02, T32_01, T17_02, T17_09, T17_10, T17_04, T17_03, T17_08, T17_06, T17_01
#> Warning: 16 testitem(s) with less than 50 valid responses: These items are nevertheless kept in the data set: 'T12_06', 'T12_07', 'T14_03', 'T14_06', 'T12_01', 'T12_03', 'T12_08', 'T12_10', 'T14_02', 'T14_05', 'T12_02', 'T12_04', 'T14_04', 'T12_05', 'T14_01', 'T12_09'
#> Warning: 2 testitem(s) are constants. Remove these items from the data set:
#> Item 'T14_04', only value '0' occurs: 20 valid responses.
#> Item 'T12_05', only value '1' occurs: 20 valid responses.
#> Remove 2 test item(s) overall.
#> Dataset is completely linked.
#> 'gauss' has been chosen for estimation method. Number of nodes was not explicitly specified. Set nodes to 20.
#> Following 14 items with less than 50 item responses:
#> item.name cases Missing valid item.p
#> factor.T12_06.1.1 T12_06 20 0 20 0.50
#> factor.T12_07.1.1 T12_07 20 0 20 0.75
#> factor.T14_03.1.1 T14_03 20 0 20 0.10
#> factor.T14_06.1.1 T14_06 20 0 20 0.35
#> factor.T12_01.1.1 T12_01 20 0 20 0.90
#> factor.T12_03.1.1 T12_03 20 0 20 0.90
#> factor.T12_08.1.1 T12_08 20 0 20 0.80
#> factor.T12_10.1.1 T12_10 20 0 20 0.70
#> factor.T14_02.1.1 T14_02 20 0 20 0.15
#> factor.T14_05.1.1 T14_05 20 0 20 0.30
#> factor.T12_02.1.1 T12_02 20 0 20 0.50
#> factor.T12_04.1.1 T12_04 20 0 20 0.70
#> factor.T14_01.1.1 T14_01 20 0 20 0.15
#> factor.T12_09.1.1 T12_09 20 0 20 0.95
#> Q matrix specifies 1 dimension(s).
#>
#>
#> ========================================================================
#> Model No. 2
#> Model name: domainlistening__group1.female_group2.countryA
#> Number of items: 16
#> Number of persons: 42
#> Number of dimensions: 1
#> ========================================================================
#>
#> Following 118 item(s) missed in data frame will be removed from Q matrix:
#> T12_11, T13_15, T15_03, T13_04, T13_07, T13_16, T13_01, T15_02, T15_01, T15_15, T15_16, T15_09, T13_12, T13_13, T15_14, T13_08, T13_06, T15_12, T13_10, T13_17, T15_11, T13_05, T13_09, T15_10, T15_13, T15_17, T15_18, T13_18, T16_08, T16_09, T16_11, T16_13, T16_01, T16_04, T16_10, T16_02, T16_12, T16_06, T16_07, T19_06, T19_08, T19_05, T19_11, T19_01, T19_02, T19_30, T19_03, T19_07, T19_04, T19_09, T21_11, T18_01, T21_01, T18_05, T21_12, T18_13, T21_10, T18_11, T21_05, T21_13, T18_08, T18_07, T21_09, T18_04, T18_02, T21_02, T21_06, T18_09, T21_04, T18_12, T21_14, T21_03, T18_10, T30_06, T29_04, T29_02, T30_02, T30_03, T30_01, T29_05, T29_03, T28_08, T28_09, T28_02, T28_03, T28_04, T28_01, T28_10, T28_06, T20_05, T20_09, T20_30, T20_08, T20_12, T20_06, T20_07, T20_02, T20_13, T20_03, T20_01, T20_04, T20_11, T20_10, T32_04, T31_03, T31_01, T31_04, T32_03, T32_02, T32_01, T17_02, T17_09, T17_10, T17_04, T17_03, T17_08, T17_06, T17_01
#> Warning: 16 testitem(s) with less than 50 valid responses: These items are nevertheless kept in the data set: 'T12_06', 'T12_07', 'T14_03', 'T14_06', 'T12_01', 'T12_03', 'T12_08', 'T12_10', 'T14_02', 'T14_05', 'T12_02', 'T12_04', 'T14_04', 'T12_05', 'T14_01', 'T12_09'
#> Warning: 1 testitem(s) are constants. Remove these items from the data set:
#> Item 'T12_05', only value '1' occurs: 42 valid responses.
#> Remove 1 test item(s) overall.
#> 1 subject(s) do not solve any item:
#> P02289 (15 false), P02289 (15 false) ...
#> Dataset is completely linked.
#> 'gauss' has been chosen for estimation method. Number of nodes was not explicitly specified. Set nodes to 20.
#> Following 15 items with less than 50 item responses:
#> item.name cases Missing valid item.p
#> factor.T12_06.1.1 T12_06 42 0 42 0.4762
#> factor.T12_07.1.1 T12_07 42 0 42 0.7381
#> factor.T14_03.1.1 T14_03 42 0 42 0.1190
#> factor.T14_06.1.1 T14_06 42 0 42 0.3810
#> factor.T12_01.1.1 T12_01 42 0 42 0.9762
#> factor.T12_03.1.1 T12_03 42 0 42 0.9048
#> factor.T12_08.1.1 T12_08 42 0 42 0.6190
#> factor.T12_10.1.1 T12_10 42 0 42 0.7857
#> factor.T14_02.1.1 T14_02 42 0 42 0.1667
#> factor.T14_05.1.1 T14_05 42 0 42 0.3095
#> factor.T12_02.1.1 T12_02 42 0 42 0.5714
#> factor.T12_04.1.1 T12_04 42 0 42 0.7619
#> factor.T14_04.1.1 T14_04 42 0 42 0.0476
#> factor.T14_01.1.1 T14_01 42 0 42 0.0714
#> factor.T12_09.1.1 T12_09 42 0 42 0.9048
#> Q matrix specifies 1 dimension(s).
#>
#>
#> ========================================================================
#> Model No. 3
#> Model name: domainlistening__group1.female_group2.countryB
#> Number of items: 16
#> Number of persons: 27
#> Number of dimensions: 1
#> ========================================================================
#>
#> Following 118 item(s) missed in data frame will be removed from Q matrix:
#> T12_11, T13_15, T15_03, T13_04, T13_07, T13_16, T13_01, T15_02, T15_01, T15_15, T15_16, T15_09, T13_12, T13_13, T15_14, T13_08, T13_06, T15_12, T13_10, T13_17, T15_11, T13_05, T13_09, T15_10, T15_13, T15_17, T15_18, T13_18, T16_08, T16_09, T16_11, T16_13, T16_01, T16_04, T16_10, T16_02, T16_12, T16_06, T16_07, T19_06, T19_08, T19_05, T19_11, T19_01, T19_02, T19_30, T19_03, T19_07, T19_04, T19_09, T21_11, T18_01, T21_01, T18_05, T21_12, T18_13, T21_10, T18_11, T21_05, T21_13, T18_08, T18_07, T21_09, T18_04, T18_02, T21_02, T21_06, T18_09, T21_04, T18_12, T21_14, T21_03, T18_10, T30_06, T29_04, T29_02, T30_02, T30_03, T30_01, T29_05, T29_03, T28_08, T28_09, T28_02, T28_03, T28_04, T28_01, T28_10, T28_06, T20_05, T20_09, T20_30, T20_08, T20_12, T20_06, T20_07, T20_02, T20_13, T20_03, T20_01, T20_04, T20_11, T20_10, T32_04, T31_03, T31_01, T31_04, T32_03, T32_02, T32_01, T17_02, T17_09, T17_10, T17_04, T17_03, T17_08, T17_06, T17_01
#> Warning: 16 testitem(s) with less than 50 valid responses: These items are nevertheless kept in the data set: 'T12_06', 'T12_07', 'T14_03', 'T14_06', 'T12_01', 'T12_03', 'T12_08', 'T12_10', 'T14_02', 'T14_05', 'T12_02', 'T12_04', 'T14_04', 'T12_05', 'T14_01', 'T12_09'
#> Dataset is completely linked.
#> 'gauss' has been chosen for estimation method. Number of nodes was not explicitly specified. Set nodes to 20.
#> Following 16 items with less than 50 item responses:
#> item.name cases Missing valid item.p
#> factor.T12_06.1.1 T12_06 27 0 27 0.222
#> factor.T12_07.1.1 T12_07 27 0 27 0.852
#> factor.T14_03.1.1 T14_03 27 0 27 0.148
#> factor.T14_06.1.1 T14_06 27 0 27 0.444
#> factor.T12_01.1.1 T12_01 27 0 27 0.963
#> factor.T12_03.1.1 T12_03 27 0 27 0.815
#> factor.T12_08.1.1 T12_08 27 0 27 0.926
#> factor.T12_10.1.1 T12_10 27 0 27 0.852
#> factor.T14_02.1.1 T14_02 27 0 27 0.222
#> factor.T14_05.1.1 T14_05 27 0 27 0.296
#> factor.T12_02.1.1 T12_02 27 0 27 0.519
#> factor.T12_04.1.1 T12_04 27 0 27 0.815
#> factor.T14_04.1.1 T14_04 27 0 27 0.111
#> factor.T12_05.1.1 T12_05 27 0 27 0.963
#> factor.T14_01.1.1 T14_01 27 0 27 0.111
#> factor.T12_09.1.1 T12_09 27 0 27 0.926
#> Q matrix specifies 1 dimension(s).
#>
#>
#> ===================================================================
#> Model No. 4
#> Model name: domainlistening__group1.female_group2.all
#> Number of items: 16
#> Number of persons: 89
#> Number of dimensions: 1
#> ===================================================================
#>
#> Following 118 item(s) missed in data frame will be removed from Q matrix:
#> T12_11, T13_15, T15_03, T13_04, T13_07, T13_16, T13_01, T15_02, T15_01, T15_15, T15_16, T15_09, T13_12, T13_13, T15_14, T13_08, T13_06, T15_12, T13_10, T13_17, T15_11, T13_05, T13_09, T15_10, T15_13, T15_17, T15_18, T13_18, T16_08, T16_09, T16_11, T16_13, T16_01, T16_04, T16_10, T16_02, T16_12, T16_06, T16_07, T19_06, T19_08, T19_05, T19_11, T19_01, T19_02, T19_30, T19_03, T19_07, T19_04, T19_09, T21_11, T18_01, T21_01, T18_05, T21_12, T18_13, T21_10, T18_11, T21_05, T21_13, T18_08, T18_07, T21_09, T18_04, T18_02, T21_02, T21_06, T18_09, T21_04, T18_12, T21_14, T21_03, T18_10, T30_06, T29_04, T29_02, T30_02, T30_03, T30_01, T29_05, T29_03, T28_08, T28_09, T28_02, T28_03, T28_04, T28_01, T28_10, T28_06, T20_05, T20_09, T20_30, T20_08, T20_12, T20_06, T20_07, T20_02, T20_13, T20_03, T20_01, T20_04, T20_11, T20_10, T32_04, T31_03, T31_01, T31_04, T32_03, T32_02, T32_01, T17_02, T17_09, T17_10, T17_04, T17_03, T17_08, T17_06, T17_01
#> Dataset is completely linked.
#> 'gauss' has been chosen for estimation method. Number of nodes was not explicitly specified. Set nodes to 20.
#> Q matrix specifies 1 dimension(s).
#>
#>
#> ======================================================================
#> Model No. 5
#> Model name: domainlistening__group1.male_group2.countryC
#> Number of items: 16
#> Number of persons: 16
#> Number of dimensions: 1
#> ======================================================================
#>
#> Following 118 item(s) missed in data frame will be removed from Q matrix:
#> T12_11, T13_15, T15_03, T13_04, T13_07, T13_16, T13_01, T15_02, T15_01, T15_15, T15_16, T15_09, T13_12, T13_13, T15_14, T13_08, T13_06, T15_12, T13_10, T13_17, T15_11, T13_05, T13_09, T15_10, T15_13, T15_17, T15_18, T13_18, T16_08, T16_09, T16_11, T16_13, T16_01, T16_04, T16_10, T16_02, T16_12, T16_06, T16_07, T19_06, T19_08, T19_05, T19_11, T19_01, T19_02, T19_30, T19_03, T19_07, T19_04, T19_09, T21_11, T18_01, T21_01, T18_05, T21_12, T18_13, T21_10, T18_11, T21_05, T21_13, T18_08, T18_07, T21_09, T18_04, T18_02, T21_02, T21_06, T18_09, T21_04, T18_12, T21_14, T21_03, T18_10, T30_06, T29_04, T29_02, T30_02, T30_03, T30_01, T29_05, T29_03, T28_08, T28_09, T28_02, T28_03, T28_04, T28_01, T28_10, T28_06, T20_05, T20_09, T20_30, T20_08, T20_12, T20_06, T20_07, T20_02, T20_13, T20_03, T20_01, T20_04, T20_11, T20_10, T32_04, T31_03, T31_01, T31_04, T32_03, T32_02, T32_01, T17_02, T17_09, T17_10, T17_04, T17_03, T17_08, T17_06, T17_01
#> Warning: 16 testitem(s) with less than 50 valid responses: These items are nevertheless kept in the data set: 'T12_06', 'T12_07', 'T14_03', 'T14_06', 'T12_01', 'T12_03', 'T12_08', 'T12_10', 'T14_02', 'T14_05', 'T12_02', 'T12_04', 'T14_04', 'T12_05', 'T14_01', 'T12_09'
#> Warning: 2 testitem(s) are constants. Remove these items from the data set:
#> Item 'T14_04', only value '0' occurs: 16 valid responses.
#> Item 'T12_05', only value '1' occurs: 16 valid responses.
#> Remove 2 test item(s) overall.
#> Dataset is completely linked.
#> 'gauss' has been chosen for estimation method. Number of nodes was not explicitly specified. Set nodes to 20.
#> Following 14 items with less than 50 item responses:
#> item.name cases Missing valid item.p
#> factor.T12_06.1.1 T12_06 16 0 16 0.6875
#> factor.T12_07.1.1 T12_07 16 0 16 0.7500
#> factor.T14_03.1.1 T14_03 16 0 16 0.0625
#> factor.T14_06.1.1 T14_06 16 0 16 0.2500
#> factor.T12_01.1.1 T12_01 16 0 16 0.6875
#> factor.T12_03.1.1 T12_03 16 0 16 0.8750
#> factor.T12_08.1.1 T12_08 16 0 16 0.4375
#> factor.T12_10.1.1 T12_10 16 0 16 0.9375
#> factor.T14_02.1.1 T14_02 16 0 16 0.1250
#> factor.T14_05.1.1 T14_05 16 0 16 0.1250
#> factor.T12_02.1.1 T12_02 16 0 16 0.6875
#> factor.T12_04.1.1 T12_04 16 0 16 0.8750
#> factor.T14_01.1.1 T14_01 16 0 16 0.0625
#> factor.T12_09.1.1 T12_09 16 0 16 0.9375
#> Q matrix specifies 1 dimension(s).
#>
#>
#> ======================================================================
#> Model No. 6
#> Model name: domainlistening__group1.male_group2.countryA
#> Number of items: 16
#> Number of persons: 32
#> Number of dimensions: 1
#> ======================================================================
#>
#> Following 118 item(s) missed in data frame will be removed from Q matrix:
#> T12_11, T13_15, T15_03, T13_04, T13_07, T13_16, T13_01, T15_02, T15_01, T15_15, T15_16, T15_09, T13_12, T13_13, T15_14, T13_08, T13_06, T15_12, T13_10, T13_17, T15_11, T13_05, T13_09, T15_10, T15_13, T15_17, T15_18, T13_18, T16_08, T16_09, T16_11, T16_13, T16_01, T16_04, T16_10, T16_02, T16_12, T16_06, T16_07, T19_06, T19_08, T19_05, T19_11, T19_01, T19_02, T19_30, T19_03, T19_07, T19_04, T19_09, T21_11, T18_01, T21_01, T18_05, T21_12, T18_13, T21_10, T18_11, T21_05, T21_13, T18_08, T18_07, T21_09, T18_04, T18_02, T21_02, T21_06, T18_09, T21_04, T18_12, T21_14, T21_03, T18_10, T30_06, T29_04, T29_02, T30_02, T30_03, T30_01, T29_05, T29_03, T28_08, T28_09, T28_02, T28_03, T28_04, T28_01, T28_10, T28_06, T20_05, T20_09, T20_30, T20_08, T20_12, T20_06, T20_07, T20_02, T20_13, T20_03, T20_01, T20_04, T20_11, T20_10, T32_04, T31_03, T31_01, T31_04, T32_03, T32_02, T32_01, T17_02, T17_09, T17_10, T17_04, T17_03, T17_08, T17_06, T17_01
#> Warning: 16 testitem(s) with less than 50 valid responses: These items are nevertheless kept in the data set: 'T12_06', 'T12_07', 'T14_03', 'T14_06', 'T12_01', 'T12_03', 'T12_08', 'T12_10', 'T14_02', 'T14_05', 'T12_02', 'T12_04', 'T14_04', 'T12_05', 'T14_01', 'T12_09'
#> Warning: 2 testitem(s) are constants. Remove these items from the data set:
#> Item 'T12_03', only value '1' occurs: 32 valid responses.
#> Item 'T12_05', only value '1' occurs: 32 valid responses.
#> Remove 2 test item(s) overall.
#> Dataset is completely linked.
#> 'gauss' has been chosen for estimation method. Number of nodes was not explicitly specified. Set nodes to 20.
#> Following 14 items with less than 50 item responses:
#> item.name cases Missing valid item.p
#> factor.T12_06.1.1 T12_06 32 0 32 0.5000
#> factor.T12_07.1.1 T12_07 32 0 32 0.8125
#> factor.T14_03.1.1 T14_03 32 0 32 0.0625
#> factor.T14_06.1.1 T14_06 32 0 32 0.2188
#> factor.T12_01.1.1 T12_01 32 0 32 0.9062
#> factor.T12_08.1.1 T12_08 32 0 32 0.8125
#> factor.T12_10.1.1 T12_10 32 0 32 0.8750
#> factor.T14_02.1.1 T14_02 32 0 32 0.0312
#> factor.T14_05.1.1 T14_05 32 0 32 0.1562
#> factor.T12_02.1.1 T12_02 32 0 32 0.6562
#> factor.T12_04.1.1 T12_04 32 0 32 0.8438
#> factor.T14_04.1.1 T14_04 32 0 32 0.1250
#> factor.T14_01.1.1 T14_01 32 0 32 0.0938
#> factor.T12_09.1.1 T12_09 32 0 32 0.9375
#> Q matrix specifies 1 dimension(s).
#>
#>
#> ======================================================================
#> Model No. 7
#> Model name: domainlistening__group1.male_group2.countryB
#> Number of items: 16
#> Number of persons: 35
#> Number of dimensions: 1
#> ======================================================================
#>
#> Following 118 item(s) missed in data frame will be removed from Q matrix:
#> T12_11, T13_15, T15_03, T13_04, T13_07, T13_16, T13_01, T15_02, T15_01, T15_15, T15_16, T15_09, T13_12, T13_13, T15_14, T13_08, T13_06, T15_12, T13_10, T13_17, T15_11, T13_05, T13_09, T15_10, T15_13, T15_17, T15_18, T13_18, T16_08, T16_09, T16_11, T16_13, T16_01, T16_04, T16_10, T16_02, T16_12, T16_06, T16_07, T19_06, T19_08, T19_05, T19_11, T19_01, T19_02, T19_30, T19_03, T19_07, T19_04, T19_09, T21_11, T18_01, T21_01, T18_05, T21_12, T18_13, T21_10, T18_11, T21_05, T21_13, T18_08, T18_07, T21_09, T18_04, T18_02, T21_02, T21_06, T18_09, T21_04, T18_12, T21_14, T21_03, T18_10, T30_06, T29_04, T29_02, T30_02, T30_03, T30_01, T29_05, T29_03, T28_08, T28_09, T28_02, T28_03, T28_04, T28_01, T28_10, T28_06, T20_05, T20_09, T20_30, T20_08, T20_12, T20_06, T20_07, T20_02, T20_13, T20_03, T20_01, T20_04, T20_11, T20_10, T32_04, T31_03, T31_01, T31_04, T32_03, T32_02, T32_01, T17_02, T17_09, T17_10, T17_04, T17_03, T17_08, T17_06, T17_01
#> Warning: 16 testitem(s) with less than 50 valid responses: These items are nevertheless kept in the data set: 'T12_06', 'T12_07', 'T14_03', 'T14_06', 'T12_01', 'T12_03', 'T12_08', 'T12_10', 'T14_02', 'T14_05', 'T12_02', 'T12_04', 'T14_04', 'T12_05', 'T14_01', 'T12_09'
#> Dataset is completely linked.
#> 'gauss' has been chosen for estimation method. Number of nodes was not explicitly specified. Set nodes to 20.
#> Following 16 items with less than 50 item responses:
#> item.name cases Missing valid item.p
#> factor.T12_06.1.1 T12_06 35 0 35 0.4571
#> factor.T12_07.1.1 T12_07 35 0 35 0.7714
#> factor.T14_03.1.1 T14_03 35 0 35 0.1714
#> factor.T14_06.1.1 T14_06 35 0 35 0.4286
#> factor.T12_01.1.1 T12_01 35 0 35 0.9143
#> factor.T12_03.1.1 T12_03 35 0 35 0.9143
#> factor.T12_08.1.1 T12_08 35 0 35 0.8000
#> factor.T12_10.1.1 T12_10 35 0 35 0.9143
#> factor.T14_02.1.1 T14_02 35 0 35 0.1429
#> factor.T14_05.1.1 T14_05 35 0 35 0.2286
#> factor.T12_02.1.1 T12_02 35 0 35 0.5714
#> factor.T12_04.1.1 T12_04 35 0 35 0.8857
#> factor.T14_04.1.1 T14_04 35 0 35 0.0571
#> factor.T12_05.1.1 T12_05 35 0 35 0.9714
#> factor.T14_01.1.1 T14_01 35 0 35 0.1714
#> factor.T12_09.1.1 T12_09 35 0 35 0.8857
#> Q matrix specifies 1 dimension(s).
#>
#>
#> =================================================================
#> Model No. 8
#> Model name: domainlistening__group1.male_group2.all
#> Number of items: 16
#> Number of persons: 83
#> Number of dimensions: 1
#> =================================================================
#>
#> Following 118 item(s) missed in data frame will be removed from Q matrix:
#> T12_11, T13_15, T15_03, T13_04, T13_07, T13_16, T13_01, T15_02, T15_01, T15_15, T15_16, T15_09, T13_12, T13_13, T15_14, T13_08, T13_06, T15_12, T13_10, T13_17, T15_11, T13_05, T13_09, T15_10, T15_13, T15_17, T15_18, T13_18, T16_08, T16_09, T16_11, T16_13, T16_01, T16_04, T16_10, T16_02, T16_12, T16_06, T16_07, T19_06, T19_08, T19_05, T19_11, T19_01, T19_02, T19_30, T19_03, T19_07, T19_04, T19_09, T21_11, T18_01, T21_01, T18_05, T21_12, T18_13, T21_10, T18_11, T21_05, T21_13, T18_08, T18_07, T21_09, T18_04, T18_02, T21_02, T21_06, T18_09, T21_04, T18_12, T21_14, T21_03, T18_10, T30_06, T29_04, T29_02, T30_02, T30_03, T30_01, T29_05, T29_03, T28_08, T28_09, T28_02, T28_03, T28_04, T28_01, T28_10, T28_06, T20_05, T20_09, T20_30, T20_08, T20_12, T20_06, T20_07, T20_02, T20_13, T20_03, T20_01, T20_04, T20_11, T20_10, T32_04, T31_03, T31_01, T31_04, T32_03, T32_02, T32_01, T17_02, T17_09, T17_10, T17_04, T17_03, T17_08, T17_06, T17_01
#> Dataset is completely linked.
#> 'gauss' has been chosen for estimation method. Number of nodes was not explicitly specified. Set nodes to 20.
#> Q matrix specifies 1 dimension(s).
#>
#>
#> =====================================================================
#> Model No. 9
#> Model name: domainlistening__group1.all_group2.countryC
#> Number of items: 16
#> Number of persons: 36
#> Number of dimensions: 1
#> =====================================================================
#>
#> Following 118 item(s) missed in data frame will be removed from Q matrix:
#> T12_11, T13_15, T15_03, T13_04, T13_07, T13_16, T13_01, T15_02, T15_01, T15_15, T15_16, T15_09, T13_12, T13_13, T15_14, T13_08, T13_06, T15_12, T13_10, T13_17, T15_11, T13_05, T13_09, T15_10, T15_13, T15_17, T15_18, T13_18, T16_08, T16_09, T16_11, T16_13, T16_01, T16_04, T16_10, T16_02, T16_12, T16_06, T16_07, T19_06, T19_08, T19_05, T19_11, T19_01, T19_02, T19_30, T19_03, T19_07, T19_04, T19_09, T21_11, T18_01, T21_01, T18_05, T21_12, T18_13, T21_10, T18_11, T21_05, T21_13, T18_08, T18_07, T21_09, T18_04, T18_02, T21_02, T21_06, T18_09, T21_04, T18_12, T21_14, T21_03, T18_10, T30_06, T29_04, T29_02, T30_02, T30_03, T30_01, T29_05, T29_03, T28_08, T28_09, T28_02, T28_03, T28_04, T28_01, T28_10, T28_06, T20_05, T20_09, T20_30, T20_08, T20_12, T20_06, T20_07, T20_02, T20_13, T20_03, T20_01, T20_04, T20_11, T20_10, T32_04, T31_03, T31_01, T31_04, T32_03, T32_02, T32_01, T17_02, T17_09, T17_10, T17_04, T17_03, T17_08, T17_06, T17_01
#> Warning: 16 testitem(s) with less than 50 valid responses: These items are nevertheless kept in the data set: 'T12_06', 'T12_07', 'T14_03', 'T14_06', 'T12_01', 'T12_03', 'T12_08', 'T12_10', 'T14_02', 'T14_05', 'T12_02', 'T12_04', 'T14_04', 'T12_05', 'T14_01', 'T12_09'
#> Warning: 2 testitem(s) are constants. Remove these items from the data set:
#> Item 'T14_04', only value '0' occurs: 36 valid responses.
#> Item 'T12_05', only value '1' occurs: 36 valid responses.
#> Remove 2 test item(s) overall.
#> Dataset is completely linked.
#> 'gauss' has been chosen for estimation method. Number of nodes was not explicitly specified. Set nodes to 20.
#> Following 14 items with less than 50 item responses:
#> item.name cases Missing valid item.p
#> factor.T12_06.1.1 T12_06 36 0 36 0.5833
#> factor.T12_07.1.1 T12_07 36 0 36 0.7500
#> factor.T14_03.1.1 T14_03 36 0 36 0.0833
#> factor.T14_06.1.1 T14_06 36 0 36 0.3056
#> factor.T12_01.1.1 T12_01 36 0 36 0.8056
#> factor.T12_03.1.1 T12_03 36 0 36 0.8889
#> factor.T12_08.1.1 T12_08 36 0 36 0.6389
#> factor.T12_10.1.1 T12_10 36 0 36 0.8056
#> factor.T14_02.1.1 T14_02 36 0 36 0.1389
#> factor.T14_05.1.1 T14_05 36 0 36 0.2222
#> factor.T12_02.1.1 T12_02 36 0 36 0.5833
#> factor.T12_04.1.1 T12_04 36 0 36 0.7778
#> factor.T14_01.1.1 T14_01 36 0 36 0.1111
#> factor.T12_09.1.1 T12_09 36 0 36 0.9444
#> Q matrix specifies 1 dimension(s).
#>
#>
#> =====================================================================
#> Model No. 10
#> Model name: domainlistening__group1.all_group2.countryA
#> Number of items: 16
#> Number of persons: 74
#> Number of dimensions: 1
#> =====================================================================
#>
#> Following 118 item(s) missed in data frame will be removed from Q matrix:
#> T12_11, T13_15, T15_03, T13_04, T13_07, T13_16, T13_01, T15_02, T15_01, T15_15, T15_16, T15_09, T13_12, T13_13, T15_14, T13_08, T13_06, T15_12, T13_10, T13_17, T15_11, T13_05, T13_09, T15_10, T15_13, T15_17, T15_18, T13_18, T16_08, T16_09, T16_11, T16_13, T16_01, T16_04, T16_10, T16_02, T16_12, T16_06, T16_07, T19_06, T19_08, T19_05, T19_11, T19_01, T19_02, T19_30, T19_03, T19_07, T19_04, T19_09, T21_11, T18_01, T21_01, T18_05, T21_12, T18_13, T21_10, T18_11, T21_05, T21_13, T18_08, T18_07, T21_09, T18_04, T18_02, T21_02, T21_06, T18_09, T21_04, T18_12, T21_14, T21_03, T18_10, T30_06, T29_04, T29_02, T30_02, T30_03, T30_01, T29_05, T29_03, T28_08, T28_09, T28_02, T28_03, T28_04, T28_01, T28_10, T28_06, T20_05, T20_09, T20_30, T20_08, T20_12, T20_06, T20_07, T20_02, T20_13, T20_03, T20_01, T20_04, T20_11, T20_10, T32_04, T31_03, T31_01, T31_04, T32_03, T32_02, T32_01, T17_02, T17_09, T17_10, T17_04, T17_03, T17_08, T17_06, T17_01
#> Warning: 1 testitem(s) are constants. Remove these items from the data set:
#> Item 'T12_05', only value '1' occurs: 74 valid responses.
#> Remove 1 test item(s) overall.
#> 1 subject(s) do not solve any item:
#> P02289 (15 false), P02289 (15 false) ...
#> Dataset is completely linked.
#> 'gauss' has been chosen for estimation method. Number of nodes was not explicitly specified. Set nodes to 20.
#> Q matrix specifies 1 dimension(s).
#>
#>
#> =====================================================================
#> Model No. 11
#> Model name: domainlistening__group1.all_group2.countryB
#> Number of items: 16
#> Number of persons: 62
#> Number of dimensions: 1
#> =====================================================================
#>
#> Following 118 item(s) missed in data frame will be removed from Q matrix:
#> T12_11, T13_15, T15_03, T13_04, T13_07, T13_16, T13_01, T15_02, T15_01, T15_15, T15_16, T15_09, T13_12, T13_13, T15_14, T13_08, T13_06, T15_12, T13_10, T13_17, T15_11, T13_05, T13_09, T15_10, T15_13, T15_17, T15_18, T13_18, T16_08, T16_09, T16_11, T16_13, T16_01, T16_04, T16_10, T16_02, T16_12, T16_06, T16_07, T19_06, T19_08, T19_05, T19_11, T19_01, T19_02, T19_30, T19_03, T19_07, T19_04, T19_09, T21_11, T18_01, T21_01, T18_05, T21_12, T18_13, T21_10, T18_11, T21_05, T21_13, T18_08, T18_07, T21_09, T18_04, T18_02, T21_02, T21_06, T18_09, T21_04, T18_12, T21_14, T21_03, T18_10, T30_06, T29_04, T29_02, T30_02, T30_03, T30_01, T29_05, T29_03, T28_08, T28_09, T28_02, T28_03, T28_04, T28_01, T28_10, T28_06, T20_05, T20_09, T20_30, T20_08, T20_12, T20_06, T20_07, T20_02, T20_13, T20_03, T20_01, T20_04, T20_11, T20_10, T32_04, T31_03, T31_01, T31_04, T32_03, T32_02, T32_01, T17_02, T17_09, T17_10, T17_04, T17_03, T17_08, T17_06, T17_01
#> Dataset is completely linked.
#> 'gauss' has been chosen for estimation method. Number of nodes was not explicitly specified. Set nodes to 20.
#> Q matrix specifies 1 dimension(s).
#>
#>
#> ================================================================
#> Model No. 12
#> Model name: domainlistening__group1.all_group2.all
#> Number of items: 16
#> Number of persons: 172
#> Number of dimensions: 1
#> ================================================================
#>
#> Following 118 item(s) missed in data frame will be removed from Q matrix:
#> T12_11, T13_15, T15_03, T13_04, T13_07, T13_16, T13_01, T15_02, T15_01, T15_15, T15_16, T15_09, T13_12, T13_13, T15_14, T13_08, T13_06, T15_12, T13_10, T13_17, T15_11, T13_05, T13_09, T15_10, T15_13, T15_17, T15_18, T13_18, T16_08, T16_09, T16_11, T16_13, T16_01, T16_04, T16_10, T16_02, T16_12, T16_06, T16_07, T19_06, T19_08, T19_05, T19_11, T19_01, T19_02, T19_30, T19_03, T19_07, T19_04, T19_09, T21_11, T18_01, T21_01, T18_05, T21_12, T18_13, T21_10, T18_11, T21_05, T21_13, T18_08, T18_07, T21_09, T18_04, T18_02, T21_02, T21_06, T18_09, T21_04, T18_12, T21_14, T21_03, T18_10, T30_06, T29_04, T29_02, T30_02, T30_03, T30_01, T29_05, T29_03, T28_08, T28_09, T28_02, T28_03, T28_04, T28_01, T28_10, T28_06, T20_05, T20_09, T20_30, T20_08, T20_12, T20_06, T20_07, T20_02, T20_13, T20_03, T20_01, T20_04, T20_11, T20_10, T32_04, T31_03, T31_01, T31_04, T32_03, T32_02, T32_01, T17_02, T17_09, T17_10, T17_04, T17_03, T17_08, T17_06, T17_01
#> Dataset is completely linked.
#> 'gauss' has been chosen for estimation method. Number of nodes was not explicitly specified. Set nodes to 20.
#> Q matrix specifies 1 dimension(s).
#>
#>
#> ======================================================================
#> Model No. 13
#> Model name: domainreading__group1.female_group2.countryC
#> Number of items: 17
#> Number of persons: 20
#> Number of dimensions: 1
#> ======================================================================
#>
#> Following 132 item(s) missed in data frame will be removed from Q matrix:
#> T07_08, T02_02, T07_01, T02_06, T07_05, T07_06, T02_07, T07_02, T07_04, T02_03, T07_09, T02_01, T02_04, T07_03, T02_05, T07_07, T07_10, T09_12, T01_08, T03_07, T03_01, T03_08, T03_05, T03_06, T03_03, T10_08, T11_06, T11_03, T11_01, T04_02, T04_01, T04_06, T04_04, T11_04, T10_01, T11_08, T10_02, T10_09, T10_07, T11_02, T04_05, T04_03, T11_05, T10_06, T11_10, T11_09, T04_07, T10_10, T04_09, T04_08, T11_0X, T06_05, T05_03, T05_06, T05_05, T06_02, T06_04, T06_03, T05_04, T05_02, T05_01, T06_01, T08_04, T08_06, T08_03, T08_07, T08_02, T08_05, T08_01, T08_09, T08_08, T03_09, T06_1X, T26_03, T26_05, T26_07, T26_09, T26_06, T26_10, T26_08, T26_02, T26_04, T27_01, T27_05, T27_07, T27_06, T27_04, T27_02, T27_03, T24_05, T24_08, T24_02, T24_10, T24_06, T24_24, T24_25, T24_09, T24_21, T24_23, T23_10, T23_13, T23_09, T23_15, T23_03, T23_04, T23_14, T23_07, T23_08, T23_02, T23_06, T23_05, T25_05, T25_22, T22_03, T25_06, T22_12, T25_09, T25_10, T25_08, T22_11, T22_02, T25_11, T25_07, T25_14, T25_12, T25_04, T22_08, T25_23, T22_10, T22_06, T25_13, T22_01
#> Warning: 17 testitem(s) with less than 50 valid responses: These items are nevertheless kept in the data set: 'T09_03', 'T09_05', 'T01_03', 'T09_02', 'T01_06', 'T01_01', 'T09_10', 'T01_02', 'T09_06', 'T01_04', 'T09_09', 'T09_11', 'T01_05', 'T09_08', 'T09_07', 'T09_04', 'T01_07'
#> Warning: 1 testitem(s) are constants. Remove these items from the data set:
#> Item 'T01_01', only value '1' occurs: 20 valid responses.
#> Remove 1 test item(s) overall.
#> Dataset is completely linked.
#> 'gauss' has been chosen for estimation method. Number of nodes was not explicitly specified. Set nodes to 20.
#> Following 16 items with less than 50 item responses:
#> item.name cases Missing valid item.p
#> factor.T09_03.1.1 T09_03 20 0 20 0.65
#> factor.T09_05.1.1 T09_05 20 0 20 0.90
#> factor.T01_03.1.1 T01_03 20 0 20 0.90
#> factor.T09_02.1.1 T09_02 20 0 20 0.60
#> factor.T01_06.1.1 T01_06 20 0 20 0.50
#> factor.T09_10.1.1 T09_10 20 0 20 0.95
#> factor.T01_02.1.1 T01_02 20 0 20 0.65
#> factor.T09_06.1.1 T09_06 20 0 20 0.85
#> factor.T01_04.1.1 T01_04 20 0 20 0.55
#> factor.T09_09.1.1 T09_09 20 0 20 0.45
#> factor.T09_11.1.1 T09_11 20 0 20 0.60
#> factor.T01_05.1.1 T01_05 20 0 20 0.55
#> factor.T09_08.1.1 T09_08 20 0 20 0.80
#> factor.T09_07.1.1 T09_07 20 0 20 0.75
#> factor.T09_04.1.1 T09_04 20 0 20 0.50
#> factor.T01_07.1.1 T01_07 20 0 20 0.55
#> Q matrix specifies 1 dimension(s).
#>
#>
#> ======================================================================
#> Model No. 14
#> Model name: domainreading__group1.female_group2.countryA
#> Number of items: 17
#> Number of persons: 42
#> Number of dimensions: 1
#> ======================================================================
#>
#> Following 132 item(s) missed in data frame will be removed from Q matrix:
#> T07_08, T02_02, T07_01, T02_06, T07_05, T07_06, T02_07, T07_02, T07_04, T02_03, T07_09, T02_01, T02_04, T07_03, T02_05, T07_07, T07_10, T09_12, T01_08, T03_07, T03_01, T03_08, T03_05, T03_06, T03_03, T10_08, T11_06, T11_03, T11_01, T04_02, T04_01, T04_06, T04_04, T11_04, T10_01, T11_08, T10_02, T10_09, T10_07, T11_02, T04_05, T04_03, T11_05, T10_06, T11_10, T11_09, T04_07, T10_10, T04_09, T04_08, T11_0X, T06_05, T05_03, T05_06, T05_05, T06_02, T06_04, T06_03, T05_04, T05_02, T05_01, T06_01, T08_04, T08_06, T08_03, T08_07, T08_02, T08_05, T08_01, T08_09, T08_08, T03_09, T06_1X, T26_03, T26_05, T26_07, T26_09, T26_06, T26_10, T26_08, T26_02, T26_04, T27_01, T27_05, T27_07, T27_06, T27_04, T27_02, T27_03, T24_05, T24_08, T24_02, T24_10, T24_06, T24_24, T24_25, T24_09, T24_21, T24_23, T23_10, T23_13, T23_09, T23_15, T23_03, T23_04, T23_14, T23_07, T23_08, T23_02, T23_06, T23_05, T25_05, T25_22, T22_03, T25_06, T22_12, T25_09, T25_10, T25_08, T22_11, T22_02, T25_11, T25_07, T25_14, T25_12, T25_04, T22_08, T25_23, T22_10, T22_06, T25_13, T22_01
#> Warning: 17 testitem(s) with less than 50 valid responses: These items are nevertheless kept in the data set: 'T09_03', 'T09_05', 'T01_03', 'T09_02', 'T01_06', 'T01_01', 'T09_10', 'T01_02', 'T09_06', 'T01_04', 'T09_09', 'T09_11', 'T01_05', 'T09_08', 'T09_07', 'T09_04', 'T01_07'
#> Dataset is completely linked.
#> 'gauss' has been chosen for estimation method. Number of nodes was not explicitly specified. Set nodes to 20.
#> Following 17 items with less than 50 item responses:
#> item.name cases Missing valid item.p
#> factor.T09_03.1.1 T09_03 42 0 42 0.643
#> factor.T09_05.1.1 T09_05 42 0 42 0.976
#> factor.T01_03.1.1 T01_03 42 0 42 0.929
#> factor.T09_02.1.1 T09_02 42 0 42 0.690
#> factor.T01_06.1.1 T01_06 42 0 42 0.333
#> factor.T01_01.1.1 T01_01 42 0 42 0.929
#> factor.T09_10.1.1 T09_10 42 0 42 0.857
#> factor.T01_02.1.1 T01_02 42 0 42 0.762
#> factor.T09_06.1.1 T09_06 42 0 42 0.857
#> factor.T01_04.1.1 T01_04 42 0 42 0.333
#> factor.T09_09.1.1 T09_09 42 0 42 0.714
#> factor.T09_11.1.1 T09_11 42 0 42 0.667
#> factor.T01_05.1.1 T01_05 42 0 42 0.405
#> factor.T09_08.1.1 T09_08 42 0 42 0.643
#> factor.T09_07.1.1 T09_07 42 0 42 0.905
#> factor.T09_04.1.1 T09_04 42 0 42 0.500
#> factor.T01_07.1.1 T01_07 42 0 42 0.595
#> Q matrix specifies 1 dimension(s).
#>
#>
#> ======================================================================
#> Model No. 15
#> Model name: domainreading__group1.female_group2.countryB
#> Number of items: 17
#> Number of persons: 27
#> Number of dimensions: 1
#> ======================================================================
#>
#> Following 132 item(s) missed in data frame will be removed from Q matrix:
#> T07_08, T02_02, T07_01, T02_06, T07_05, T07_06, T02_07, T07_02, T07_04, T02_03, T07_09, T02_01, T02_04, T07_03, T02_05, T07_07, T07_10, T09_12, T01_08, T03_07, T03_01, T03_08, T03_05, T03_06, T03_03, T10_08, T11_06, T11_03, T11_01, T04_02, T04_01, T04_06, T04_04, T11_04, T10_01, T11_08, T10_02, T10_09, T10_07, T11_02, T04_05, T04_03, T11_05, T10_06, T11_10, T11_09, T04_07, T10_10, T04_09, T04_08, T11_0X, T06_05, T05_03, T05_06, T05_05, T06_02, T06_04, T06_03, T05_04, T05_02, T05_01, T06_01, T08_04, T08_06, T08_03, T08_07, T08_02, T08_05, T08_01, T08_09, T08_08, T03_09, T06_1X, T26_03, T26_05, T26_07, T26_09, T26_06, T26_10, T26_08, T26_02, T26_04, T27_01, T27_05, T27_07, T27_06, T27_04, T27_02, T27_03, T24_05, T24_08, T24_02, T24_10, T24_06, T24_24, T24_25, T24_09, T24_21, T24_23, T23_10, T23_13, T23_09, T23_15, T23_03, T23_04, T23_14, T23_07, T23_08, T23_02, T23_06, T23_05, T25_05, T25_22, T22_03, T25_06, T22_12, T25_09, T25_10, T25_08, T22_11, T22_02, T25_11, T25_07, T25_14, T25_12, T25_04, T22_08, T25_23, T22_10, T22_06, T25_13, T22_01
#> Warning: 17 testitem(s) with less than 50 valid responses: These items are nevertheless kept in the data set: 'T09_03', 'T09_05', 'T01_03', 'T09_02', 'T01_06', 'T01_01', 'T09_10', 'T01_02', 'T09_06', 'T01_04', 'T09_09', 'T09_11', 'T01_05', 'T09_08', 'T09_07', 'T09_04', 'T01_07'
#> Warning: 1 testitem(s) are constants. Remove these items from the data set:
#> Item 'T01_03', only value '1' occurs: 27 valid responses.
#> Remove 1 test item(s) overall.
#> Dataset is completely linked.
#> 'gauss' has been chosen for estimation method. Number of nodes was not explicitly specified. Set nodes to 20.
#> Following 16 items with less than 50 item responses:
#> item.name cases Missing valid item.p
#> factor.T09_03.1.1 T09_03 27 0 27 0.630
#> factor.T09_05.1.1 T09_05 27 0 27 0.889
#> factor.T09_02.1.1 T09_02 27 0 27 0.630
#> factor.T01_06.1.1 T01_06 27 0 27 0.519
#> factor.T01_01.1.1 T01_01 27 0 27 0.889
#> factor.T09_10.1.1 T09_10 27 0 27 0.926
#> factor.T01_02.1.1 T01_02 27 0 27 0.926
#> factor.T09_06.1.1 T09_06 27 0 27 0.963
#> factor.T01_04.1.1 T01_04 27 0 27 0.444
#> factor.T09_09.1.1 T09_09 27 0 27 0.593
#> factor.T09_11.1.1 T09_11 27 0 27 0.667
#> factor.T01_05.1.1 T01_05 27 0 27 0.444
#> factor.T09_08.1.1 T09_08 27 0 27 0.704
#> factor.T09_07.1.1 T09_07 27 0 27 0.963
#> factor.T09_04.1.1 T09_04 27 0 27 0.407
#> factor.T01_07.1.1 T01_07 27 0 27 0.852
#> Q matrix specifies 1 dimension(s).
#>
#>
#> =================================================================
#> Model No. 16
#> Model name: domainreading__group1.female_group2.all
#> Number of items: 17
#> Number of persons: 89
#> Number of dimensions: 1
#> =================================================================
#>
#> Following 132 item(s) missed in data frame will be removed from Q matrix:
#> T07_08, T02_02, T07_01, T02_06, T07_05, T07_06, T02_07, T07_02, T07_04, T02_03, T07_09, T02_01, T02_04, T07_03, T02_05, T07_07, T07_10, T09_12, T01_08, T03_07, T03_01, T03_08, T03_05, T03_06, T03_03, T10_08, T11_06, T11_03, T11_01, T04_02, T04_01, T04_06, T04_04, T11_04, T10_01, T11_08, T10_02, T10_09, T10_07, T11_02, T04_05, T04_03, T11_05, T10_06, T11_10, T11_09, T04_07, T10_10, T04_09, T04_08, T11_0X, T06_05, T05_03, T05_06, T05_05, T06_02, T06_04, T06_03, T05_04, T05_02, T05_01, T06_01, T08_04, T08_06, T08_03, T08_07, T08_02, T08_05, T08_01, T08_09, T08_08, T03_09, T06_1X, T26_03, T26_05, T26_07, T26_09, T26_06, T26_10, T26_08, T26_02, T26_04, T27_01, T27_05, T27_07, T27_06, T27_04, T27_02, T27_03, T24_05, T24_08, T24_02, T24_10, T24_06, T24_24, T24_25, T24_09, T24_21, T24_23, T23_10, T23_13, T23_09, T23_15, T23_03, T23_04, T23_14, T23_07, T23_08, T23_02, T23_06, T23_05, T25_05, T25_22, T22_03, T25_06, T22_12, T25_09, T25_10, T25_08, T22_11, T22_02, T25_11, T25_07, T25_14, T25_12, T25_04, T22_08, T25_23, T22_10, T22_06, T25_13, T22_01
#> Dataset is completely linked.
#> 'gauss' has been chosen for estimation method. Number of nodes was not explicitly specified. Set nodes to 20.
#> Q matrix specifies 1 dimension(s).
#>
#>
#> ====================================================================
#> Model No. 17
#> Model name: domainreading__group1.male_group2.countryC
#> Number of items: 17
#> Number of persons: 16
#> Number of dimensions: 1
#> ====================================================================
#>
#> Following 132 item(s) missed in data frame will be removed from Q matrix:
#> T07_08, T02_02, T07_01, T02_06, T07_05, T07_06, T02_07, T07_02, T07_04, T02_03, T07_09, T02_01, T02_04, T07_03, T02_05, T07_07, T07_10, T09_12, T01_08, T03_07, T03_01, T03_08, T03_05, T03_06, T03_03, T10_08, T11_06, T11_03, T11_01, T04_02, T04_01, T04_06, T04_04, T11_04, T10_01, T11_08, T10_02, T10_09, T10_07, T11_02, T04_05, T04_03, T11_05, T10_06, T11_10, T11_09, T04_07, T10_10, T04_09, T04_08, T11_0X, T06_05, T05_03, T05_06, T05_05, T06_02, T06_04, T06_03, T05_04, T05_02, T05_01, T06_01, T08_04, T08_06, T08_03, T08_07, T08_02, T08_05, T08_01, T08_09, T08_08, T03_09, T06_1X, T26_03, T26_05, T26_07, T26_09, T26_06, T26_10, T26_08, T26_02, T26_04, T27_01, T27_05, T27_07, T27_06, T27_04, T27_02, T27_03, T24_05, T24_08, T24_02, T24_10, T24_06, T24_24, T24_25, T24_09, T24_21, T24_23, T23_10, T23_13, T23_09, T23_15, T23_03, T23_04, T23_14, T23_07, T23_08, T23_02, T23_06, T23_05, T25_05, T25_22, T22_03, T25_06, T22_12, T25_09, T25_10, T25_08, T22_11, T22_02, T25_11, T25_07, T25_14, T25_12, T25_04, T22_08, T25_23, T22_10, T22_06, T25_13, T22_01
#> Warning: 17 testitem(s) with less than 50 valid responses: These items are nevertheless kept in the data set: 'T09_03', 'T09_05', 'T01_03', 'T09_02', 'T01_06', 'T01_01', 'T09_10', 'T01_02', 'T09_06', 'T01_04', 'T09_09', 'T09_11', 'T01_05', 'T09_08', 'T09_07', 'T09_04', 'T01_07'
#> Dataset is completely linked.
#> 'gauss' has been chosen for estimation method. Number of nodes was not explicitly specified. Set nodes to 20.
#> Following 17 items with less than 50 item responses:
#> item.name cases Missing valid item.p
#> factor.T09_03.1.1 T09_03 16 0 16 0.500
#> factor.T09_05.1.1 T09_05 16 0 16 0.875
#> factor.T01_03.1.1 T01_03 16 0 16 0.875
#> factor.T09_02.1.1 T09_02 16 0 16 0.812
#> factor.T01_06.1.1 T01_06 16 0 16 0.500
#> factor.T01_01.1.1 T01_01 16 0 16 0.938
#> factor.T09_10.1.1 T09_10 16 0 16 0.812
#> factor.T01_02.1.1 T01_02 16 0 16 0.500
#> factor.T09_06.1.1 T09_06 16 0 16 0.625
#> factor.T01_04.1.1 T01_04 16 0 16 0.375
#> factor.T09_09.1.1 T09_09 16 0 16 0.625
#> factor.T09_11.1.1 T09_11 16 0 16 0.500
#> factor.T01_05.1.1 T01_05 16 0 16 0.562
#> factor.T09_08.1.1 T09_08 16 0 16 0.625
#> factor.T09_07.1.1 T09_07 16 0 16 0.625
#> factor.T09_04.1.1 T09_04 16 0 16 0.438
#> factor.T01_07.1.1 T01_07 16 0 16 0.250
#> Q matrix specifies 1 dimension(s).
#>
#>
#> ====================================================================
#> Model No. 18
#> Model name: domainreading__group1.male_group2.countryA
#> Number of items: 17
#> Number of persons: 32
#> Number of dimensions: 1
#> ====================================================================
#>
#> Following 132 item(s) missed in data frame will be removed from Q matrix:
#> T07_08, T02_02, T07_01, T02_06, T07_05, T07_06, T02_07, T07_02, T07_04, T02_03, T07_09, T02_01, T02_04, T07_03, T02_05, T07_07, T07_10, T09_12, T01_08, T03_07, T03_01, T03_08, T03_05, T03_06, T03_03, T10_08, T11_06, T11_03, T11_01, T04_02, T04_01, T04_06, T04_04, T11_04, T10_01, T11_08, T10_02, T10_09, T10_07, T11_02, T04_05, T04_03, T11_05, T10_06, T11_10, T11_09, T04_07, T10_10, T04_09, T04_08, T11_0X, T06_05, T05_03, T05_06, T05_05, T06_02, T06_04, T06_03, T05_04, T05_02, T05_01, T06_01, T08_04, T08_06, T08_03, T08_07, T08_02, T08_05, T08_01, T08_09, T08_08, T03_09, T06_1X, T26_03, T26_05, T26_07, T26_09, T26_06, T26_10, T26_08, T26_02, T26_04, T27_01, T27_05, T27_07, T27_06, T27_04, T27_02, T27_03, T24_05, T24_08, T24_02, T24_10, T24_06, T24_24, T24_25, T24_09, T24_21, T24_23, T23_10, T23_13, T23_09, T23_15, T23_03, T23_04, T23_14, T23_07, T23_08, T23_02, T23_06, T23_05, T25_05, T25_22, T22_03, T25_06, T22_12, T25_09, T25_10, T25_08, T22_11, T22_02, T25_11, T25_07, T25_14, T25_12, T25_04, T22_08, T25_23, T22_10, T22_06, T25_13, T22_01
#> Warning: 17 testitem(s) with less than 50 valid responses: These items are nevertheless kept in the data set: 'T09_03', 'T09_05', 'T01_03', 'T09_02', 'T01_06', 'T01_01', 'T09_10', 'T01_02', 'T09_06', 'T01_04', 'T09_09', 'T09_11', 'T01_05', 'T09_08', 'T09_07', 'T09_04', 'T01_07'
#> 1 subject(s) solved each item: P02635 (17 correct), P02635 (17 correct) ...
#> Dataset is completely linked.
#> 'gauss' has been chosen for estimation method. Number of nodes was not explicitly specified. Set nodes to 20.
#> Following 17 items with less than 50 item responses:
#> item.name cases Missing valid item.p
#> factor.T09_03.1.1 T09_03 32 0 32 0.719
#> factor.T09_05.1.1 T09_05 32 0 32 0.875
#> factor.T01_03.1.1 T01_03 32 0 32 0.750
#> factor.T09_02.1.1 T09_02 32 0 32 0.688
#> factor.T01_06.1.1 T01_06 32 0 32 0.312
#> factor.T01_01.1.1 T01_01 32 0 32 0.875
#> factor.T09_10.1.1 T09_10 32 0 32 0.750
#> factor.T01_02.1.1 T01_02 32 0 32 0.750
#> factor.T09_06.1.1 T09_06 32 0 32 0.812
#> factor.T01_04.1.1 T01_04 32 0 32 0.375
#> factor.T09_09.1.1 T09_09 32 0 32 0.656
#> factor.T09_11.1.1 T09_11 32 0 32 0.438
#> factor.T01_05.1.1 T01_05 32 0 32 0.281
#> factor.T09_08.1.1 T09_08 32 0 32 0.688
#> factor.T09_07.1.1 T09_07 32 0 32 0.781
#> factor.T09_04.1.1 T09_04 32 0 32 0.562
#> factor.T01_07.1.1 T01_07 32 0 32 0.469
#> Q matrix specifies 1 dimension(s).
#>
#>
#> ====================================================================
#> Model No. 19
#> Model name: domainreading__group1.male_group2.countryB
#> Number of items: 17
#> Number of persons: 35
#> Number of dimensions: 1
#> ====================================================================
#>
#> Following 132 item(s) missed in data frame will be removed from Q matrix:
#> T07_08, T02_02, T07_01, T02_06, T07_05, T07_06, T02_07, T07_02, T07_04, T02_03, T07_09, T02_01, T02_04, T07_03, T02_05, T07_07, T07_10, T09_12, T01_08, T03_07, T03_01, T03_08, T03_05, T03_06, T03_03, T10_08, T11_06, T11_03, T11_01, T04_02, T04_01, T04_06, T04_04, T11_04, T10_01, T11_08, T10_02, T10_09, T10_07, T11_02, T04_05, T04_03, T11_05, T10_06, T11_10, T11_09, T04_07, T10_10, T04_09, T04_08, T11_0X, T06_05, T05_03, T05_06, T05_05, T06_02, T06_04, T06_03, T05_04, T05_02, T05_01, T06_01, T08_04, T08_06, T08_03, T08_07, T08_02, T08_05, T08_01, T08_09, T08_08, T03_09, T06_1X, T26_03, T26_05, T26_07, T26_09, T26_06, T26_10, T26_08, T26_02, T26_04, T27_01, T27_05, T27_07, T27_06, T27_04, T27_02, T27_03, T24_05, T24_08, T24_02, T24_10, T24_06, T24_24, T24_25, T24_09, T24_21, T24_23, T23_10, T23_13, T23_09, T23_15, T23_03, T23_04, T23_14, T23_07, T23_08, T23_02, T23_06, T23_05, T25_05, T25_22, T22_03, T25_06, T22_12, T25_09, T25_10, T25_08, T22_11, T22_02, T25_11, T25_07, T25_14, T25_12, T25_04, T22_08, T25_23, T22_10, T22_06, T25_13, T22_01
#> Warning: 17 testitem(s) with less than 50 valid responses: These items are nevertheless kept in the data set: 'T09_03', 'T09_05', 'T01_03', 'T09_02', 'T01_06', 'T01_01', 'T09_10', 'T01_02', 'T09_06', 'T01_04', 'T09_09', 'T09_11', 'T01_05', 'T09_08', 'T09_07', 'T09_04', 'T01_07'
#> 2 subject(s) solved each item: P03303 (17 correct), P03306 (17 correct), P03306 (17 correct) ...
#> Dataset is completely linked.
#> 'gauss' has been chosen for estimation method. Number of nodes was not explicitly specified. Set nodes to 20.
#> Following 17 items with less than 50 item responses:
#> item.name cases Missing valid item.p
#> factor.T09_03.1.1 T09_03 35 0 35 0.714
#> factor.T09_05.1.1 T09_05 35 0 35 0.971
#> factor.T01_03.1.1 T01_03 35 0 35 0.914
#> factor.T09_02.1.1 T09_02 35 0 35 0.629
#> factor.T01_06.1.1 T01_06 35 0 35 0.457
#> factor.T01_01.1.1 T01_01 35 0 35 0.914
#> factor.T09_10.1.1 T09_10 35 0 35 0.971
#> factor.T01_02.1.1 T01_02 35 0 35 0.857
#> factor.T09_06.1.1 T09_06 35 0 35 0.857
#> factor.T01_04.1.1 T01_04 35 0 35 0.457
#> factor.T09_09.1.1 T09_09 35 0 35 0.571
#> factor.T09_11.1.1 T09_11 35 0 35 0.743
#> factor.T01_05.1.1 T01_05 35 0 35 0.514
#> factor.T09_08.1.1 T09_08 35 0 35 0.829
#> factor.T09_07.1.1 T09_07 35 0 35 0.857
#> factor.T09_04.1.1 T09_04 35 0 35 0.486
#> factor.T01_07.1.1 T01_07 35 0 35 0.686
#> Q matrix specifies 1 dimension(s).
#>
#>
#> ===============================================================
#> Model No. 20
#> Model name: domainreading__group1.male_group2.all
#> Number of items: 17
#> Number of persons: 83
#> Number of dimensions: 1
#> ===============================================================
#>
#> Following 132 item(s) missed in data frame will be removed from Q matrix:
#> T07_08, T02_02, T07_01, T02_06, T07_05, T07_06, T02_07, T07_02, T07_04, T02_03, T07_09, T02_01, T02_04, T07_03, T02_05, T07_07, T07_10, T09_12, T01_08, T03_07, T03_01, T03_08, T03_05, T03_06, T03_03, T10_08, T11_06, T11_03, T11_01, T04_02, T04_01, T04_06, T04_04, T11_04, T10_01, T11_08, T10_02, T10_09, T10_07, T11_02, T04_05, T04_03, T11_05, T10_06, T11_10, T11_09, T04_07, T10_10, T04_09, T04_08, T11_0X, T06_05, T05_03, T05_06, T05_05, T06_02, T06_04, T06_03, T05_04, T05_02, T05_01, T06_01, T08_04, T08_06, T08_03, T08_07, T08_02, T08_05, T08_01, T08_09, T08_08, T03_09, T06_1X, T26_03, T26_05, T26_07, T26_09, T26_06, T26_10, T26_08, T26_02, T26_04, T27_01, T27_05, T27_07, T27_06, T27_04, T27_02, T27_03, T24_05, T24_08, T24_02, T24_10, T24_06, T24_24, T24_25, T24_09, T24_21, T24_23, T23_10, T23_13, T23_09, T23_15, T23_03, T23_04, T23_14, T23_07, T23_08, T23_02, T23_06, T23_05, T25_05, T25_22, T22_03, T25_06, T22_12, T25_09, T25_10, T25_08, T22_11, T22_02, T25_11, T25_07, T25_14, T25_12, T25_04, T22_08, T25_23, T22_10, T22_06, T25_13, T22_01
#> 3 subject(s) solved each item: P02635 (17 correct), P03303 (17 correct), P03306 (17 correct) ...
#> Dataset is completely linked.
#> 'gauss' has been chosen for estimation method. Number of nodes was not explicitly specified. Set nodes to 20.
#> Q matrix specifies 1 dimension(s).
#>
#>
#> ===================================================================
#> Model No. 21
#> Model name: domainreading__group1.all_group2.countryC
#> Number of items: 17
#> Number of persons: 36
#> Number of dimensions: 1
#> ===================================================================
#>
#> Following 132 item(s) missed in data frame will be removed from Q matrix:
#> T07_08, T02_02, T07_01, T02_06, T07_05, T07_06, T02_07, T07_02, T07_04, T02_03, T07_09, T02_01, T02_04, T07_03, T02_05, T07_07, T07_10, T09_12, T01_08, T03_07, T03_01, T03_08, T03_05, T03_06, T03_03, T10_08, T11_06, T11_03, T11_01, T04_02, T04_01, T04_06, T04_04, T11_04, T10_01, T11_08, T10_02, T10_09, T10_07, T11_02, T04_05, T04_03, T11_05, T10_06, T11_10, T11_09, T04_07, T10_10, T04_09, T04_08, T11_0X, T06_05, T05_03, T05_06, T05_05, T06_02, T06_04, T06_03, T05_04, T05_02, T05_01, T06_01, T08_04, T08_06, T08_03, T08_07, T08_02, T08_05, T08_01, T08_09, T08_08, T03_09, T06_1X, T26_03, T26_05, T26_07, T26_09, T26_06, T26_10, T26_08, T26_02, T26_04, T27_01, T27_05, T27_07, T27_06, T27_04, T27_02, T27_03, T24_05, T24_08, T24_02, T24_10, T24_06, T24_24, T24_25, T24_09, T24_21, T24_23, T23_10, T23_13, T23_09, T23_15, T23_03, T23_04, T23_14, T23_07, T23_08, T23_02, T23_06, T23_05, T25_05, T25_22, T22_03, T25_06, T22_12, T25_09, T25_10, T25_08, T22_11, T22_02, T25_11, T25_07, T25_14, T25_12, T25_04, T22_08, T25_23, T22_10, T22_06, T25_13, T22_01
#> Warning: 17 testitem(s) with less than 50 valid responses: These items are nevertheless kept in the data set: 'T09_03', 'T09_05', 'T01_03', 'T09_02', 'T01_06', 'T01_01', 'T09_10', 'T01_02', 'T09_06', 'T01_04', 'T09_09', 'T09_11', 'T01_05', 'T09_08', 'T09_07', 'T09_04', 'T01_07'
#> Dataset is completely linked.
#> 'gauss' has been chosen for estimation method. Number of nodes was not explicitly specified. Set nodes to 20.
#> Following 17 items with less than 50 item responses:
#> item.name cases Missing valid item.p
#> factor.T09_03.1.1 T09_03 36 0 36 0.583
#> factor.T09_05.1.1 T09_05 36 0 36 0.889
#> factor.T01_03.1.1 T01_03 36 0 36 0.889
#> factor.T09_02.1.1 T09_02 36 0 36 0.694
#> factor.T01_06.1.1 T01_06 36 0 36 0.500
#> factor.T01_01.1.1 T01_01 36 0 36 0.972
#> factor.T09_10.1.1 T09_10 36 0 36 0.889
#> factor.T01_02.1.1 T01_02 36 0 36 0.583
#> factor.T09_06.1.1 T09_06 36 0 36 0.750
#> factor.T01_04.1.1 T01_04 36 0 36 0.472
#> factor.T09_09.1.1 T09_09 36 0 36 0.528
#> factor.T09_11.1.1 T09_11 36 0 36 0.556
#> factor.T01_05.1.1 T01_05 36 0 36 0.556
#> factor.T09_08.1.1 T09_08 36 0 36 0.722
#> factor.T09_07.1.1 T09_07 36 0 36 0.694
#> factor.T09_04.1.1 T09_04 36 0 36 0.472
#> factor.T01_07.1.1 T01_07 36 0 36 0.417
#> Q matrix specifies 1 dimension(s).
#>
#>
#> ===================================================================
#> Model No. 22
#> Model name: domainreading__group1.all_group2.countryA
#> Number of items: 17
#> Number of persons: 74
#> Number of dimensions: 1
#> ===================================================================
#>
#> Following 132 item(s) missed in data frame will be removed from Q matrix:
#> T07_08, T02_02, T07_01, T02_06, T07_05, T07_06, T02_07, T07_02, T07_04, T02_03, T07_09, T02_01, T02_04, T07_03, T02_05, T07_07, T07_10, T09_12, T01_08, T03_07, T03_01, T03_08, T03_05, T03_06, T03_03, T10_08, T11_06, T11_03, T11_01, T04_02, T04_01, T04_06, T04_04, T11_04, T10_01, T11_08, T10_02, T10_09, T10_07, T11_02, T04_05, T04_03, T11_05, T10_06, T11_10, T11_09, T04_07, T10_10, T04_09, T04_08, T11_0X, T06_05, T05_03, T05_06, T05_05, T06_02, T06_04, T06_03, T05_04, T05_02, T05_01, T06_01, T08_04, T08_06, T08_03, T08_07, T08_02, T08_05, T08_01, T08_09, T08_08, T03_09, T06_1X, T26_03, T26_05, T26_07, T26_09, T26_06, T26_10, T26_08, T26_02, T26_04, T27_01, T27_05, T27_07, T27_06, T27_04, T27_02, T27_03, T24_05, T24_08, T24_02, T24_10, T24_06, T24_24, T24_25, T24_09, T24_21, T24_23, T23_10, T23_13, T23_09, T23_15, T23_03, T23_04, T23_14, T23_07, T23_08, T23_02, T23_06, T23_05, T25_05, T25_22, T22_03, T25_06, T22_12, T25_09, T25_10, T25_08, T22_11, T22_02, T25_11, T25_07, T25_14, T25_12, T25_04, T22_08, T25_23, T22_10, T22_06, T25_13, T22_01
#> 1 subject(s) solved each item: P02635 (17 correct), P02635 (17 correct) ...
#> Dataset is completely linked.
#> 'gauss' has been chosen for estimation method. Number of nodes was not explicitly specified. Set nodes to 20.
#> Q matrix specifies 1 dimension(s).
#>
#>
#> ===================================================================
#> Model No. 23
#> Model name: domainreading__group1.all_group2.countryB
#> Number of items: 17
#> Number of persons: 62
#> Number of dimensions: 1
#> ===================================================================
#>
#> Following 132 item(s) missed in data frame will be removed from Q matrix:
#> T07_08, T02_02, T07_01, T02_06, T07_05, T07_06, T02_07, T07_02, T07_04, T02_03, T07_09, T02_01, T02_04, T07_03, T02_05, T07_07, T07_10, T09_12, T01_08, T03_07, T03_01, T03_08, T03_05, T03_06, T03_03, T10_08, T11_06, T11_03, T11_01, T04_02, T04_01, T04_06, T04_04, T11_04, T10_01, T11_08, T10_02, T10_09, T10_07, T11_02, T04_05, T04_03, T11_05, T10_06, T11_10, T11_09, T04_07, T10_10, T04_09, T04_08, T11_0X, T06_05, T05_03, T05_06, T05_05, T06_02, T06_04, T06_03, T05_04, T05_02, T05_01, T06_01, T08_04, T08_06, T08_03, T08_07, T08_02, T08_05, T08_01, T08_09, T08_08, T03_09, T06_1X, T26_03, T26_05, T26_07, T26_09, T26_06, T26_10, T26_08, T26_02, T26_04, T27_01, T27_05, T27_07, T27_06, T27_04, T27_02, T27_03, T24_05, T24_08, T24_02, T24_10, T24_06, T24_24, T24_25, T24_09, T24_21, T24_23, T23_10, T23_13, T23_09, T23_15, T23_03, T23_04, T23_14, T23_07, T23_08, T23_02, T23_06, T23_05, T25_05, T25_22, T22_03, T25_06, T22_12, T25_09, T25_10, T25_08, T22_11, T22_02, T25_11, T25_07, T25_14, T25_12, T25_04, T22_08, T25_23, T22_10, T22_06, T25_13, T22_01
#> 2 subject(s) solved each item: P03303 (17 correct), P03306 (17 correct), P03306 (17 correct) ...
#> Dataset is completely linked.
#> 'gauss' has been chosen for estimation method. Number of nodes was not explicitly specified. Set nodes to 20.
#> Q matrix specifies 1 dimension(s).
#>
#>
#> ==============================================================
#> Model No. 24
#> Model name: domainreading__group1.all_group2.all
#> Number of items: 17
#> Number of persons: 172
#> Number of dimensions: 1
#> ==============================================================
#>
#> Following 132 item(s) missed in data frame will be removed from Q matrix:
#> T07_08, T02_02, T07_01, T02_06, T07_05, T07_06, T02_07, T07_02, T07_04, T02_03, T07_09, T02_01, T02_04, T07_03, T02_05, T07_07, T07_10, T09_12, T01_08, T03_07, T03_01, T03_08, T03_05, T03_06, T03_03, T10_08, T11_06, T11_03, T11_01, T04_02, T04_01, T04_06, T04_04, T11_04, T10_01, T11_08, T10_02, T10_09, T10_07, T11_02, T04_05, T04_03, T11_05, T10_06, T11_10, T11_09, T04_07, T10_10, T04_09, T04_08, T11_0X, T06_05, T05_03, T05_06, T05_05, T06_02, T06_04, T06_03, T05_04, T05_02, T05_01, T06_01, T08_04, T08_06, T08_03, T08_07, T08_02, T08_05, T08_01, T08_09, T08_08, T03_09, T06_1X, T26_03, T26_05, T26_07, T26_09, T26_06, T26_10, T26_08, T26_02, T26_04, T27_01, T27_05, T27_07, T27_06, T27_04, T27_02, T27_03, T24_05, T24_08, T24_02, T24_10, T24_06, T24_24, T24_25, T24_09, T24_21, T24_23, T23_10, T23_13, T23_09, T23_15, T23_03, T23_04, T23_14, T23_07, T23_08, T23_02, T23_06, T23_05, T25_05, T25_22, T22_03, T25_06, T22_12, T25_09, T25_10, T25_08, T22_11, T22_02, T25_11, T25_07, T25_14, T25_12, T25_04, T22_08, T25_23, T22_10, T22_06, T25_13, T22_01
#> 3 subject(s) solved each item: P02635 (17 correct), P03303 (17 correct), P03306 (17 correct) ...
#> Dataset is completely linked.
#> 'gauss' has been chosen for estimation method. Number of nodes was not explicitly specified. Set nodes to 20.
#> Q matrix specifies 1 dimension(s).
# run all models
runMul<- runModel(modMul)
# get results of all models
resMul<- getResults(runMul)
#> |*****|
#> |-----|
#> |*****|
#> |-----|
#> |*****|
#> |-----|
#> |*****|
#> |-----|
#> |*****|
#> |-----|
#> |*****|
#> |-----|
#> |*****|
#> |-----|
#> |*****|
#> |-----|
#> |*****|
#> |-----|
#> |*****|
#> |-----|
#> |*****|
#> |-----|
#> |*****|
#> |-----|
#> |*****|
#> |-----|
#> |*****|
#> |-----|
#> |*****|
#> |-----|
#> |*****|
#> |-----|
#> |*****|
#> |-----|
#> |*****|
#> |-----|
#> |*****|
#> |-----|
#> |*****|
#> |-----|
#> |*****|
#> |-----|
#> |*****|
#> |-----|
#> |*****|
#> |-----|
#> |*****|
#> |-----|
################################################################################
### Example 5: Linking and equating for multiple models ###
################################################################################
# Example 5: define und run multiple models according to different domains (item groups)
# and further linking/equating. This example mimics the routines necessary for the
# 'Vergleichsarbeiten' (VERA) at the Institute of Educational Progress (IQB)
# Let's assume that the 2015 cohort serves as the subsample for the 'Vergleichsarbeiten'.
vera <- subset(trends, booklet == "Bo02" & year == 2015) |>
reshape2::dcast(idstud+language+sex+ses+country~item, value.var = "value")
# specify two models according to the two domains 'reading' and 'listening'
l2 <- splitModels ( qMatrix = qMat, nCores = 1)
#> --------------------------------
#> splitModels: generating 2 models
#> ..
#> see <returned>$models
#> number of cores: 1
#> --------------------------------
# define 2 models. free estimation without anchoring
mods <- defineModel(dat = vera, id = "idstud", check.for.linking = TRUE,
splittedModels = l2, software = "tam")
#>
#> Specification of 'qMatrix' and 'person.groups' results in 2 model(s).
#> Warning: Model split preparation for model No. 1, model name domainlistening: 118 items from 134 items listed the Q matrix not found in data:
#> ℹ 'T12_02', 'T13_15', 'T15_03', 'T13_04', 'T13_07', 'T13_16', 'T13_01', 'T15_02', 'T15_01', 'T15_15', 'T15_16', 'T15_09', 'T13_12', 'T13_13', 'T15_14', 'T13_08', 'T13_06', 'T15_12', 'T13_10', 'T13_17', 'T15_11', 'T13_05', 'T13_09', 'T15_10', 'T15_13', 'T15_17', 'T15_18', 'T13_18', 'T16_08', 'T16_09', 'T16_11', 'T16_13', 'T16_01', 'T16_04', 'T16_10', 'T16_02', 'T16_12', 'T16_06', 'T16_07', 'T19_06', 'T19_08', 'T19_05', 'T19_11', 'T19_01', 'T19_02', 'T19_30', 'T19_03', 'T19_07', 'T19_04', 'T19_09', 'T21_11', 'T18_01', 'T21_01', 'T18_05', 'T21_12', 'T18_13', 'T21_10', 'T18_11', 'T21_05', 'T21_13', 'T18_08', 'T18_07', 'T21_09', 'T18_04', 'T18_02', 'T21_02', 'T21_06', 'T18_09', 'T21_04', 'T18_12', 'T21_14', 'T21_03', 'T18_10', 'T30_06', 'T29_04', 'T29_02', 'T30_02', 'T30_03', 'T30_01', 'T29_05', 'T29_03', 'T28_08', 'T28_09', 'T28_02', 'T28_03', 'T28_04', 'T28_01', 'T28_10', 'T28_06', 'T20_05', 'T20_09', 'T20_30', 'T20_08', 'T20_12', 'T20_06', 'T20_07', 'T20_02', 'T20_13', 'T20_03',
#> 'T20_01', 'T20_04', 'T20_11', 'T20_10', 'T32_04', 'T31_03', 'T31_01', 'T31_04', 'T32_03', 'T32_02', 'T32_01', 'T17_02', 'T17_09', 'T17_10', 'T17_04', 'T17_03', 'T17_08', 'T17_06', 'T17_01'
#> Warning: Model split preparation for model No. 2, model name domainreading: 132 items from 149 items listed the Q matrix not found in data:
#> ℹ 'T07_08', 'T02_02', 'T07_01', 'T02_06', 'T07_05', 'T07_06', 'T02_07', 'T07_02', 'T07_04', 'T02_03', 'T07_09', 'T02_01', 'T02_04', 'T07_03', 'T02_05', 'T07_07', 'T07_10', 'T09_11', 'T01_05', 'T03_07', 'T03_01', 'T03_08', 'T03_05', 'T03_06', 'T03_03', 'T10_08', 'T11_06', 'T11_03', 'T11_01', 'T04_02', 'T04_01', 'T04_06', 'T04_04', 'T11_04', 'T10_01', 'T11_08', 'T10_02', 'T10_09', 'T10_07', 'T11_02', 'T04_05', 'T04_03', 'T11_05', 'T10_06', 'T11_10', 'T11_09', 'T04_07', 'T10_10', 'T04_09', 'T04_08', 'T11_0X', 'T06_05', 'T05_03', 'T05_06', 'T05_05', 'T06_02', 'T06_04', 'T06_03', 'T05_04', 'T05_02', 'T05_01', 'T06_01', 'T08_04', 'T08_06', 'T08_03', 'T08_07', 'T08_02', 'T08_05', 'T08_01', 'T08_09', 'T08_08', 'T03_09', 'T06_1X', 'T26_03', 'T26_05', 'T26_07', 'T26_09', 'T26_06', 'T26_10', 'T26_08', 'T26_02', 'T26_04', 'T27_01', 'T27_05', 'T27_07', 'T27_06', 'T27_04', 'T27_02', 'T27_03', 'T24_05', 'T24_08', 'T24_02', 'T24_10', 'T24_06', 'T24_24', 'T24_25', 'T24_09', 'T24_21', 'T24_23',
#> 'T23_10', 'T23_13', 'T23_09', 'T23_15', 'T23_03', 'T23_04', 'T23_14', 'T23_07', 'T23_08', 'T23_02', 'T23_06', 'T23_05', 'T25_05', 'T25_22', 'T22_03', 'T25_06', 'T22_12', 'T25_09', 'T25_10', 'T25_08', 'T22_11', 'T22_02', 'T25_11', 'T25_07', 'T25_14', 'T25_12', 'T25_04', 'T22_08', 'T25_23', 'T22_10', 'T22_06', 'T25_13', 'T22_01'
#>
#>
#> =========================================
#> Model No. 1
#> Model name: domainlistening
#> Number of items: 16
#> Number of persons: 133
#> Number of dimensions: 1
#> =========================================
#>
#> Following 118 item(s) missed in data frame will be removed from Q matrix:
#> T12_02, T13_15, T15_03, T13_04, T13_07, T13_16, T13_01, T15_02, T15_01, T15_15, T15_16, T15_09, T13_12, T13_13, T15_14, T13_08, T13_06, T15_12, T13_10, T13_17, T15_11, T13_05, T13_09, T15_10, T15_13, T15_17, T15_18, T13_18, T16_08, T16_09, T16_11, T16_13, T16_01, T16_04, T16_10, T16_02, T16_12, T16_06, T16_07, T19_06, T19_08, T19_05, T19_11, T19_01, T19_02, T19_30, T19_03, T19_07, T19_04, T19_09, T21_11, T18_01, T21_01, T18_05, T21_12, T18_13, T21_10, T18_11, T21_05, T21_13, T18_08, T18_07, T21_09, T18_04, T18_02, T21_02, T21_06, T18_09, T21_04, T18_12, T21_14, T21_03, T18_10, T30_06, T29_04, T29_02, T30_02, T30_03, T30_01, T29_05, T29_03, T28_08, T28_09, T28_02, T28_03, T28_04, T28_01, T28_10, T28_06, T20_05, T20_09, T20_30, T20_08, T20_12, T20_06, T20_07, T20_02, T20_13, T20_03, T20_01, T20_04, T20_11, T20_10, T32_04, T31_03, T31_01, T31_04, T32_03, T32_02, T32_01, T17_02, T17_09, T17_10, T17_04, T17_03, T17_08, T17_06, T17_01
#> Dataset is completely linked.
#> 'gauss' has been chosen for estimation method. Number of nodes was not explicitly specified. Set nodes to 20.
#> Q matrix specifies 1 dimension(s).
#>
#>
#> =======================================
#> Model No. 2
#> Model name: domainreading
#> Number of items: 17
#> Number of persons: 133
#> Number of dimensions: 1
#> =======================================
#>
#> Following 132 item(s) missed in data frame will be removed from Q matrix:
#> T07_08, T02_02, T07_01, T02_06, T07_05, T07_06, T02_07, T07_02, T07_04, T02_03, T07_09, T02_01, T02_04, T07_03, T02_05, T07_07, T07_10, T09_11, T01_05, T03_07, T03_01, T03_08, T03_05, T03_06, T03_03, T10_08, T11_06, T11_03, T11_01, T04_02, T04_01, T04_06, T04_04, T11_04, T10_01, T11_08, T10_02, T10_09, T10_07, T11_02, T04_05, T04_03, T11_05, T10_06, T11_10, T11_09, T04_07, T10_10, T04_09, T04_08, T11_0X, T06_05, T05_03, T05_06, T05_05, T06_02, T06_04, T06_03, T05_04, T05_02, T05_01, T06_01, T08_04, T08_06, T08_03, T08_07, T08_02, T08_05, T08_01, T08_09, T08_08, T03_09, T06_1X, T26_03, T26_05, T26_07, T26_09, T26_06, T26_10, T26_08, T26_02, T26_04, T27_01, T27_05, T27_07, T27_06, T27_04, T27_02, T27_03, T24_05, T24_08, T24_02, T24_10, T24_06, T24_24, T24_25, T24_09, T24_21, T24_23, T23_10, T23_13, T23_09, T23_15, T23_03, T23_04, T23_14, T23_07, T23_08, T23_02, T23_06, T23_05, T25_05, T25_22, T22_03, T25_06, T22_12, T25_09, T25_10, T25_08, T22_11, T22_02, T25_11, T25_07, T25_14, T25_12, T25_04, T22_08, T25_23, T22_10, T22_06, T25_13, T22_01
#> 1 subject(s) solved each item: P04491 (17 correct), P04491 (17 correct) ...
#> Dataset is completely linked.
#> 'gauss' has been chosen for estimation method. Number of nodes was not explicitly specified. Set nodes to 20.
#> Q matrix specifies 1 dimension(s).
# run 2 models
runs <- runModel(mods)
# get the results
ress <- getResults(runs)
#> |*****|
#> |-----|
#> |*****|
#> |-----|
# norm parameters are the item parameters from example 1
aPar <- itemFromRes(res1)[,c("item", "est")]
#> Error: object 'res1' not found
# anchoring without exclusion of linking DIF items (DIF items will only be identified)
# the linking error is well above the critical threshold of 0.1
anch <- equat1pl ( results = ress, prmNorm = aPar, excludeLinkingDif = FALSE,
difBound = 0.64)
#> Found 2 model(s).
#> Equating is executed for each dimension in each model separately.
#> Error: object 'aPar' not found
# anchoring with exclusion of linking DIF items: especially for listening, the
# linking constant changes substantially when linking DIF items are excluded
anch2 <- equat1pl ( results = ress, prmNorm = aPar, excludeLinkingDif = TRUE,
difBound = 0.64, iterativ = FALSE)
#> Found 2 model(s).
#> Equating is executed for each dimension in each model separately.
#> Error: object 'aPar' not found
# anchoring with iterative exclusion of linking DIF items
anch3 <- equat1pl ( results = ress, prmNorm = aPar, excludeLinkingDif = TRUE,
difBound = 0.64, iterativ = TRUE)
#> Found 2 model(s).
#> Equating is executed for each dimension in each model separately.
#> Error: object 'aPar' not found
# transformation to the Bista metric
# first we arbitrarily define mean and standard deviation of the reference
# population according to both dimensions (defined in the Q matrix):
# reading and listening
# Note that the the first column of the 'refPop' data frame must include the
# domain names. Domain names must match the names defined in the Q matrix
refPop<- data.frame ( domain = c("domainreading", "domainlistening"),
m = c(0.03890191, 0.03587727), sd= c(1.219, 0.8978912))
# second, we specify a list with cut scores. Values must be in ascending order.
# Labels of the competence stages are optional. If no labels are specified,
# the will be defaulted to 1, 2, 3 ... etc.
# Note: if labels are specified, there must be one label more than cut scores.
# (i.e. 4 cut scores need 5 labels, etc.)
cuts <- list ( domainreading = list ( values = 390+0:3*75),
domainlistening = list ( values = 360+0:3*85))
# transformation
dfr <- transformToBista ( equatingList = anch3, refPop = refPop, cuts=cuts )
#> Error: object 'anch3' not found
head(dfr$itempars)
#> Error: object 'dfr' not found
head(dfr$personpars)
#> Error: object 'dfr' not found
################################################################################
### Example 5a: Linking for multiple models, including global domain ###
################################################################################
# Example 5a: define und run multiple models according to different domains (item groups)
# and further linking/equating. Same as example 5, but extended for the 'global'
# domain.
# add the 'global' domain (reading and listening together) in the Q matrix
qMat2 <- qMat |> dplyr::mutate(global = 1)
# specify two models according to the two domains 'reading' and 'listening'
l3 <- splitModels ( qMatrix = qMat2, nCores = 1)
#> --------------------------------
#> splitModels: generating 3 models
#> ...
#> see <returned>$models
#> number of cores: 1
#> --------------------------------
# define 3 models
mods5 <- defineModel(dat = vera, id = "idstud", check.for.linking = TRUE,
splittedModels = l3, software = "tam")
#>
#> Specification of 'qMatrix' and 'person.groups' results in 3 model(s).
#> Warning: Model split preparation for model No. 1, model name domainlistening: 118 items from 134 items listed the Q matrix not found in data:
#> ℹ 'T12_02', 'T13_15', 'T15_03', 'T13_04', 'T13_07', 'T13_16', 'T13_01', 'T15_02', 'T15_01', 'T15_15', 'T15_16', 'T15_09', 'T13_12', 'T13_13', 'T15_14', 'T13_08', 'T13_06', 'T15_12', 'T13_10', 'T13_17', 'T15_11', 'T13_05', 'T13_09', 'T15_10', 'T15_13', 'T15_17', 'T15_18', 'T13_18', 'T16_08', 'T16_09', 'T16_11', 'T16_13', 'T16_01', 'T16_04', 'T16_10', 'T16_02', 'T16_12', 'T16_06', 'T16_07', 'T19_06', 'T19_08', 'T19_05', 'T19_11', 'T19_01', 'T19_02', 'T19_30', 'T19_03', 'T19_07', 'T19_04', 'T19_09', 'T21_11', 'T18_01', 'T21_01', 'T18_05', 'T21_12', 'T18_13', 'T21_10', 'T18_11', 'T21_05', 'T21_13', 'T18_08', 'T18_07', 'T21_09', 'T18_04', 'T18_02', 'T21_02', 'T21_06', 'T18_09', 'T21_04', 'T18_12', 'T21_14', 'T21_03', 'T18_10', 'T30_06', 'T29_04', 'T29_02', 'T30_02', 'T30_03', 'T30_01', 'T29_05', 'T29_03', 'T28_08', 'T28_09', 'T28_02', 'T28_03', 'T28_04', 'T28_01', 'T28_10', 'T28_06', 'T20_05', 'T20_09', 'T20_30', 'T20_08', 'T20_12', 'T20_06', 'T20_07', 'T20_02', 'T20_13', 'T20_03',
#> 'T20_01', 'T20_04', 'T20_11', 'T20_10', 'T32_04', 'T31_03', 'T31_01', 'T31_04', 'T32_03', 'T32_02', 'T32_01', 'T17_02', 'T17_09', 'T17_10', 'T17_04', 'T17_03', 'T17_08', 'T17_06', 'T17_01'
#> Warning: Model split preparation for model No. 2, model name domainreading: 132 items from 149 items listed the Q matrix not found in data:
#> ℹ 'T07_08', 'T02_02', 'T07_01', 'T02_06', 'T07_05', 'T07_06', 'T02_07', 'T07_02', 'T07_04', 'T02_03', 'T07_09', 'T02_01', 'T02_04', 'T07_03', 'T02_05', 'T07_07', 'T07_10', 'T09_11', 'T01_05', 'T03_07', 'T03_01', 'T03_08', 'T03_05', 'T03_06', 'T03_03', 'T10_08', 'T11_06', 'T11_03', 'T11_01', 'T04_02', 'T04_01', 'T04_06', 'T04_04', 'T11_04', 'T10_01', 'T11_08', 'T10_02', 'T10_09', 'T10_07', 'T11_02', 'T04_05', 'T04_03', 'T11_05', 'T10_06', 'T11_10', 'T11_09', 'T04_07', 'T10_10', 'T04_09', 'T04_08', 'T11_0X', 'T06_05', 'T05_03', 'T05_06', 'T05_05', 'T06_02', 'T06_04', 'T06_03', 'T05_04', 'T05_02', 'T05_01', 'T06_01', 'T08_04', 'T08_06', 'T08_03', 'T08_07', 'T08_02', 'T08_05', 'T08_01', 'T08_09', 'T08_08', 'T03_09', 'T06_1X', 'T26_03', 'T26_05', 'T26_07', 'T26_09', 'T26_06', 'T26_10', 'T26_08', 'T26_02', 'T26_04', 'T27_01', 'T27_05', 'T27_07', 'T27_06', 'T27_04', 'T27_02', 'T27_03', 'T24_05', 'T24_08', 'T24_02', 'T24_10', 'T24_06', 'T24_24', 'T24_25', 'T24_09', 'T24_21', 'T24_23',
#> 'T23_10', 'T23_13', 'T23_09', 'T23_15', 'T23_03', 'T23_04', 'T23_14', 'T23_07', 'T23_08', 'T23_02', 'T23_06', 'T23_05', 'T25_05', 'T25_22', 'T22_03', 'T25_06', 'T22_12', 'T25_09', 'T25_10', 'T25_08', 'T22_11', 'T22_02', 'T25_11', 'T25_07', 'T25_14', 'T25_12', 'T25_04', 'T22_08', 'T25_23', 'T22_10', 'T22_06', 'T25_13', 'T22_01'
#> Warning: Model split preparation for model No. 3, model name global: 250 items from 283 items listed the Q matrix not found in data:
#> ℹ 'T12_02', 'T07_08', 'T02_02', 'T07_01', 'T02_06', 'T07_05', 'T07_06', 'T02_07', 'T07_02', 'T07_04', 'T02_03', 'T07_09', 'T02_01', 'T02_04', 'T07_03', 'T02_05', 'T07_07', 'T07_10', 'T09_11', 'T01_05', 'T13_15', 'T15_03', 'T13_04', 'T13_07', 'T13_16', 'T13_01', 'T15_02', 'T15_01', 'T15_15', 'T15_16', 'T15_09', 'T13_12', 'T13_13', 'T15_14', 'T13_08', 'T13_06', 'T15_12', 'T13_10', 'T13_17', 'T15_11', 'T13_05', 'T13_09', 'T15_10', 'T15_13', 'T03_07', 'T03_01', 'T03_08', 'T03_05', 'T03_06', 'T03_03', 'T10_08', 'T11_06', 'T11_03', 'T11_01', 'T04_02', 'T04_01', 'T04_06', 'T04_04', 'T11_04', 'T10_01', 'T11_08', 'T10_02', 'T10_09', 'T10_07', 'T11_02', 'T04_05', 'T04_03', 'T11_05', 'T10_06', 'T11_10', 'T11_09', 'T04_07', 'T15_17', 'T15_18', 'T13_18', 'T10_10', 'T04_09', 'T04_08', 'T11_0X', 'T16_08', 'T16_09', 'T16_11', 'T16_13', 'T16_01', 'T16_04', 'T16_10', 'T16_02', 'T16_12', 'T16_06', 'T16_07', 'T06_05', 'T05_03', 'T05_06', 'T05_05', 'T06_02', 'T06_04', 'T06_03', 'T05_04', 'T05_02',
#> 'T05_01', 'T06_01', 'T08_04', 'T08_06', 'T08_03', 'T08_07', 'T08_02', 'T08_05', 'T08_01', 'T08_09', 'T08_08', 'T03_09', 'T06_1X', 'T19_06', 'T19_08', 'T19_05', 'T19_11', 'T19_01', 'T19_02', 'T19_30', 'T19_03', 'T19_07', 'T19_04', 'T19_09', 'T26_03', 'T26_05', 'T26_07', 'T26_09', 'T26_06', 'T26_10', 'T26_08', 'T26_02', 'T26_04', 'T21_11', 'T18_01', 'T21_01', 'T18_05', 'T21_12', 'T18_13', 'T21_10', 'T18_11', 'T21_05', 'T21_13', 'T18_08', 'T18_07', 'T21_09', 'T18_04', 'T18_02', 'T21_02', 'T21_06', 'T18_09', 'T21_04', 'T18_12', 'T21_14', 'T21_03', 'T18_10', 'T27_01', 'T27_05', 'T27_07', 'T27_06', 'T27_04', 'T27_02', 'T27_03', 'T30_06', 'T29_04', 'T29_02', 'T30_02', 'T30_03', 'T30_01', 'T29_05', 'T29_03', 'T24_05', 'T24_08', 'T24_02', 'T24_10', 'T24_06', 'T24_24', 'T24_25', 'T24_09', 'T24_21', 'T24_23', 'T28_08', 'T28_09', 'T28_02', 'T28_03', 'T28_04', 'T28_01', 'T28_10', 'T28_06', 'T23_10', 'T23_13', 'T23_09', 'T23_15', 'T23_03', 'T23_04', 'T23_14', 'T23_07', 'T23_08', 'T23_02',
#> 'T23_06', 'T23_05', 'T25_05', 'T25_22', 'T22_03', 'T25_06', 'T22_12', 'T25_09', 'T25_10', 'T25_08', 'T22_11', 'T22_02', 'T25_11', 'T25_07', 'T25_14', 'T25_12', 'T25_04', 'T22_08', 'T25_23', 'T22_10', 'T22_06', 'T25_13', 'T22_01', 'T20_05', 'T20_09', 'T20_30', 'T20_08', 'T20_12', 'T20_06', 'T20_07', 'T20_02', 'T20_13', 'T20_03', 'T20_01', 'T20_04', 'T20_11', 'T20_10', 'T32_04', 'T31_03', 'T31_01', 'T31_04', 'T32_03', 'T32_02', 'T32_01', 'T17_02', 'T17_09', 'T17_10', 'T17_04', 'T17_03', 'T17_08', 'T17_06', 'T17_01'
#>
#>
#> =========================================
#> Model No. 1
#> Model name: domainlistening
#> Number of items: 16
#> Number of persons: 133
#> Number of dimensions: 1
#> =========================================
#>
#> Following 118 item(s) missed in data frame will be removed from Q matrix:
#> T12_02, T13_15, T15_03, T13_04, T13_07, T13_16, T13_01, T15_02, T15_01, T15_15, T15_16, T15_09, T13_12, T13_13, T15_14, T13_08, T13_06, T15_12, T13_10, T13_17, T15_11, T13_05, T13_09, T15_10, T15_13, T15_17, T15_18, T13_18, T16_08, T16_09, T16_11, T16_13, T16_01, T16_04, T16_10, T16_02, T16_12, T16_06, T16_07, T19_06, T19_08, T19_05, T19_11, T19_01, T19_02, T19_30, T19_03, T19_07, T19_04, T19_09, T21_11, T18_01, T21_01, T18_05, T21_12, T18_13, T21_10, T18_11, T21_05, T21_13, T18_08, T18_07, T21_09, T18_04, T18_02, T21_02, T21_06, T18_09, T21_04, T18_12, T21_14, T21_03, T18_10, T30_06, T29_04, T29_02, T30_02, T30_03, T30_01, T29_05, T29_03, T28_08, T28_09, T28_02, T28_03, T28_04, T28_01, T28_10, T28_06, T20_05, T20_09, T20_30, T20_08, T20_12, T20_06, T20_07, T20_02, T20_13, T20_03, T20_01, T20_04, T20_11, T20_10, T32_04, T31_03, T31_01, T31_04, T32_03, T32_02, T32_01, T17_02, T17_09, T17_10, T17_04, T17_03, T17_08, T17_06, T17_01
#> Dataset is completely linked.
#> 'gauss' has been chosen for estimation method. Number of nodes was not explicitly specified. Set nodes to 20.
#> Q matrix specifies 1 dimension(s).
#>
#>
#> =======================================
#> Model No. 2
#> Model name: domainreading
#> Number of items: 17
#> Number of persons: 133
#> Number of dimensions: 1
#> =======================================
#>
#> Following 132 item(s) missed in data frame will be removed from Q matrix:
#> T07_08, T02_02, T07_01, T02_06, T07_05, T07_06, T02_07, T07_02, T07_04, T02_03, T07_09, T02_01, T02_04, T07_03, T02_05, T07_07, T07_10, T09_11, T01_05, T03_07, T03_01, T03_08, T03_05, T03_06, T03_03, T10_08, T11_06, T11_03, T11_01, T04_02, T04_01, T04_06, T04_04, T11_04, T10_01, T11_08, T10_02, T10_09, T10_07, T11_02, T04_05, T04_03, T11_05, T10_06, T11_10, T11_09, T04_07, T10_10, T04_09, T04_08, T11_0X, T06_05, T05_03, T05_06, T05_05, T06_02, T06_04, T06_03, T05_04, T05_02, T05_01, T06_01, T08_04, T08_06, T08_03, T08_07, T08_02, T08_05, T08_01, T08_09, T08_08, T03_09, T06_1X, T26_03, T26_05, T26_07, T26_09, T26_06, T26_10, T26_08, T26_02, T26_04, T27_01, T27_05, T27_07, T27_06, T27_04, T27_02, T27_03, T24_05, T24_08, T24_02, T24_10, T24_06, T24_24, T24_25, T24_09, T24_21, T24_23, T23_10, T23_13, T23_09, T23_15, T23_03, T23_04, T23_14, T23_07, T23_08, T23_02, T23_06, T23_05, T25_05, T25_22, T22_03, T25_06, T22_12, T25_09, T25_10, T25_08, T22_11, T22_02, T25_11, T25_07, T25_14, T25_12, T25_04, T22_08, T25_23, T22_10, T22_06, T25_13, T22_01
#> 1 subject(s) solved each item: P04491 (17 correct), P04491 (17 correct) ...
#> Dataset is completely linked.
#> 'gauss' has been chosen for estimation method. Number of nodes was not explicitly specified. Set nodes to 20.
#> Q matrix specifies 1 dimension(s).
#>
#>
#> ================================
#> Model No. 3
#> Model name: global
#> Number of items: 33
#> Number of persons: 133
#> Number of dimensions: 1
#> ================================
#>
#> Following 250 item(s) missed in data frame will be removed from Q matrix:
#> T12_02, T07_08, T02_02, T07_01, T02_06, T07_05, T07_06, T02_07, T07_02, T07_04, T02_03, T07_09, T02_01, T02_04, T07_03, T02_05, T07_07, T07_10, T09_11, T01_05, T13_15, T15_03, T13_04, T13_07, T13_16, T13_01, T15_02, T15_01, T15_15, T15_16, T15_09, T13_12, T13_13, T15_14, T13_08, T13_06, T15_12, T13_10, T13_17, T15_11, T13_05, T13_09, T15_10, T15_13, T03_07, T03_01, T03_08, T03_05, T03_06, T03_03, T10_08, T11_06, T11_03, T11_01, T04_02, T04_01, T04_06, T04_04, T11_04, T10_01, T11_08, T10_02, T10_09, T10_07, T11_02, T04_05, T04_03, T11_05, T10_06, T11_10, T11_09, T04_07, T15_17, T15_18, T13_18, T10_10, T04_09, T04_08, T11_0X, T16_08, T16_09, T16_11, T16_13, T16_01, T16_04, T16_10, T16_02, T16_12, T16_06, T16_07, T06_05, T05_03, T05_06, T05_05, T06_02, T06_04, T06_03, T05_04, T05_02, T05_01, T06_01, T08_04, T08_06, T08_03, T08_07, T08_02, T08_05, T08_01, T08_09, T08_08, T03_09, T06_1X, T19_06, T19_08, T19_05, T19_11, T19_01, T19_02, T19_30, T19_03, T19_07, T19_04, T19_09, T26_03, T26_05, T26_07, T26_09, T26_06, T26_10, T26_08, T26_02, T26_04, T21_11, T18_01, T21_01, T18_05, T21_12, T18_13, T21_10, T18_11, T21_05, T21_13, T18_08, T18_07, T21_09, T18_04, T18_02, T21_02, T21_06, T18_09, T21_04, T18_12, T21_14, T21_03, T18_10, T27_01, T27_05, T27_07, T27_06, T27_04, T27_02, T27_03, T30_06, T29_04, T29_02, T30_02, T30_03, T30_01, T29_05, T29_03, T24_05, T24_08, T24_02, T24_10, T24_06, T24_24, T24_25, T24_09, T24_21, T24_23, T28_08, T28_09, T28_02, T28_03, T28_04, T28_01, T28_10, T28_06, T23_10, T23_13, T23_09, T23_15, T23_03, T23_04, T23_14, T23_07, T23_08, T23_02, T23_06, T23_05, T25_05, T25_22, T22_03, T25_06, T22_12, T25_09, T25_10, T25_08, T22_11, T22_02, T25_11, T25_07, T25_14, T25_12, T25_04, T22_08, T25_23, T22_10, T22_06, T25_13, T22_01, T20_05, T20_09, T20_30, T20_08, T20_12, T20_06, T20_07, T20_02, T20_13, T20_03, T20_01, T20_04, T20_11, T20_10, T32_04, T31_03, T31_01, T31_04, T32_03, T32_02, T32_01, T17_02, T17_09, T17_10, T17_04, T17_03, T17_08, T17_06, T17_01
#> Dataset is completely linked.
#> 'gauss' has been chosen for estimation method. Number of nodes was not explicitly specified. Set nodes to 20.
#> Q matrix specifies 1 dimension(s).
# run 3 models
runs5 <- runModel(mods5)
# get the results
res5 <- getResults(runs5)
#> |*****|
#> |-----|
#> |*****|
#> |-----|
#> |*****|
#> |-----|
# only for illustration, we create arbitrary 'normed' parameters for anchoring.
# Each item now has two item parameter: one is domain-specific, one is the global
# item parameter. Hence, each item occurs two times in 'prmNrm'
prmNrm<- do.call("rbind", by(data = itemFromRes(ress),
INDICES = itemFromRes(ress)[,"dimension"], FUN = function (d) {
d <- d[sample(1:nrow(d), size = round(nrow(d)/3), replace=FALSE), c("dimension", "item", "est")]
const<- ifelse(d[1,"dimension"] == "domainlistening", 0.6, 0.4)
err <- ifelse(d[1,"dimension"] == "domainlistening", 0.75, 0.6)
d[,"est"] <- d[,"est"] - const + rnorm ( nrow(d), 0, err)
return(d) }))
prmGlo<- prmNrm |> dplyr::mutate(dimension = "global")
prmNrm<- rbind ( prmNrm, prmGlo)
# if the item identifier in 'prmNrm' is not unique, 'equat1pl' has to know which
# parameter in 'prmNrm' belongs to which dimension/domain. Hence, the dimension
# is added to 'prmNrm'.
# anchoring: if 'prmNrm' has more than 2 columns, the columns of 'prmNrm' must be
# specified in 'equat1pl'
anch5 <- equat1pl ( results = res5, prmNorm = prmNrm, item = "item", value = "est",
domain = "dimension", excludeLinkingDif = FALSE, difBound = 0.64)
#> Found 3 model(s).
#> Equating is executed for each dimension in each model separately.
#>
#> ====================================================================================================
#>
#> Model No. 1
#> Model name: domainlistening
#> Number of dimension(s): 1
#> Name(s) of dimension(s): domainlistening
#> Number of linking items: 5
#> Linking method: Mean.Mean
#>
#> Dimension 'domainlistening': 4 of 5 items with linking |DIF| > 0.64 identified.
#>
#> item dif linking.constant linkerror
#> 1 T12_03 -1.250 -0.908 0.463
#> 2 T12_06 1.030 -0.908 0.463
#> 3 T12_09 -0.649 -0.908 0.463
#> 4 T12_10 1.074 -0.908 0.463
#>
#>
#> ====================================================================================================
#>
#> Model No. 2
#> Model name: domainreading
#> Number of dimension(s): 1
#> Name(s) of dimension(s): domainreading
#> Number of linking items: 6
#> Linking method: Mean.Mean
#>
#> Dimension 'domainreading': 1 of 6 items with linking |DIF| > 0.64 identified.
#>
#> item dif linking.constant linkerror
#> 1 T01_06 -0.83 -0.278 0.188
#>
#>
#> ====================================================================================================
#>
#> Model No. 3
#> Model name: global
#> Number of dimension(s): 1
#> Name(s) of dimension(s): global
#> Number of linking items: 11
#> Linking method: Mean.Mean
#>
#> Dimension 'global': 4 of 11 items with linking |DIF| > 0.64 identified.
#>
#> item dif linking.constant linkerror
#> 1 T01_06 -1.153 -0.593 0.241
#> 2 T12_03 -0.893 -0.593 0.241
#> 3 T12_06 1.332 -0.593 0.241
#> 4 T12_10 1.409 -0.593 0.241
#>
# for illustration: consequences of linking the global domain if linking constants
# differ between domains
anch5a<- equat1pl ( results = res5, prmNorm = prmNrm, item = "item", value = "est",
domain = "dimension", excludeLinkingDif = TRUE, iterativ = TRUE, difBound = 0.64)
#> Found 3 model(s).
#> Equating is executed for each dimension in each model separately.
#>
#> ====================================================================================================
#>
#> Model No. 1
#> Model name: domainlistening
#> Number of dimension(s): 1
#> Name(s) of dimension(s): domainlistening
#> Number of linking items: 5
#> Linking method: Mean.Mean
#>
#> Dimension 'domainlistening': 4 of 5 items with linking |DIF| > 0.64 identified.
#> Iteration 1: Exclude item 'T12_03'.
#> Iteration 2: Exclude item 'T12_09'.
#> Iteration 3: Exclude item 'T12_05'.
#>
#> method iter itemExcluded DIF.excluded linking.constant linkerror
#> 1 iterativ 0 -0.908 0.463
#> 2 iterativ 1 T12_03 -1.25 -1.216 0.446
#> 3 iterativ 2 T12_09 -1.003 -1.529 0.449
#> 4 iterativ 3 T12_05 -1.04 -1.978 0.022
#>
#>
#> ====================================================================================================
#>
#> Model No. 2
#> Model name: domainreading
#> Number of dimension(s): 1
#> Name(s) of dimension(s): domainreading
#> Number of linking items: 6
#> Linking method: Mean.Mean
#>
#> Dimension 'domainreading': 1 of 6 items with linking |DIF| > 0.64 identified.
#> Iteration 1: Exclude item 'T01_06'.
#>
#> method iter itemExcluded DIF.excluded linking.constant linkerror
#> 1 iterativ 0 -0.278 0.188
#> 2 iterativ 1 T01_06 -0.83 -0.438 0.119
#>
#>
#> ====================================================================================================
#>
#> Model No. 3
#> Model name: global
#> Number of dimension(s): 1
#> Name(s) of dimension(s): global
#> Number of linking items: 11
#> Linking method: Mean.Mean
#>
#> Dimension 'global': 4 of 11 items with linking |DIF| > 0.64 identified.
#> Iteration 1: Exclude item 'T12_10'.
#> Iteration 2: Exclude item 'T12_06'.
#> Iteration 3: Exclude item 'T01_06'.
#>
#> method iter itemExcluded DIF.excluded linking.constant linkerror
#> 1 iterativ 0 -0.593 0.241
#> 2 iterativ 1 T12_10 1.409 -0.449 0.213
#> 3 iterativ 2 T12_06 1.489 -0.282 0.148
#> 4 iterativ 3 T01_06 -0.808 -0.384 0.123
#>
# transformation to the Bista metric
# first we arbitrarily define mean and standard deviation of the reference
# population according to the three dimensions (defined in the Q matrix):
# reading, listening, and global
# Note that the the first column of the 'refPop' data frame must include the
# domain names. Domain names must match the names defined in the Q matrix
refPop<- data.frame ( domain = c("domainreading", "domainlistening", "global"),
m = c(0.03890191, 0.03587727, 0.03), sd= c(1.219, 0.8978912, 1.05))
# second, we specify a list with cut scores. Values must be in ascending order.
# Labels of the competence stages are optional. If no labels are specified,
# the will be defaulted to 1, 2, 3 ... etc.
# Note: if labels are specified, there must be one label more than cut scores.
# (i.e. 4 cut scores need 5 labels, etc.)
cuts <- list ( domainreading = list ( values = 390+0:3*75),
domainlistening = list ( values = 360+0:3*85),
global = list ( values = 400+0:2*100, labels = c("A1", "A2", "B1", "B2")))
# transformation
dfr <- transformToBista(equatingList = anch5, refPop = refPop, cuts=cuts )
#> The 'refPop' data.frame does not include information about reference population mean/SD on Bista metric. Values will be defaulted to mean = 500 and SD = 100.
#> Registered S3 method overwritten by 'lme4':
#> method from
#> na.action.merMod car
#> Warning: Model 'domainlistening', dimension 'domainlistening': No plausible values on trait level(s) '5'.
#> 1 of 5 trait levels(s) of merging variable 'traitLevel' from data set 'linking error list' not included in data set 'plausible values'.
#> Found duplicated entries in 'item-ID' column. This should only occur for subject 'math' in grade 3.
#> Cannot find 'global' entry in the 'domain' column. Cancel reshaping.
head(dfr$itempars)
#> model item infit outfit itemDiscrim Nvalid est se itemP dimension estTransf estTransf625 estTransfBista traitLevel linkingConstant linkingMethod nLinkitems linkingError linkingErrorTransfBista linkingErrorTraitLevel refMean refSD refTransfMean refTransfSD
#> 1 domainlistening T12_01 1.0463783 1.2299030 0.02099527 133 -3.015202 0.3725945 0.9398496 domainlistening -3.923596 -3.412770 115.91699 1 -0.9083935 Mean.Mean 5 0.4628182 51.54502 0.1611926 0.03587727 0.8978912 500 100
#> 2 domainlistening T12_03 0.9182225 0.7734278 0.37141881 133 -2.290056 0.2836654 0.8872180 domainlistening -3.198449 -2.687623 196.67809 1 -0.9083935 Mean.Mean 5 0.4628182 51.54502 0.1611926 0.03587727 0.8978912 500 100
#> 3 domainlistening T12_04 0.9827313 0.9135211 0.23727591 133 -2.211710 0.2762515 0.8796992 domainlistening -3.120104 -2.609278 205.40357 1 -0.9083935 Mean.Mean 5 0.4628182 51.54502 0.1611926 0.03587727 0.8978912 500 100
#> 4 domainlistening T12_05 1.0000024 0.7405255 0.10028410 133 -4.485206 0.7170350 0.9849624 domainlistening -5.393599 -4.882774 -47.80032 1 -0.9083935 Mean.Mean 5 0.4628182 51.54502 0.1611926 0.03587727 0.8978912 500 100
#> 5 domainlistening T12_10 0.9238162 0.8834016 0.38203159 133 -1.247520 0.2119197 0.7518797 domainlistening -2.155913 -1.645088 312.78747 1 -0.9083935 Mean.Mean 5 0.4628182 51.54502 0.1611926 0.03587727 0.8978912 500 100
#> 6 domainlistening T12_07 0.9202637 0.8904835 0.33255757 133 -2.066257 0.2634642 0.8646617 domainlistening -2.974651 -2.463825 221.60300 1 -0.9083935 Mean.Mean 5 0.4628182 51.54502 0.1611926 0.03587727 0.8978912 500 100
head(dfr$personpars)
#> model idstud group imp value valueTransfBista traitLevel dimension linkingError linkingErrorTransfBista linkingErrorTraitLevel
#> 1 domainlistening P04477 domainlistening pv1 -0.6731876 319.8603 1 domainlistening 0.4628182 51.54502 0.1611926
#> 2 domainlistening P08343 domainlistening pv1 -1.4303260 235.5362 1 domainlistening 0.4628182 51.54502 0.1611926
#> 3 domainlistening P08344 domainlistening pv1 -0.9794447 285.7518 1 domainlistening 0.4628182 51.54502 0.1611926
#> 4 domainlistening P04480 domainlistening pv1 -0.9794447 285.7518 1 domainlistening 0.4628182 51.54502 0.1611926
#> 5 domainlistening P08346 domainlistening pv1 -1.0598387 276.7982 1 domainlistening 0.4628182 51.54502 0.1611926
#> 6 domainlistening P04482 domainlistening pv1 -0.5975374 328.2856 1 domainlistening 0.4628182 51.54502 0.1611926
################################################################################
### Example 6: Scaling, linking and equating for multiple models (II) ###
################################################################################
# Example 6 and 6a: define und run multiple models according to different domains
# (item groups) and different person groups. This example mimics the routines
# necessary for the 'Laendervergleich/Bildungstrend' at the Institute for
# Educational Progress (IQB). Example 6 demonstrates routines without trend
# estimation, i.e. for the first (or solely) measurement occasion. Example 6a
# demonstrates routines for the second measurement occasion (t2) which is linked
# to t1. Example 6b demonstrates routines for the third measurement occasion (t3)
# which is linked to the common scale of t1 and t2.
# Preparation: assume time of measurement 't1' corresponds to the year 2010.
# This is the year of the reference population. We consider both domains,
# reading and listening in a single call. Hence, the data.frame 'datT1' contains
# items of both domains. We add weights to the data as the definition of mean
# and sd of the reference population (population of 't1') necessitates weights.
datT1<- reshape2::dcast(subset ( trends, year == 2010),
idstud+country+sex+ses+language+wgt~item, value.var="value")
# generate Q matrix
qMat <- unique(trends[ ,c("item","domain")])
qMat <- data.frame ( qMat[,"item", drop=FALSE], model.matrix(~domain-1, data = qMat))
# First step: item calibration in separate unidimensional models for each domain
# (the models are short and simple, so we don't need multicore)
modsT1<- splitModels ( qMatrix = qMat, nCores = 1)
#> --------------------------------
#> splitModels: generating 2 models
#> ..
#> see <returned>$models
#> number of cores: 1
#> --------------------------------
# define 2 models. Note: not all items of the Q matrix are present in the data.
# Items which occur only in the Q matrix will be ignored.
defT1 <- defineModel(dat = datT1, id = "idstud", check.for.linking = TRUE,
splittedModels = modsT1, software = "tam")
#>
#> Specification of 'qMatrix' and 'person.groups' results in 2 model(s).
#> Warning: Model split preparation for model No. 1, model name domainlistening: 83 items from 134 items listed the Q matrix not found in data:
#> ℹ 'T12_11', 'T15_17', 'T15_18', 'T13_18', 'T19_06', 'T19_08', 'T19_05', 'T19_11', 'T19_01', 'T19_02', 'T19_30', 'T19_03', 'T19_07', 'T19_04', 'T19_09', 'T21_11', 'T18_01', 'T21_01', 'T18_05', 'T21_12', 'T18_13', 'T21_10', 'T18_11', 'T21_05', 'T21_13', 'T18_08', 'T18_07', 'T21_09', 'T18_04', 'T18_02', 'T21_02', 'T21_06', 'T18_09', 'T21_04', 'T18_12', 'T21_14', 'T21_03', 'T18_10', 'T30_06', 'T29_04', 'T29_02', 'T30_02', 'T30_03', 'T30_01', 'T29_05', 'T29_03', 'T28_08', 'T28_09', 'T28_02', 'T28_03', 'T28_04', 'T28_01', 'T28_10', 'T28_06', 'T20_05', 'T20_09', 'T20_30', 'T20_08', 'T20_12', 'T20_06', 'T20_07', 'T20_02', 'T20_13', 'T20_03', 'T20_01', 'T20_04', 'T20_11', 'T20_10', 'T32_04', 'T31_03', 'T31_01', 'T31_04', 'T32_03', 'T32_02', 'T32_01', 'T17_02', 'T17_09', 'T17_10', 'T17_04', 'T17_03', 'T17_08', 'T17_06', 'T17_01'
#> Warning: Model split preparation for model No. 2, model name domainreading: 69 items from 149 items listed the Q matrix not found in data:
#> ℹ 'T09_12', 'T01_08', 'T10_10', 'T04_09', 'T04_08', 'T11_0X', 'T08_09', 'T08_08', 'T03_09', 'T06_1X', 'T26_03', 'T26_05', 'T26_07', 'T26_09', 'T26_06', 'T26_10', 'T26_08', 'T26_02', 'T26_04', 'T27_01', 'T27_05', 'T27_07', 'T27_06', 'T27_04', 'T27_02', 'T27_03', 'T24_05', 'T24_08', 'T24_02', 'T24_10', 'T24_06', 'T24_24', 'T24_25', 'T24_09', 'T24_21', 'T24_23', 'T23_10', 'T23_13', 'T23_09', 'T23_15', 'T23_03', 'T23_04', 'T23_14', 'T23_07', 'T23_08', 'T23_02', 'T23_06', 'T23_05', 'T25_05', 'T25_22', 'T22_03', 'T25_06', 'T22_12', 'T25_09', 'T25_10', 'T25_08', 'T22_11', 'T22_02', 'T25_11', 'T25_07', 'T25_14', 'T25_12', 'T25_04', 'T22_08', 'T25_23', 'T22_10', 'T22_06', 'T25_13', 'T22_01'
#>
#>
#> =========================================
#> Model No. 1
#> Model name: domainlistening
#> Number of items: 51
#> Number of persons: 4476
#> Number of dimensions: 1
#> =========================================
#>
#> Following 83 item(s) missed in data frame will be removed from Q matrix:
#> T12_11, T15_17, T15_18, T13_18, T19_06, T19_08, T19_05, T19_11, T19_01, T19_02, T19_30, T19_03, T19_07, T19_04, T19_09, T21_11, T18_01, T21_01, T18_05, T21_12, T18_13, T21_10, T18_11, T21_05, T21_13, T18_08, T18_07, T21_09, T18_04, T18_02, T21_02, T21_06, T18_09, T21_04, T18_12, T21_14, T21_03, T18_10, T30_06, T29_04, T29_02, T30_02, T30_03, T30_01, T29_05, T29_03, T28_08, T28_09, T28_02, T28_03, T28_04, T28_01, T28_10, T28_06, T20_05, T20_09, T20_30, T20_08, T20_12, T20_06, T20_07, T20_02, T20_13, T20_03, T20_01, T20_04, T20_11, T20_10, T32_04, T31_03, T31_01, T31_04, T32_03, T32_02, T32_01, T17_02, T17_09, T17_10, T17_04, T17_03, T17_08, T17_06, T17_01
#> 76 subject(s) do not solve any item:
#> P00009 (6 false), P00864 (6 false), P02759 (21 false) ...
#> 55 subject(s) solved each item: P00099 (10 correct), P00404 (10 correct), P02659 (11 correct) ...
#> Dataset is completely linked.
#> 'gauss' has been chosen for estimation method. Number of nodes was not explicitly specified. Set nodes to 20.
#> Q matrix specifies 1 dimension(s).
#>
#>
#> =======================================
#> Model No. 2
#> Model name: domainreading
#> Number of items: 80
#> Number of persons: 4476
#> Number of dimensions: 1
#> =======================================
#>
#> Following 69 item(s) missed in data frame will be removed from Q matrix:
#> T09_12, T01_08, T10_10, T04_09, T04_08, T11_0X, T08_09, T08_08, T03_09, T06_1X, T26_03, T26_05, T26_07, T26_09, T26_06, T26_10, T26_08, T26_02, T26_04, T27_01, T27_05, T27_07, T27_06, T27_04, T27_02, T27_03, T24_05, T24_08, T24_02, T24_10, T24_06, T24_24, T24_25, T24_09, T24_21, T24_23, T23_10, T23_13, T23_09, T23_15, T23_03, T23_04, T23_14, T23_07, T23_08, T23_02, T23_06, T23_05, T25_05, T25_22, T22_03, T25_06, T22_12, T25_09, T25_10, T25_08, T22_11, T22_02, T25_11, T25_07, T25_14, T25_12, T25_04, T22_08, T25_23, T22_10, T22_06, T25_13, T22_01
#> 17 subject(s) do not solve any item:
#> P00173 (6 false), P01137 (7 false), P02863 (23 false) ...
#> 97 subject(s) solved each item: P00078 (6 correct), P00389 (10 correct), P03338 (27 correct) ...
#> Dataset is completely linked.
#> 'gauss' has been chosen for estimation method. Number of nodes was not explicitly specified. Set nodes to 20.
#> Q matrix specifies 1 dimension(s).
# run (calibrate) the 2 models subsequently
runT1 <- runModel(defT1)
# get the results of the two unidimensional models
resT1 <- getResults(runT1, omitWle = TRUE, Q3 = FALSE)
#> Getting standard errors with the tam.se function: 0.4 secs
#> |*****|
#> |-----|
#> Getting PVs calling tam.pv from getTamPVs: 1.5 secs
#> Getting standard errors with the tam.se function: 0.8 secs
#> |*****|
#> |-----|
#> Getting PVs calling tam.pv from getTamPVs: 1.1 secs
# extract item parameters from the 'results' object
# t1 is the reference measurement occasion, i.e. no linking/equating is necessary
itemT1<- itemFromRes(resT1)
# The measurement point 't1' defines the reference population. We need to specify
# mean and SD, using weights.
eqRef <- equat1pl(resT1)
#> Found 2 model(s).
#> Equating is executed for each dimension in each model separately.
#> No norm parameter defined ('prmNorm' is missing). Treat current sample as drawn from the reference population.
# transformation to the 'bista' metric
# Note: if the sample was drawn from the reference population in a weighted sample,
# mean and SD are not yet known. So we ignore the 'refPop' argument in 'transformToBista'
# and simply define the cut scores. The function then assumes that the parameter
# stem from the reference population and estimates its mean and sd.
cuts <- list ( domainreading = list ( values = 390+0:3*75),
domainlistening = list ( values = 360+0:3*85))
tfRef <- transformToBista ( equatingList = eqRef, cuts=cuts, vera=FALSE,
weights = datT1[,c("idstud", "wgt")] )
#> 'refPop' was not defined. Treat current sample as drawn from the reference population.
#> The 'refPop' data.frame does not include information about reference population mean/SD on Bista metric. Values will be defaulted to mean = 500 and SD = 100.
# Second step: drawing plausible values separately for each country.
# A two-dimensional (reading/listening) model is specified separately for each
# person group (= each country) with item parameters fixed at their calibration
# values. Moreover, a latent regression model is used (in the actual 'Laendervergleich',
# regressors are principal components of previously imputed background variables). We use 'sex',
# 'ses' and 'language' as regressors. For convenience, 'ses' is scaled (mean = 0, sd = 1)
datT1[,"ses_scaled"] <- scale(datT1[,"ses"])[,1]
# have a look at 'sex' and 'language at home' in each country:
table(datT1[,c("country", "sex")])
#> sex
#> country female male
#> countryA 787 811
#> countryB 663 646
#> countryC 795 774
table(datT1[,c("country", "language")])
#> language
#> country native nativeAndOther other
#> countryA 1143 437 18
#> countryB 1218 81 10
#> countryC 1248 308 13
# Running second step: split models according to person groups
# ('all.persons' must be FALSE, otherwise the whole group would be treated as
# a separate distinct group.)
modT1P<- splitModels ( person.groups = datT1[,c("idstud", "country")],
all.persons = FALSE, nCores = 1)
#> --------------------------------
#> splitModels: generating 3 models
#> ...
#> see <returned>$models
#> number of cores: 1
#> --------------------------------
# define the 3 country-specific 2-dimensional models, specifying latent regression
# model and fixed item parameters.
defT1P<- defineModel(dat = datT1, items = itemT1[,"item"], id = "idstud",
check.for.linking = TRUE, splittedModels = modT1P, qMatrix = qMat,
anchor = itemT1[,c("item", "est")],
HG.var = c("sex", "ses_scaled", "language"), software = "tam")
#>
#> Specification of 'qMatrix' and 'person.groups' results in 3 model(s).
#>
#>
#> ==========================================
#> Model No. 1
#> Model name: country.countryC
#> Number of items: 131
#> Number of persons: 1569
#> Number of dimensions: 2
#> ==========================================
#>
#> Following 152 item(s) missed in data frame will be removed from Q matrix:
#> T12_11, T09_12, T01_08, T15_17, T15_18, T13_18, T10_10, T04_09, T04_08, T11_0X, T08_09, T08_08, T03_09, T06_1X, T19_06, T19_08, T19_05, T19_11, T19_01, T19_02, T19_30, T19_03, T19_07, T19_04, T19_09, T26_03, T26_05, T26_07, T26_09, T26_06, T26_10, T26_08, T26_02, T26_04, T21_11, T18_01, T21_01, T18_05, T21_12, T18_13, T21_10, T18_11, T21_05, T21_13, T18_08, T18_07, T21_09, T18_04, T18_02, T21_02, T21_06, T18_09, T21_04, T18_12, T21_14, T21_03, T18_10, T27_01, T27_05, T27_07, T27_06, T27_04, T27_02, T27_03, T30_06, T29_04, T29_02, T30_02, T30_03, T30_01, T29_05, T29_03, T24_05, T24_08, T24_02, T24_10, T24_06, T24_24, T24_25, T24_09, T24_21, T24_23, T28_08, T28_09, T28_02, T28_03, T28_04, T28_01, T28_10, T28_06, T23_10, T23_13, T23_09, T23_15, T23_03, T23_04, T23_14, T23_07, T23_08, T23_02, T23_06, T23_05, T25_05, T25_22, T22_03, T25_06, T22_12, T25_09, T25_10, T25_08, T22_11, T22_02, T25_11, T25_07, T25_14, T25_12, T25_04, T22_08, T25_23, T22_10, T22_06, T25_13, T22_01, T20_05, T20_09, T20_30, T20_08, T20_12, T20_06, T20_07, T20_02, T20_13, T20_03, T20_01, T20_04, T20_11, T20_10, T32_04, T31_03, T31_01, T31_04, T32_03, T32_02, T32_01, T17_02, T17_09, T17_10, T17_04, T17_03, T17_08, T17_06, T17_01
#> Background variable(s) 'sex', 'language' of class
#> 'factor', 'factor' will be converted to indicator variables.
#> Variable 'sex' was converted to 1 indicator(s) with name(s) 'sexmale'.
#> Variable 'language' was converted to 2 indicator(s) with name(s) 'languagenativeAndOther', 'languageother'.
#> 5 subject(s) solved each item: P00106 (17 correct), P00939 (17 correct), P00393 (20 correct) ...
#> Dataset is completely linked.
#> 'gauss' has been chosen for estimation method. Number of nodes was not explicitly specified. Set nodes to 20.
#> 131 common items found in 'anchor' list and data frame.
#> Q matrix specifies 2 dimension(s).
#>
#>
#> ==========================================
#> Model No. 2
#> Model name: country.countryA
#> Number of items: 131
#> Number of persons: 1598
#> Number of dimensions: 2
#> ==========================================
#>
#> Following 152 item(s) missed in data frame will be removed from Q matrix:
#> T12_11, T09_12, T01_08, T15_17, T15_18, T13_18, T10_10, T04_09, T04_08, T11_0X, T08_09, T08_08, T03_09, T06_1X, T19_06, T19_08, T19_05, T19_11, T19_01, T19_02, T19_30, T19_03, T19_07, T19_04, T19_09, T26_03, T26_05, T26_07, T26_09, T26_06, T26_10, T26_08, T26_02, T26_04, T21_11, T18_01, T21_01, T18_05, T21_12, T18_13, T21_10, T18_11, T21_05, T21_13, T18_08, T18_07, T21_09, T18_04, T18_02, T21_02, T21_06, T18_09, T21_04, T18_12, T21_14, T21_03, T18_10, T27_01, T27_05, T27_07, T27_06, T27_04, T27_02, T27_03, T30_06, T29_04, T29_02, T30_02, T30_03, T30_01, T29_05, T29_03, T24_05, T24_08, T24_02, T24_10, T24_06, T24_24, T24_25, T24_09, T24_21, T24_23, T28_08, T28_09, T28_02, T28_03, T28_04, T28_01, T28_10, T28_06, T23_10, T23_13, T23_09, T23_15, T23_03, T23_04, T23_14, T23_07, T23_08, T23_02, T23_06, T23_05, T25_05, T25_22, T22_03, T25_06, T22_12, T25_09, T25_10, T25_08, T22_11, T22_02, T25_11, T25_07, T25_14, T25_12, T25_04, T22_08, T25_23, T22_10, T22_06, T25_13, T22_01, T20_05, T20_09, T20_30, T20_08, T20_12, T20_06, T20_07, T20_02, T20_13, T20_03, T20_01, T20_04, T20_11, T20_10, T32_04, T31_03, T31_01, T31_04, T32_03, T32_02, T32_01, T17_02, T17_09, T17_10, T17_04, T17_03, T17_08, T17_06, T17_01
#> Background variable(s) 'sex', 'language' of class
#> 'factor', 'factor' will be converted to indicator variables.
#> Variable 'sex' was converted to 1 indicator(s) with name(s) 'sexmale'.
#> Variable 'language' was converted to 2 indicator(s) with name(s) 'languagenativeAndOther', 'languageother'.
#> Dataset is completely linked.
#> 'gauss' has been chosen for estimation method. Number of nodes was not explicitly specified. Set nodes to 20.
#> 131 common items found in 'anchor' list and data frame.
#> Q matrix specifies 2 dimension(s).
#>
#>
#> ==========================================
#> Model No. 3
#> Model name: country.countryB
#> Number of items: 131
#> Number of persons: 1309
#> Number of dimensions: 2
#> ==========================================
#>
#> Following 152 item(s) missed in data frame will be removed from Q matrix:
#> T12_11, T09_12, T01_08, T15_17, T15_18, T13_18, T10_10, T04_09, T04_08, T11_0X, T08_09, T08_08, T03_09, T06_1X, T19_06, T19_08, T19_05, T19_11, T19_01, T19_02, T19_30, T19_03, T19_07, T19_04, T19_09, T26_03, T26_05, T26_07, T26_09, T26_06, T26_10, T26_08, T26_02, T26_04, T21_11, T18_01, T21_01, T18_05, T21_12, T18_13, T21_10, T18_11, T21_05, T21_13, T18_08, T18_07, T21_09, T18_04, T18_02, T21_02, T21_06, T18_09, T21_04, T18_12, T21_14, T21_03, T18_10, T27_01, T27_05, T27_07, T27_06, T27_04, T27_02, T27_03, T30_06, T29_04, T29_02, T30_02, T30_03, T30_01, T29_05, T29_03, T24_05, T24_08, T24_02, T24_10, T24_06, T24_24, T24_25, T24_09, T24_21, T24_23, T28_08, T28_09, T28_02, T28_03, T28_04, T28_01, T28_10, T28_06, T23_10, T23_13, T23_09, T23_15, T23_03, T23_04, T23_14, T23_07, T23_08, T23_02, T23_06, T23_05, T25_05, T25_22, T22_03, T25_06, T22_12, T25_09, T25_10, T25_08, T22_11, T22_02, T25_11, T25_07, T25_14, T25_12, T25_04, T22_08, T25_23, T22_10, T22_06, T25_13, T22_01, T20_05, T20_09, T20_30, T20_08, T20_12, T20_06, T20_07, T20_02, T20_13, T20_03, T20_01, T20_04, T20_11, T20_10, T32_04, T31_03, T31_01, T31_04, T32_03, T32_02, T32_01, T17_02, T17_09, T17_10, T17_04, T17_03, T17_08, T17_06, T17_01
#> Background variable(s) 'sex', 'language' of class
#> 'factor', 'factor' will be converted to indicator variables.
#> Variable 'sex' was converted to 1 indicator(s) with name(s) 'sexmale'.
#> Variable 'language' was converted to 2 indicator(s) with name(s) 'languagenativeAndOther', 'languageother'.
#> Dataset is completely linked.
#> 'gauss' has been chosen for estimation method. Number of nodes was not explicitly specified. Set nodes to 20.
#> 131 common items found in 'anchor' list and data frame.
#> Q matrix specifies 2 dimension(s).
# run the 3 models (estimation needs approx. 20 seconds)
runT1P<- runModel(defT1P)
# get the results (to save time, item fit estimation is skipped)
resT1P<- getResults(runT1P, omitWle = TRUE, Q3 = FALSE)
#> Getting standard errors with the tam.se function: 12.6 secs
#> Getting infit parameters calling tam.fit from getTamInfit: 2.9 secs
#> |*****|
#> |-----|
#> Getting PVs calling tam.pv from getTamPVs: 0.5 secs
#> Getting standard errors with the tam.se function: 15.3 secs
#> Getting infit parameters calling tam.fit from getTamInfit: 3 secs
#> |*****|
#> |-----|
#> Getting PVs calling tam.pv from getTamPVs: 0.5 secs
#> Getting Q3 statistic calling tam.modelfit from getTamQ3: 0.3 secs
#> Getting standard errors with the tam.se function: 12.3 secs
#> Getting infit parameters calling tam.fit from getTamInfit: 2.2 secs
#> |*****|
#> |-----|
#> Getting PVs calling tam.pv from getTamPVs: 0.4 secs
# latent regression coefficients for the three countries and two dimensions
regcoefFromRes(resT1P, digits = 3)
#> $`model: 'country.countryA', group: 'domainlistening'`
#> parameter est se p sig
#> 1 (Intercept) -0.011 0.021 0.577
#> 2 languagenativeAndOther -0.353 0.039 0.000 ***
#> 3 languageother -0.608 0.195 0.002 **
#> 4 ses_scaled 0.377 0.020 0.000 ***
#> 5 sexmale -0.009 0.029 0.755
#>
#> $`model: 'country.countryB', group: 'domainlistening'`
#> parameter est se p sig
#> 1 (Intercept) 0.075 0.024 0.002 **
#> 2 languagenativeAndOther -0.343 0.095 0.000 ***
#> 3 languageother -0.285 0.271 0.292
#> 4 ses_scaled 0.275 0.026 0.000 ***
#> 5 sexmale 0.079 0.034 0.021 *
#>
#> $`model: 'country.countryC', group: 'domainlistening'`
#> parameter est se p sig
#> 1 (Intercept) 0.133 0.023 0.000 ***
#> 2 languagenativeAndOther -0.381 0.053 0.000 ***
#> 3 languageother -0.007 0.281 0.982
#> 4 ses_scaled 0.298 0.024 0.000 ***
#> 5 sexmale -0.008 0.033 0.811
#>
#> $`model: 'country.countryA', group: 'domainreading'`
#> parameter est se p sig
#> 1 (Intercept) -0.033 0.023 0.147
#> 2 languagenativeAndOther -0.308 0.043 0.000 ***
#> 3 languageother -0.451 0.214 0.035 *
#> 4 ses_scaled 0.400 0.022 0.000 ***
#> 5 sexmale -0.233 0.032 0.000 ***
#>
#> $`model: 'country.countryB', group: 'domainreading'`
#> parameter est se p sig
#> 1 (Intercept) 0.412 0.027 0.000 ***
#> 2 languagenativeAndOther -0.147 0.106 0.165
#> 3 languageother -0.467 0.296 0.115
#> 4 ses_scaled 0.288 0.029 0.000 ***
#> 5 sexmale -0.194 0.038 0.000 ***
#>
#> $`model: 'country.countryC', group: 'domainreading'`
#> parameter est se p sig
#> 1 (Intercept) 0.167 0.026 0.000 ***
#> 2 languagenativeAndOther -0.298 0.059 0.000 ***
#> 3 languageother -0.505 0.310 0.103
#> 4 ses_scaled 0.384 0.027 0.000 ***
#> 5 sexmale -0.233 0.037 0.000 ***
#>
# equating is not necessary, as the models run with fixed item parameters
# However, to prepare for the transformation on the 'bista' metric, run
# 'equat1pl' with empty arguments
ankT1P<- equat1pl ( results = resT1P)
#> Found 3 model(s).
#> Equating is executed for each dimension in each model separately.
#> No norm parameter defined ('prmNorm' is missing). Treat current sample as drawn from the reference population.
# transformation to the 'bista' metric
# Note: if the sample was drawn from the reference population, mean and SD
# were just computed and captured in 'tfRef'.
dfrT1P<- transformToBista ( equatingList = ankT1P, refPop = tfRef[["refPop"]][,-2], cuts=cuts, vera=FALSE )
################################################################################
### Example 6a: Extend example 6 with trend estimation (now for t2) ###
################################################################################
# Example 6a needs the objects (Q matrix, item parameters, ...) created in example 6
# Preparation: assume time of measurement 't2'.
datT2<- reshape2::dcast(subset ( trends, year == 2015),
idstud+country+sex+ses+language~item, value.var="value")
# First step: item calibration in separate unidimensional models for each domain
modsT2<- splitModels ( qMatrix = qMat, nCores = 1)
#> --------------------------------
#> splitModels: generating 2 models
#> ..
#> see <returned>$models
#> number of cores: 1
#> --------------------------------
# define 2 models. Items which occur only in the Q matrix but not in the data
# will be ignored.
defT2 <- defineModel(dat = datT2, id = "idstud", check.for.linking = TRUE,
splittedModels = modsT2, software = "tam")
#>
#> Specification of 'qMatrix' and 'person.groups' results in 2 model(s).
#> Warning: Model split preparation for model No. 1, model name domainlistening: 38 items from 134 items listed the Q matrix not found in data:
#> ℹ 'T12_02', 'T13_16', 'T15_12', 'T15_11', 'T16_08', 'T16_09', 'T16_11', 'T16_13', 'T16_01', 'T16_04', 'T16_10', 'T16_02', 'T16_12', 'T16_06', 'T16_07', 'T30_06', 'T29_04', 'T29_02', 'T30_02', 'T30_03', 'T30_01', 'T29_05', 'T29_03', 'T28_08', 'T28_09', 'T28_02', 'T28_03', 'T28_04', 'T28_01', 'T28_10', 'T28_06', 'T32_04', 'T31_03', 'T31_01', 'T31_04', 'T32_03', 'T32_02', 'T32_01'
#> Warning: Model split preparation for model No. 2, model name domainreading: 30 items from 149 items listed the Q matrix not found in data:
#> ℹ 'T09_11', 'T01_05', 'T03_07', 'T10_08', 'T11_03', 'T04_01', 'T11_04', 'T04_07', 'T11_0X', 'T06_04', 'T06_03', 'T08_04', 'T08_05', 'T06_1X', 'T26_03', 'T26_05', 'T26_07', 'T26_09', 'T26_06', 'T26_10', 'T26_08', 'T26_02', 'T26_04', 'T27_01', 'T27_05', 'T27_07', 'T27_06', 'T27_04', 'T27_02', 'T27_03'
#>
#>
#> =========================================
#> Model No. 1
#> Model name: domainlistening
#> Number of items: 96
#> Number of persons: 4516
#> Number of dimensions: 1
#> =========================================
#>
#> Following 38 item(s) missed in data frame will be removed from Q matrix:
#> T12_02, T13_16, T15_12, T15_11, T16_08, T16_09, T16_11, T16_13, T16_01, T16_04, T16_10, T16_02, T16_12, T16_06, T16_07, T30_06, T29_04, T29_02, T30_02, T30_03, T30_01, T29_05, T29_03, T28_08, T28_09, T28_02, T28_03, T28_04, T28_01, T28_10, T28_06, T32_04, T31_03, T31_01, T31_04, T32_03, T32_02, T32_01
#> Found 105 cases with missings on all items.
#> Cases with missings on all items will be deleted.
#> 149 subject(s) do not solve any item:
#> P04521 (6 false), P07332 (6 false), P08038 (19 false) ...
#> 77 subject(s) solved each item: P04599 (10 correct), P06913 (10 correct), P06282 (13 correct) ...
#> W A R N I N G ! Dataset is NOT completely linked (even if cases with missings on all items are removed).
#> 501 cases unconnected. Following items are unconnected:
#> T20_05, T20_09, T20_30, T20_08, T20_12, T20_06, T20_07, T20_02, T20_13, T20_03, T20_01, T20_04, T20_11, T20_10
#> 'gauss' has been chosen for estimation method. Number of nodes was not explicitly specified. Set nodes to 20.
#> Q matrix specifies 1 dimension(s).
#>
#>
#> =======================================
#> Model No. 2
#> Model name: domainreading
#> Number of items: 119
#> Number of persons: 4516
#> Number of dimensions: 1
#> =======================================
#>
#> Following 30 item(s) missed in data frame will be removed from Q matrix:
#> T09_11, T01_05, T03_07, T10_08, T11_03, T04_01, T11_04, T04_07, T11_0X, T06_04, T06_03, T08_04, T08_05, T06_1X, T26_03, T26_05, T26_07, T26_09, T26_06, T26_10, T26_08, T26_02, T26_04, T27_01, T27_05, T27_07, T27_06, T27_04, T27_02, T27_03
#> 56 subject(s) do not solve any item:
#> P04511 (6 false), P05616 (10 false), P08143 (14 false) ...
#> 114 subject(s) solved each item: P04509 (6 correct), P08241 (8 correct), P05805 (20 correct) ...
#> W A R N I N G ! Dataset is NOT completely linked (even if cases with missings on all items are removed).
#> 4035 cases unconnected. Following items are unconnected:
#> T02_02, T02_06, T02_07, T02_03, T02_01, T02_04, T02_05, T09_03, T09_05, T01_03, T09_02, T01_06, T01_01, T09_10, T01_02, T09_06, T01_04, T09_09, T09_08, T09_07, T09_04, T01_07, T09_12, T01_08, T03_01, T03_08, T03_05, T03_06, T03_03, T11_06, T11_01, T04_02, T04_06, T04_04, T10_01, T11_08, T10_02, T10_09, T10_07, T11_02, T04_05, T04_03, T11_05, T10_06, T11_10, T11_09, T10_10, T04_09, T04_08, T06_05, T05_03, T05_06, T05_05, T06_02, T05_04, T05_02, T05_01, T06_01, T08_06, T08_03, T08_07, T08_02, T08_01, T08_09, T08_08, T03_09, T24_05, T24_08, T24_02, T24_10, T24_06, T24_24, T24_25, T24_09, T24_21, T24_23, T23_10, T23_13, T23_09, T23_15, T23_03, T23_04, T23_14, T23_07, T23_08, T23_02, T23_06, T23_05, T25_05, T25_22, T22_03, T25_06, T22_12, T25_09, T25_10, T25_08, T22_11, T22_02, T25_11, T25_07, T25_14, T25_12, T25_04, T22_08, T25_23, T22_10, T22_06, T25_13, T22_01
#> 'gauss' has been chosen for estimation method. Number of nodes was not explicitly specified. Set nodes to 20.
#> Q matrix specifies 1 dimension(s).
# run 2 models for calibration
runT2 <- runModel(defT2)
# get the results
resT2 <- getResults(runT2)
#> Getting standard errors with the tam.se function: 1.1 secs
#> Getting infit parameters calling tam.fit from getTamInfit: 0.4 secs
#> Getting WLEs calling tam.wle from getTamWles: 1.6 secs
#> |*****|
#> |-----|
#> Getting PVs calling tam.pv from getTamPVs: 1.6 secs
#> Getting Q3 statistic calling tam.modelfit from getTamQ3: 2.7 secs
#> Getting standard errors with the tam.se function: 1.4 secs
#> Getting infit parameters calling tam.fit from getTamInfit: 0.3 secs
#> Getting WLEs calling tam.wle from getTamWles: 1.2 secs
#> |*****|
#> |-----|
#> Getting PVs calling tam.pv from getTamPVs: 1.1 secs
#> Getting Q3 statistic calling tam.modelfit from getTamQ3: 4.3 secs
# collect item parameters
itemT2<- itemFromRes(resT2)
# Second step: compute linking constant between 't1' and 't2' with the iterative
# exclusion of linking DIF items and computation of linking error. We use the
# 'itemT1' object created in example 6 for reference item parameters. The linking
# procedure is executed consecutively for listening and reading.
L.t1t2<- equat1pl ( results = resT2, prmNorm = itemT1[,c("item", "est")],
excludeLinkingDif = TRUE, difBound = 0.64, iterativ = TRUE)
#> Found 2 model(s).
#> Equating is executed for each dimension in each model separately.
#>
#> ====================================================================================================
#>
#> Model No. 1
#> Model name: domainlistening
#> Number of dimension(s): 1
#> Name(s) of dimension(s): domainlistening
#> Number of linking items: 36
#> Linking method: Mean.Mean
#>
#> Dimension 'domainlistening': 2 of 36 items with linking |DIF| > 0.64 identified.
#> Iteration 1: Exclude item 'T15_02'.
#> Iteration 2: Exclude item 'T14_04'.
#>
#> method iter itemExcluded DIF.excluded linking.constant linkerror
#> 1 iterativ 0 -0.243 0.082
#> 2 iterativ 1 T15_02 -1.873 -0.298 0.063
#> 3 iterativ 2 T14_04 1.675 -0.250 0.042
#>
#>
#> ====================================================================================================
#>
#> Model No. 2
#> Model name: domainreading
#> Number of dimension(s): 1
#> Name(s) of dimension(s): domainreading
#> Number of linking items: 68
#> Linking method: Mean.Mean
#>
#> Dimension 'domainreading': 4 of 68 items with linking |DIF| > 0.64 identified.
#> Iteration 1: Exclude item 'T02_07'.
#> Iteration 2: Exclude item 'T01_07'.
#> Iteration 3: Exclude item 'T02_04'.
#> Iteration 4: Exclude item 'T05_04'.
#>
#> method iter itemExcluded DIF.excluded linking.constant linkerror
#> 1 iterativ 0 -0.232 0.061
#> 2 iterativ 1 T02_07 2.505 -0.195 0.050
#> 3 iterativ 2 T01_07 2.267 -0.160 0.036
#> 4 iterativ 3 T02_04 1.176 -0.142 0.032
#> 5 iterativ 4 T05_04 0.867 -0.129 0.029
#>
# linking constant is negative: students performance at T2 is worse than T1
# see that DIF is non symmetrical for reading: the four items with the highest
# amount of DIF have all positive DIF values. DIF exclusion hence shrinks linking
# constant towards zero.
# Third step: transform item parameters of 't2' to the metric of 't1'
# We now need to specify the 'refPop' argument. We use the values from 't1' which
# serves as the reference. To capture linking errors in a separate data.frame
# within the returned list, we define the years of assessment
ref <- tfRef[["refPop"]][,-2]
T.t1t2<- transformToBista ( equatingList = L.t1t2, refPop=ref, cuts = cuts,
vera=FALSE, years = c(2010,2015))
# The object 'T.t1t2' now contains transformed person and item parameters with
# original and transformed linking errors. See for example person parameter:
head(T.t1t2$personpars)
#> model idstud group imp value valueTransfBista traitLevel dimension linkingError linkingErrorTransfBista linkingErrorTraitLevel
#> 1 domainlistening P04477 domainlistening pv1 0.45849465 519.6722 3 domainlistening 0.04212599 5.031752 0.002074141
#> 2 domainlistening P04478 domainlistening pv1 0.21930774 491.1025 3 domainlistening 0.04212599 5.031752 0.002074141
#> 3 domainlistening P08882 domainlistening pv3 -0.16394757 445.3244 3 domainlistening 0.04212599 5.031752 0.002074141
#> 4 domainlistening P07848 domainlistening pv1 0.30686409 501.5607 3 domainlistening 0.04212599 5.031752 0.002074141
#> 5 domainlistening P06698 domainlistening pv2 -0.16098073 445.6788 3 domainlistening 0.04212599 5.031752 0.002074141
#> 6 domainlistening P04482 domainlistening pv1 0.03072918 468.5776 3 domainlistening 0.04212599 5.031752 0.002074141
# Fourth step: drawing plausible values for 't2'. We use the transformed item
# parameters (captured in 'T.t1t2') for anchoring
# Running second step: split models according to person groups (countries)
# ('all.persons' must be FALSE, otherwise the whole group would be treated as
# a separate distinct group.)
modT2P<- splitModels ( person.groups = datT2[,c("idstud", "country")] ,
all.persons = FALSE, nCores = 1)
#> --------------------------------
#> splitModels: generating 3 models
#> ...
#> see <returned>$models
#> number of cores: 1
#> --------------------------------
# define the 3 country-specific 2-dimensional models, specifying latent regression
# model and fixed item parameters. We used the transformed item parameters (captured
# in 'T.t1t2[["itempars"]]' --- using the 'estTransf' column) for anchoring.
# Again, 'ses' is scaled (mean = 0, sd = 1)
datT2[,"ses_scaled"] <- scale(datT2[,"ses"])[,1]
defT2P<- defineModel(dat = datT2, items = itemT2[,"item"], id = "idstud",
check.for.linking = TRUE, splittedModels = modT2P, qMatrix = qMat,
anchor = T.t1t2[["itempars"]][,c("item", "estTransf")],
HG.var = c("sex", "ses_scaled", "language"), software = "tam")
#>
#> Specification of 'qMatrix' and 'person.groups' results in 3 model(s).
#>
#>
#> ==========================================
#> Model No. 1
#> Model name: country.countryC
#> Number of items: 215
#> Number of persons: 1777
#> Number of dimensions: 2
#> ==========================================
#>
#> Following 68 item(s) missed in data frame will be removed from Q matrix:
#> T12_02, T09_11, T01_05, T13_16, T15_12, T15_11, T03_07, T10_08, T11_03, T04_01, T11_04, T04_07, T11_0X, T16_08, T16_09, T16_11, T16_13, T16_01, T16_04, T16_10, T16_02, T16_12, T16_06, T16_07, T06_04, T06_03, T08_04, T08_05, T06_1X, T26_03, T26_05, T26_07, T26_09, T26_06, T26_10, T26_08, T26_02, T26_04, T27_01, T27_05, T27_07, T27_06, T27_04, T27_02, T27_03, T30_06, T29_04, T29_02, T30_02, T30_03, T30_01, T29_05, T29_03, T28_08, T28_09, T28_02, T28_03, T28_04, T28_01, T28_10, T28_06, T32_04, T31_03, T31_01, T31_04, T32_03, T32_02, T32_01
#> Background variable(s) 'sex', 'language' of class
#> 'factor', 'factor' will be converted to indicator variables.
#> Variable 'sex' was converted to 1 indicator(s) with name(s) 'sexmale'.
#> Variable 'language' was converted to 2 indicator(s) with name(s) 'languagenativeAndOther', 'languageother'.
#> 3 subject(s) do not solve any item:
#> P05705 (18 false), P05714 (18 false), P05853 (19 false) ...
#> 1 subject(s) solved each item: P05805 (20 correct), P05805 (20 correct) ...
#> Dataset is completely linked.
#> 'gauss' has been chosen for estimation method. Number of nodes was not explicitly specified. Set nodes to 20.
#> 215 common items found in 'anchor' list and data frame.
#> Q matrix specifies 2 dimension(s).
#>
#>
#> ==========================================
#> Model No. 2
#> Model name: country.countryA
#> Number of items: 215
#> Number of persons: 1502
#> Number of dimensions: 2
#> ==========================================
#>
#> Following 68 item(s) missed in data frame will be removed from Q matrix:
#> T12_02, T09_11, T01_05, T13_16, T15_12, T15_11, T03_07, T10_08, T11_03, T04_01, T11_04, T04_07, T11_0X, T16_08, T16_09, T16_11, T16_13, T16_01, T16_04, T16_10, T16_02, T16_12, T16_06, T16_07, T06_04, T06_03, T08_04, T08_05, T06_1X, T26_03, T26_05, T26_07, T26_09, T26_06, T26_10, T26_08, T26_02, T26_04, T27_01, T27_05, T27_07, T27_06, T27_04, T27_02, T27_03, T30_06, T29_04, T29_02, T30_02, T30_03, T30_01, T29_05, T29_03, T28_08, T28_09, T28_02, T28_03, T28_04, T28_01, T28_10, T28_06, T32_04, T31_03, T31_01, T31_04, T32_03, T32_02, T32_01
#> Warning: 1 testitem(s) are constants. Remove these items from the data set:
#> Item 'T19_11', only value '0' occurs: 139 valid responses.
#> Background variable(s) 'sex', 'language' of class
#> 'factor', 'factor' will be converted to indicator variables.
#> Variable 'sex' was converted to 1 indicator(s) with name(s) 'sexmale'.
#> Variable 'language' was converted to 2 indicator(s) with name(s) 'languagenativeAndOther', 'languageother'.
#> Remove 1 test item(s) overall.
#> 1 subject(s) solved each item: P07096 (17 correct), P07096 (17 correct) ...
#> Dataset is completely linked.
#> 'gauss' has been chosen for estimation method. Number of nodes was not explicitly specified. Set nodes to 20.
#> 214 common items found in 'anchor' list and data frame.
#> 1 of 215 item(s) of merging variable 'item' from data set 'anchor list' not included in data set 'item response data'.
#> Q matrix specifies 2 dimension(s).
#>
#>
#> ==========================================
#> Model No. 3
#> Model name: country.countryB
#> Number of items: 215
#> Number of persons: 1237
#> Number of dimensions: 2
#> ==========================================
#>
#> Following 68 item(s) missed in data frame will be removed from Q matrix:
#> T12_02, T09_11, T01_05, T13_16, T15_12, T15_11, T03_07, T10_08, T11_03, T04_01, T11_04, T04_07, T11_0X, T16_08, T16_09, T16_11, T16_13, T16_01, T16_04, T16_10, T16_02, T16_12, T16_06, T16_07, T06_04, T06_03, T08_04, T08_05, T06_1X, T26_03, T26_05, T26_07, T26_09, T26_06, T26_10, T26_08, T26_02, T26_04, T27_01, T27_05, T27_07, T27_06, T27_04, T27_02, T27_03, T30_06, T29_04, T29_02, T30_02, T30_03, T30_01, T29_05, T29_03, T28_08, T28_09, T28_02, T28_03, T28_04, T28_01, T28_10, T28_06, T32_04, T31_03, T31_01, T31_04, T32_03, T32_02, T32_01
#> Warning: 1 testitem(s) are constants. Remove these items from the data set:
#> Item 'T02_07', only value '0' occurs: 147 valid responses.
#> Background variable(s) 'sex', 'language' of class
#> 'factor', 'factor' will be converted to indicator variables.
#> Variable 'sex' was converted to 1 indicator(s) with name(s) 'sexmale'.
#> Variable 'language' was converted to 2 indicator(s) with name(s) 'languagenativeAndOther', 'languageother'.
#> Remove 1 test item(s) overall.
#> Dataset is completely linked.
#> 'gauss' has been chosen for estimation method. Number of nodes was not explicitly specified. Set nodes to 20.
#> 214 common items found in 'anchor' list and data frame.
#> 1 of 215 item(s) of merging variable 'item' from data set 'anchor list' not included in data set 'item response data'.
#> Q matrix specifies 2 dimension(s).
# run the 3 models (estimation takes approx. 29 seconds)
runT2P<- runModel(defT2P)
# get the results
resT2P<- getResults(runT2P)
#> Q3 is only available for unidimensional models. Estimation will be skipped.
#> Getting standard errors with the tam.se function: 22.6 secs
#> Getting infit parameters calling tam.fit from getTamInfit: 4.1 secs
#> Getting WLEs calling tam.wle from getTamWles: 1.8 secs
#> |*****|
#> |-----|
#> Getting PVs calling tam.pv from getTamPVs: 0.5 secs
#> Q3 is only available for unidimensional models. Estimation will be skipped.
#> Getting standard errors with the tam.se function: 19.4 secs
#> Getting infit parameters calling tam.fit from getTamInfit: 3.1 secs
#> Getting WLEs calling tam.wle from getTamWles: 1.1 secs
#> |*****|
#> |-----|
#> Getting PVs calling tam.pv from getTamPVs: 0.4 secs
#> Q3 is only available for unidimensional models. Estimation will be skipped.
#> Getting standard errors with the tam.se function: 16.3 secs
#> Getting infit parameters calling tam.fit from getTamInfit: 2.9 secs
#> Getting WLEs calling tam.wle from getTamWles: 0.6 secs
#> |*****|
#> |-----|
#> Getting PVs calling tam.pv from getTamPVs: 0.3 secs
# equating is not necessary, as the models run with fixed item parameters
# However, to prepare for the transformation on the 'bista' metric, run
# 'equat1pl' with empty arguments
ankT2P<- equat1pl ( results = resT2P)
#> Found 3 model(s).
#> Equating is executed for each dimension in each model separately.
#> No norm parameter defined ('prmNorm' is missing). Treat current sample as drawn from the reference population.
# transformation to the 'bista' metric, using the previously defined cut scores
# and the reference population mean and sd from 't1'
dfrT2P<- transformToBista ( equatingList = ankT2P, refPop=ref, cuts=cuts, vera=FALSE)
################################################################################
### Example 6b: Extend example 6a with trend estimation (now for t3) ###
################################################################################
# Example 6b needs the objects (Q matrix, item parameters, ...) created in example 6
# and 6a. Preparation: assume time of measurement 't3'.
datT3<- reshape2::dcast(subset ( trends, year == 2020),
idstud+country+sex+ses+language~item, value.var="value")
# First step: item calibration in separate unidimensional models for each domain
modsT3<- splitModels ( qMatrix = qMat, nCores = 1)
#> --------------------------------
#> splitModels: generating 2 models
#> ..
#> see <returned>$models
#> number of cores: 1
#> --------------------------------
# define 2 models. Items which occur only in the Q matrix but not in the data
# will be ignored.
defT3 <- defineModel(dat = datT3, id = "idstud", check.for.linking = TRUE,
splittedModels = modsT3, software = "tam")
#>
#> Specification of 'qMatrix' and 'person.groups' results in 2 model(s).
#> Warning: Model split preparation for model No. 1, model name domainlistening: 15 items from 134 items listed the Q matrix not found in data:
#> ℹ 'T12_02', 'T13_16', 'T15_12', 'T15_11', 'T16_08', 'T16_09', 'T16_11', 'T16_13', 'T16_01', 'T16_04', 'T16_10', 'T16_02', 'T16_12', 'T16_06', 'T16_07'
#> Warning: Model split preparation for model No. 2, model name domainreading: 12 items from 149 items listed the Q matrix not found in data:
#> ℹ 'T09_11', 'T01_05', 'T03_07', 'T10_08', 'T11_03', 'T04_01', 'T11_04', 'T04_07', 'T06_04', 'T06_03', 'T08_04', 'T08_05'
#>
#>
#> =========================================
#> Model No. 1
#> Model name: domainlistening
#> Number of items: 119
#> Number of persons: 4532
#> Number of dimensions: 1
#> =========================================
#>
#> Following 15 item(s) missed in data frame will be removed from Q matrix:
#> T12_02, T13_16, T15_12, T15_11, T16_08, T16_09, T16_11, T16_13, T16_01, T16_04, T16_10, T16_02, T16_12, T16_06, T16_07
#> Found 212 cases with missings on all items.
#> Cases with missings on all items will be deleted.
#> 185 subject(s) do not solve any item:
#> P09749 (6 false), P12158 (6 false), P11510 (24 false) ...
#> 87 subject(s) solved each item: P10065 (7 correct), P09684 (10 correct), P11607 (21 correct) ...
#> Dataset is completely linked.
#> 'gauss' has been chosen for estimation method. Number of nodes was not explicitly specified. Set nodes to 20.
#> Q matrix specifies 1 dimension(s).
#>
#>
#> =======================================
#> Model No. 2
#> Model name: domainreading
#> Number of items: 137
#> Number of persons: 4532
#> Number of dimensions: 1
#> =======================================
#>
#> Following 12 item(s) missed in data frame will be removed from Q matrix:
#> T09_11, T01_05, T03_07, T10_08, T11_03, T04_01, T11_04, T04_07, T06_04, T06_03, T08_04, T08_05
#> Found 59 cases with missings on all items.
#> Cases with missings on all items will be deleted.
#> 83 subject(s) do not solve any item:
#> P09337 (6 false), P09202 (10 false), P10569 (17 false) ...
#> 137 subject(s) solved each item: P09247 (6 correct), P09425 (7 correct), P12928 (24 correct) ...
#> Dataset is completely linked.
#> 'gauss' has been chosen for estimation method. Number of nodes was not explicitly specified. Set nodes to 20.
#> Q matrix specifies 1 dimension(s).
# run 2 models
runT3 <- runModel(defT3)
# get the results
resT3 <- getResults(runT3)
#> Getting standard errors with the tam.se function: 1.4 secs
#> Getting infit parameters calling tam.fit from getTamInfit: 0.4 secs
#> Getting WLEs calling tam.wle from getTamWles: 0.9 secs
#> |*****|
#> |-----|
#> Getting PVs calling tam.pv from getTamPVs: 0.8 secs
#> Getting Q3 statistic calling tam.modelfit from getTamQ3: 4.4 secs
#> Getting standard errors with the tam.se function: 1.7 secs
#> Getting infit parameters calling tam.fit from getTamInfit: 0.4 secs
#> Getting WLEs calling tam.wle from getTamWles: 1.4 secs
#> |*****|
#> |-----|
#> Getting PVs calling tam.pv from getTamPVs: 0.8 secs
#> Getting Q3 statistic calling tam.modelfit from getTamQ3: 5.6 secs
# collect item parameters
itemT3<- itemFromRes(resT3)
# Second step: compute linking constant. We link the items of 't3' to the items
# of 't2' which are already transformed to the metric of 't1' (we use the item
# parameters which were used for plausible values imputation at 't2' as norm
# parameters)
L.t2t3<- equat1pl ( results = resT3, prmNorm = T.t1t2[["itempars"]][,c("item", "estTransf")],
excludeLinkingDif = TRUE, difBound = 0.64, iterativ = TRUE)
#> Found 2 model(s).
#> Equating is executed for each dimension in each model separately.
#>
#> ====================================================================================================
#>
#> Model No. 1
#> Model name: domainlistening
#> Number of dimension(s): 1
#> Name(s) of dimension(s): domainlistening
#> Number of linking items: 96
#> Linking method: Mean.Mean
#>
#> Dimension 'domainlistening': 6 of 96 items with linking |DIF| > 0.64 identified.
#> Iteration 1: Exclude item 'T19_11'.
#> Iteration 2: Exclude item 'T14_04'.
#> Iteration 3: Exclude item 'T20_30'.
#> Iteration 4: Exclude item 'T15_18'.
#> Iteration 5: Exclude item 'T20_03'.
#> Iteration 6: Exclude item 'T12_05'.
#> Iteration 7: Exclude item 'T20_04'.
#>
#> method iter itemExcluded DIF.excluded linking.constant linkerror
#> 1 iterativ 0 -0.302 0.052
#> 2 iterativ 1 T19_11 -2.536 -0.328 0.046
#> 3 iterativ 2 T14_04 -2.531 -0.355 0.037
#> 4 iterativ 3 T20_30 -1.749 -0.374 0.033
#> 5 iterativ 4 T15_18 -1.436 -0.389 0.029
#> 6 iterativ 5 T20_03 -0.704 -0.397 0.028
#> 7 iterativ 6 T12_05 0.678 -0.390 0.028
#> 8 iterativ 7 T20_04 -0.655 -0.397 0.027
#>
#>
#> ====================================================================================================
#>
#> Model No. 2
#> Model name: domainreading
#> Number of dimension(s): 1
#> Name(s) of dimension(s): domainreading
#> Number of linking items: 119
#> Linking method: Mean.Mean
#>
#> Dimension 'domainreading': 7 of 119 items with linking |DIF| > 0.64 identified.
#> Iteration 1: Exclude item 'T02_07'.
#> Iteration 2: Exclude item 'T05_01'.
#> Iteration 3: Exclude item 'T23_14'.
#> Iteration 4: Exclude item 'T04_08'.
#> Iteration 5: Exclude item 'T03_03'.
#> Iteration 6: Exclude item 'T10_10'.
#> Iteration 7: Exclude item 'T02_04'.
#>
#> method iter itemExcluded DIF.excluded linking.constant linkerror
#> 1 iterativ 0 -0.227 0.040
#> 2 iterativ 1 T02_07 -3.381 -0.255 0.029
#> 3 iterativ 2 T05_01 1.045 -0.246 0.028
#> 4 iterativ 3 T23_14 0.992 -0.238 0.026
#> 5 iterativ 4 T04_08 -0.915 -0.246 0.025
#> 6 iterativ 5 T03_03 0.687 -0.240 0.025
#> 7 iterativ 6 T10_10 -0.68 -0.246 0.024
#> 8 iterativ 7 T02_04 -0.654 -0.251 0.024
#>
# linking constant is negative: students performance at T3 is worse than T1
# Third step: transform item parameters of 't3' to the common metric of 't1' and 't2'
# We already know the 'refPop' values.
ref <- tfRef[["refPop"]][,-2]
T.t2t3<- transformToBista ( equatingList = L.t2t3, refPop=ref, cuts = cuts,
vera=FALSE, years = c(2015,2020))
# Fourth step: drawing plausible values for 't3'. We use the transformed item
# parameters (captured in 'T.t2t3') for anchoring
# Running second step: split models according to person groups (countries)
# ('all.persons' must be FALSE, otherwise the whole group would be treated as
# a separate distinct group.)
modT3P<- splitModels ( person.groups = datT3[,c("idstud", "country")] ,
all.persons = FALSE, nCores = 1)
#> --------------------------------
#> splitModels: generating 3 models
#> ...
#> see <returned>$models
#> number of cores: 1
#> --------------------------------
# define the 3 country-specific 2-dimensional models, specifying latent regression
# model and fixed item parameters. We used the transformed item parameters (captured
# in 'T.t2t3[["itempars"]]' --- using the 'estTransf' column) for anchoring.
# Again, 'ses' is scaled (mean = 0, sd = 1)
datT3[,"ses_scaled"] <- scale(datT3[,"ses"])[,1]
defT3P<- defineModel(dat = datT3, items = itemT3[,"item"], id = "idstud",
check.for.linking = TRUE, splittedModels = modT3P, qMatrix = qMat,
anchor = T.t2t3[["itempars"]][,c("item", "estTransf")],
HG.var = c("sex", "ses_scaled", "language"), software = "tam")
#>
#> Specification of 'qMatrix' and 'person.groups' results in 3 model(s).
#>
#>
#> ==========================================
#> Model No. 1
#> Model name: country.countryC
#> Number of items: 256
#> Number of persons: 1924
#> Number of dimensions: 2
#> ==========================================
#>
#> Following 27 item(s) missed in data frame will be removed from Q matrix:
#> T12_02, T09_11, T01_05, T13_16, T15_12, T15_11, T03_07, T10_08, T11_03, T04_01, T11_04, T04_07, T16_08, T16_09, T16_11, T16_13, T16_01, T16_04, T16_10, T16_02, T16_12, T16_06, T16_07, T06_04, T06_03, T08_04, T08_05
#> Background variable(s) 'sex', 'language' of class
#> 'factor', 'factor' will be converted to indicator variables.
#> Variable 'sex' was converted to 1 indicator(s) with name(s) 'sexmale'.
#> Variable 'language' was converted to 2 indicator(s) with name(s) 'languagenativeAndOther', 'languageother'.
#> 10 subject(s) do not solve any item:
#> P10086 (6 false), P09365 (21 false), P09127 (33 false) ...
#> 4 subject(s) solved each item: P10097 (6 correct), P10105 (6 correct), P10712 (6 correct) ...
#> Dataset is completely linked.
#> 'gauss' has been chosen for estimation method. Number of nodes was not explicitly specified. Set nodes to 20.
#> 256 common items found in 'anchor' list and data frame.
#> Q matrix specifies 2 dimension(s).
#>
#>
#> ==========================================
#> Model No. 2
#> Model name: country.countryA
#> Number of items: 256
#> Number of persons: 1363
#> Number of dimensions: 2
#> ==========================================
#>
#> Following 27 item(s) missed in data frame will be removed from Q matrix:
#> T12_02, T09_11, T01_05, T13_16, T15_12, T15_11, T03_07, T10_08, T11_03, T04_01, T11_04, T04_07, T16_08, T16_09, T16_11, T16_13, T16_01, T16_04, T16_10, T16_02, T16_12, T16_06, T16_07, T06_04, T06_03, T08_04, T08_05
#> Background variable(s) 'sex', 'language' of class
#> 'factor', 'factor' will be converted to indicator variables.
#> Variable 'sex' was converted to 1 indicator(s) with name(s) 'sexmale'.
#> Variable 'language' was converted to 2 indicator(s) with name(s) 'languagenativeAndOther', 'languageother'.
#> 7 subject(s) do not solve any item:
#> P11450 (8 false), P11264 (12 false), P11035 (20 false) ...
#> Dataset is completely linked.
#> 'gauss' has been chosen for estimation method. Number of nodes was not explicitly specified. Set nodes to 20.
#> 256 common items found in 'anchor' list and data frame.
#> Q matrix specifies 2 dimension(s).
#>
#>
#> ==========================================
#> Model No. 3
#> Model name: country.countryB
#> Number of items: 256
#> Number of persons: 1245
#> Number of dimensions: 2
#> ==========================================
#>
#> Following 27 item(s) missed in data frame will be removed from Q matrix:
#> T12_02, T09_11, T01_05, T13_16, T15_12, T15_11, T03_07, T10_08, T11_03, T04_01, T11_04, T04_07, T16_08, T16_09, T16_11, T16_13, T16_01, T16_04, T16_10, T16_02, T16_12, T16_06, T16_07, T06_04, T06_03, T08_04, T08_05
#> Background variable(s) 'sex', 'language' of class
#> 'factor', 'factor' will be converted to indicator variables.
#> Variable 'sex' was converted to 1 indicator(s) with name(s) 'sexmale'.
#> Variable 'language' was converted to 2 indicator(s) with name(s) 'languagenativeAndOther', 'languageother'.
#> 2 subject(s) do not solve any item:
#> P12905 (13 false), P13484 (19 false), P13484 (19 false) ...
#> 1 subject(s) solved each item: P13133 (17 correct), P13133 (17 correct) ...
#> Dataset is completely linked.
#> 'gauss' has been chosen for estimation method. Number of nodes was not explicitly specified. Set nodes to 20.
#> 256 common items found in 'anchor' list and data frame.
#> Q matrix specifies 2 dimension(s).
# run the 3 models (estimation takes approx. 20 seconds)
runT3P<- runModel(defT3P)
# get the results
resT3P<- getResults(runT3P)
#> Q3 is only available for unidimensional models. Estimation will be skipped.
#> Getting standard errors with the tam.se function: 1.2 mins
#> Getting infit parameters calling tam.fit from getTamInfit: 5.3 secs
#> Getting WLEs calling tam.wle from getTamWles: 1.8 secs
#> |*****|
#> |-----|
#> Getting PVs calling tam.pv from getTamPVs: 1.1 secs
#> Q3 is only available for unidimensional models. Estimation will be skipped.
#> Getting standard errors with the tam.se function: 54.1 secs
#> Getting infit parameters calling tam.fit from getTamInfit: 3.6 secs
#> Getting WLEs calling tam.wle from getTamWles: 1.4 secs
#> |*****|
#> |-----|
#> Getting PVs calling tam.pv from getTamPVs: 0.9 secs
#> Q3 is only available for unidimensional models. Estimation will be skipped.
#> Getting standard errors with the tam.se function: 53.1 secs
#> Getting infit parameters calling tam.fit from getTamInfit: 3.3 secs
#> Getting WLEs calling tam.wle from getTamWles: 0.6 secs
#> |*****|
#> |-----|
#> Getting PVs calling tam.pv from getTamPVs: 0.8 secs
# equating is not necessary, as the models run with fixed item parameters
# However, to prepare for the transformation on the 'bista' metric, run
# 'equat1pl' with empty arguments
ankT3P<- equat1pl ( results = resT3P)
#> Found 3 model(s).
#> Equating is executed for each dimension in each model separately.
#> No norm parameter defined ('prmNorm' is missing). Treat current sample as drawn from the reference population.
# transformation to the 'bista' metric, using the previously defined cut scores
# and the reference population mean and sd from 't1'
dfrT3P<- transformToBista ( equatingList = ankT3P, refPop=ref, cuts=cuts, vera=FALSE)
################################################################################
### Example 6c: Prepare trend estimation ###
################################################################################
# Example 6c needs the objects (Q matrix, item parameters, ...) created in example 6,
# 6a, and 6b.
# We now collect the person parameter estimates (plausible values) from t1, t2, and t3
# in a common data.frame. The person estimates are already collected in the objects
# previously created by 'transformToBista()'. Not all columns are necessary.
# plausibles values of measurement occasion 1 ('t1'): add the year to the data.frame
persT1<- data.frame ( year = 2010,
dfrT1P[["personpars"]][,c("idstud", "dimension", "imp", "value", "valueTransfBista", "traitLevel")],
stringsAsFactors = FALSE)
# plausibles values of measurement occasion 2 ('t2'): add the year to the data.frame
persT2<- data.frame ( year = 2015,
dfrT2P[["personpars"]][,c("idstud", "dimension", "imp", "value", "valueTransfBista", "traitLevel")],
stringsAsFactors = FALSE)
# plausibles values of measurement occasion 3 ('t3'): add the year to the data.frame
persT3<- data.frame ( year = 2020,
dfrT3P[["personpars"]][,c("idstud", "dimension", "imp", "value", "valueTransfBista", "traitLevel")],
stringsAsFactors = FALSE)
# bind together in a common data.frame
pers <- rbind(persT1, persT2, persT3)
# merge background variables to plausible values data
# first we have to create the 'domain' column in plausible values data
pers[,"domain"] <- car::recode(pers[,"dimension"], "'domainlistening'='listening'; 'domainreading'='reading'")
pers[,"dimension"] <- NULL
pers <- eatTools::mergeAttr(unique(trends[,c("year", "idclass", "idstud", "wgt", "jkzone", "jkrep", "domain", "country", "language", "ses", "sex")]),
pers, by = c("year", "idstud", "domain"), all = FALSE, setAttr=FALSE)
#> Merging levels are not unique in data set 'y'.
#> 376 of 27048 unit(s) of merging variable combination 'year'+'idstud'+'domain' from data set 'y' not included in data set 'x'.
# collect original linking errors
# t1 vs. t2: linking errors were computed in example 6a.
let1t2<- T.t1t2[["linkingErrors"]]
# t2 vs. t3: linking errors were computed in example 6b.
let2t3<- T.t2t3[["linkingErrors"]]
# t1 vs. t3: linking errors were not yet computed: link t3 to t1 to create linking error template
L.t1t3<- equat1pl ( results = resT3, prmNorm = itemT1[,c("item", "est")],
excludeLinkingDif = TRUE, difBound = 0.64, iterativ = TRUE)
#> Found 2 model(s).
#> Equating is executed for each dimension in each model separately.
#>
#> ====================================================================================================
#>
#> Model No. 1
#> Model name: domainlistening
#> Number of dimension(s): 1
#> Name(s) of dimension(s): domainlistening
#> Number of linking items: 36
#> Linking method: Mean.Mean
#>
#> Dimension 'domainlistening': 4 of 36 items with linking |DIF| > 0.64 identified.
#> Iteration 1: Exclude item 'T15_02'.
#> Iteration 2: Exclude item 'T12_05'.
#> Iteration 3: Exclude item 'T14_04'.
#>
#> method iter itemExcluded DIF.excluded linking.constant linkerror
#> 1 iterativ 0 -0.355 0.078
#> 2 iterativ 1 T15_02 -1.746 -0.406 0.061
#> 3 iterativ 2 T12_05 1.242 -0.370 0.051
#> 4 iterativ 3 T14_04 -0.886 -0.396 0.045
#>
#>
#> ====================================================================================================
#>
#> Model No. 2
#> Model name: domainreading
#> Number of dimension(s): 1
#> Name(s) of dimension(s): domainreading
#> Number of linking items: 68
#> Linking method: Mean.Mean
#>
#> Dimension 'domainreading': 4 of 68 items with linking |DIF| > 0.64 identified.
#> Iteration 1: Exclude item 'T01_07'.
#> Iteration 2: Exclude item 'T02_07'.
#> Iteration 3: Exclude item 'T09_07'.
#> Iteration 4: Exclude item 'T04_04'.
#> Iteration 5: Exclude item 'T07_10'.
#>
#> method iter itemExcluded DIF.excluded linking.constant linkerror
#> 1 iterativ 0 -0.305 0.048
#> 2 iterativ 1 T01_07 1.656 -0.280 0.041
#> 3 iterativ 2 T02_07 -0.849 -0.293 0.040
#> 4 iterativ 3 T09_07 -0.735 -0.304 0.039
#> 5 iterativ 4 T04_04 -0.67 -0.314 0.038
#> 6 iterativ 5 T07_10 -0.644 -0.325 0.037
#>
# indirect linking ('chained' linking)
chain <- multiEquatError (eq.1_2=L.t1t2, eq.2_3=L.t2t3, eq.1_3=L.t1t3)
#>
#> Dimension 'domainlistening': Direct linking error of the three combinations of measurements:
#> mzp1 mzp2 N.Items SD Var linkerror chained mzp_1.vs.3 mzp_1.vs.2.vs.3
#> 1 1 2 34 0.246 0.060 0.042 NA NA NA
#> 2 1 3 33 0.256 0.065 0.045 0.05 -0.396 -0.647
#> 3 2 3 89 0.253 0.064 0.027 NA NA NA
#>
#> Dimension 'domainreading': Direct linking error of the three combinations of measurements:
#> mzp1 mzp2 N.Items SD Var linkerror chained mzp_1.vs.3 mzp_1.vs.2.vs.3
#> 1 1 2 64 0.234 0.055 0.029 NA NA NA
#> 2 1 3 63 0.295 0.087 0.037 0.038 -0.325 -0.38
#> 3 2 3 112 0.253 0.064 0.024 NA NA NA
# replace direct linking errors with indirect linking errors
L.t1t3<- replaceLinkingError (equatingList =L.t1t3, multiEquatError_output=chain)
#> Dimension 'domainlistening': Replace old linking error 0.0445 with 0.0499
#> Dimension 'domainreading': Replace old linking error 0.0372 with 0.0378
# transform linking errors
ref <- tfRef[["refPop"]][,-2]
tle <- transformToBista ( equatingList = L.t1t3, refPop=ref, cuts = cuts,
vera=FALSE, years = c(2010,2020))
let1t3<- tle[["linkingErrors"]]
# bind all linking errors in a common data.frame
lErr <- rbind(let1t2, let2t3, let1t3)
lErr[,"domain"] <- car::recode(lErr[,"domain"], "'domainlistening'='listening'; 'domainreading'='reading'")
################################################################################
### Example 6d: eatRep estimation with plausible values and linking errors ###
################################################################################
# Example 6d needs the objects ('pers', 'lErr') created in example 6c
# load the 'eatRep' package ... note: needs eatRep version 0.14.0 or higher
library(eatRep)
#> Loading required package: survey
#> Loading required package: grid
#> Loading required package: Matrix
#> Loading required package: survival
#>
#> Attaching package: ‘survey’
#> The following object is masked from ‘package:graphics’:
#>
#> dotchart
#> Loading required package: BIFIEsurvey
#> |-----------------------------------------------------------------
#> | BIFIEsurvey 3.8.0 (2026-01-10 07:53:04 UTC)
#> | http://www.iqs.gv.at
#> |-----------------------------------------------------------------
#> Loading required package: progress
#> Loading required package: lavaan
#> This is lavaan 0.6-21
#> lavaan is FREE software! Please report any bugs.
# compute means for both countries with trend, for both domains separately,
# using replications methods (jackknife-1)
means <- by(data = pers, INDICES = pers[,"domain"], FUN = function ( dim ) {
m <- repMean(datL = dim, ID="idstud", PSU = "idclass", type = "jk1",
imp = "imp", groups = "country", dependent = "valueTransfBista",
trend = "year", linkErr = lErr[which(lErr[,"domain"] == dim[1,"domain"]),],
engine="BIFIEsurvey")
r <- report(m, add = list(domain = dim[1,"domain"]))
return(r)})
#>
#> Trend group: '2010'
#> 1 analyse(s) overall according to: 'group.splits = 1'.
#> Assume unnested structure with 5 imputations.
#>
#> `BIFIEsurvey::BIFIE.data.jack`(data = "datL", wgt = "wgtOne",
#> jktype = "JK_GROUP", jkzone = "idclass", jkrep = NULL, jkfac = NULL,
#> fayfac = 0.995833333333333, cdata = FALSE)
#> MI data with 5 datasets || 240 replication weights with fayfac=0.996 || 4476 cases and 9 variables
#> 'BIFIE.univar' for 'call = mean'. dependent = 'valueTransfBista'. group(s) = 'country'.
#>
#>
#> Trend group: '2015'
#> 1 analyse(s) overall according to: 'group.splits = 1'.
#> Assume unnested structure with 5 imputations.
#>
#> `BIFIEsurvey::BIFIE.data.jack`(data = "datL", wgt = "wgtOne",
#> jktype = "JK_GROUP", jkzone = "idclass", jkrep = NULL, jkfac = NULL,
#> fayfac = 0.995967741935484, cdata = FALSE)
#> MI data with 5 datasets || 248 replication weights with fayfac=0.996 || 4411 cases and 9 variables
#> 'BIFIE.univar' for 'call = mean'. dependent = 'valueTransfBista'. group(s) = 'country'.
#>
#>
#> Trend group: '2020'
#> 1 analyse(s) overall according to: 'group.splits = 1'.
#> Assume unnested structure with 5 imputations.
#>
#> `BIFIEsurvey::BIFIE.data.jack`(data = "datL", wgt = "wgtOne",
#> jktype = "JK_GROUP", jkzone = "idclass", jkrep = NULL, jkfac = NULL,
#> fayfac = 0.995951417004049, cdata = FALSE)
#> MI data with 5 datasets || 247 replication weights with fayfac=0.996 || 4320 cases and 9 variables
#> 'BIFIE.univar' for 'call = mean'. dependent = 'valueTransfBista'. group(s) = 'country'.
#>
#> Warning: `report()` was deprecated in eatRep 0.15.0.
#> ℹ For the original behavior of report() please use eatRep version 0.14.7: 'https://cran.r-project.org/src/contrib/Archive/eatRep/'
#> Warning: No linking errors for parameters 'NcasesValid', 'Ncases', 'sd'. Linking errors for these parameters will be defaulted to 0.
#> Warning: Found 3 missing linking errors for dependent variable 'valueTransfBista' and parameter(s) 'sd'. Assume linking error of 0 for these cases.
#> Warning: Found 3 missing linking errors for dependent variable 'valueTransfBista' and parameter(s) 'sd'. Assume linking error of 0 for these cases.
#> Warning: Found 3 missing linking errors for dependent variable 'valueTransfBista' and parameter(s) 'sd'. Assume linking error of 0 for these cases.
#>
#> Trend group: '2010'
#> 1 analyse(s) overall according to: 'group.splits = 1'.
#> Assume unnested structure with 5 imputations.
#>
#> `BIFIEsurvey::BIFIE.data.jack`(data = "datL", wgt = "wgtOne",
#> jktype = "JK_GROUP", jkzone = "idclass", jkrep = NULL, jkfac = NULL,
#> fayfac = 0.995833333333333, cdata = FALSE)
#> MI data with 5 datasets || 240 replication weights with fayfac=0.996 || 4476 cases and 9 variables
#> 'BIFIE.univar' for 'call = mean'. dependent = 'valueTransfBista'. group(s) = 'country'.
#>
#>
#> Trend group: '2015'
#> 1 analyse(s) overall according to: 'group.splits = 1'.
#> Assume unnested structure with 5 imputations.
#>
#> `BIFIEsurvey::BIFIE.data.jack`(data = "datL", wgt = "wgtOne",
#> jktype = "JK_GROUP", jkzone = "idclass", jkrep = NULL, jkfac = NULL,
#> fayfac = 0.996062992125984, cdata = FALSE)
#> MI data with 5 datasets || 254 replication weights with fayfac=0.996 || 4516 cases and 9 variables
#> 'BIFIE.univar' for 'call = mean'. dependent = 'valueTransfBista'. group(s) = 'country'.
#>
#>
#> Trend group: '2020'
#> 1 analyse(s) overall according to: 'group.splits = 1'.
#> Assume unnested structure with 5 imputations.
#>
#> `BIFIEsurvey::BIFIE.data.jack`(data = "datL", wgt = "wgtOne",
#> jktype = "JK_GROUP", jkzone = "idclass", jkrep = NULL, jkfac = NULL,
#> fayfac = 0.99609375, cdata = FALSE)
#> MI data with 5 datasets || 256 replication weights with fayfac=0.996 || 4473 cases and 9 variables
#> 'BIFIE.univar' for 'call = mean'. dependent = 'valueTransfBista'. group(s) = 'country'.
#>
#> Warning: No linking errors for parameters 'NcasesValid', 'Ncases', 'sd'. Linking errors for these parameters will be defaulted to 0.
#> Warning: Found 3 missing linking errors for dependent variable 'valueTransfBista' and parameter(s) 'sd'. Assume linking error of 0 for these cases.
#> Warning: Found 3 missing linking errors for dependent variable 'valueTransfBista' and parameter(s) 'sd'. Assume linking error of 0 for these cases.
#> Warning: Found 3 missing linking errors for dependent variable 'valueTransfBista' and parameter(s) 'sd'. Assume linking error of 0 for these cases.
means <- do.call("rbind", means)
# additionally: differ the sex-specific means in each country from the sex-specific means
# in the whole population? Are the differences (male vs. female) in each country different
# from the difference (male vs. female) in the whole population?
means2<- by(data = pers, INDICES = pers[,"domain"], FUN = function ( dim ) {
m <- repMean(datL = dim, ID="idstud", PSU = "idclass", type = "jk1",
imp = "imp", groups = c("country","sex"), group.differences.by = "sex",
group.splits = 0:1, cross.differences = TRUE,crossDiffSE.engine= "lm",
dependent = "valueTransfBista", trend = "year",
linkErr = lErr[which(lErr[,"domain"] == dim[1,"domain"]),],
engine="BIFIEsurvey")
r <- report(m, add = list(domain = dim[1,"domain"]))
return(r)})
#>
#> Trend group: '2010'
#> 3 analyse(s) overall according to: 'group.splits = 0 1'.
#>
#> analysis.number hierarchy.level dependent.var imp.var groups.divided.by group.differences.by
#> 1 0 valueTransfBista imp <NA>
#> 2 1 valueTransfBista imp country <NA>
#> 3 1 valueTransfBista imp sex sex
#>
#> Assume unnested structure with 5 imputations.
#>
#> `BIFIEsurvey::BIFIE.data.jack`(data = "datL", wgt = "wgtOne",
#> jktype = "JK_GROUP", jkzone = "idclass", jkrep = NULL, jkfac = NULL,
#> fayfac = 0.995833333333333, cdata = FALSE)
#> MI data with 5 datasets || 240 replication weights with fayfac=0.996 || 4476 cases and 11 variables
#> 'BIFIE.univar' for 'call = mean'. dependent = 'valueTransfBista'. group(s) = 'dummyGroup'.
#>
#> `BIFIEsurvey::BIFIE.data.jack`(data = "datL", wgt = "wgtOne",
#> jktype = "JK_GROUP", jkzone = "idclass", jkrep = NULL, jkfac = NULL,
#> fayfac = 0.995833333333333, cdata = FALSE)
#> MI data with 5 datasets || 240 replication weights with fayfac=0.996 || 4476 cases and 10 variables
#> 'BIFIE.univar' for 'call = mean'. dependent = 'valueTransfBista'. group(s) = 'country'.
#>
#> `BIFIEsurvey::BIFIE.data.jack`(data = "datL", wgt = "wgtOne",
#> jktype = "JK_GROUP", jkzone = "idclass", jkrep = NULL, jkfac = NULL,
#> fayfac = 0.995833333333333, cdata = FALSE)
#> MI data with 5 datasets || 240 replication weights with fayfac=0.996 || 4476 cases and 10 variables
#> 'BIFIE.univar' for 'call = mean'. dependent = 'valueTransfBista'. group(s) = 'sex'.
#>
#>
#> Trend group: '2015'
#> 3 analyse(s) overall according to: 'group.splits = 0 1'.
#>
#> analysis.number hierarchy.level dependent.var imp.var groups.divided.by group.differences.by
#> 1 0 valueTransfBista imp <NA>
#> 2 1 valueTransfBista imp country <NA>
#> 3 1 valueTransfBista imp sex sex
#>
#> Assume unnested structure with 5 imputations.
#>
#> `BIFIEsurvey::BIFIE.data.jack`(data = "datL", wgt = "wgtOne",
#> jktype = "JK_GROUP", jkzone = "idclass", jkrep = NULL, jkfac = NULL,
#> fayfac = 0.995967741935484, cdata = FALSE)
#> MI data with 5 datasets || 248 replication weights with fayfac=0.996 || 4411 cases and 11 variables
#> 'BIFIE.univar' for 'call = mean'. dependent = 'valueTransfBista'. group(s) = 'dummyGroup'.
#>
#> `BIFIEsurvey::BIFIE.data.jack`(data = "datL", wgt = "wgtOne",
#> jktype = "JK_GROUP", jkzone = "idclass", jkrep = NULL, jkfac = NULL,
#> fayfac = 0.995967741935484, cdata = FALSE)
#> MI data with 5 datasets || 248 replication weights with fayfac=0.996 || 4411 cases and 10 variables
#> 'BIFIE.univar' for 'call = mean'. dependent = 'valueTransfBista'. group(s) = 'country'.
#>
#> `BIFIEsurvey::BIFIE.data.jack`(data = "datL", wgt = "wgtOne",
#> jktype = "JK_GROUP", jkzone = "idclass", jkrep = NULL, jkfac = NULL,
#> fayfac = 0.995967741935484, cdata = FALSE)
#> MI data with 5 datasets || 248 replication weights with fayfac=0.996 || 4411 cases and 10 variables
#> 'BIFIE.univar' for 'call = mean'. dependent = 'valueTransfBista'. group(s) = 'sex'.
#>
#>
#> Trend group: '2020'
#> 3 analyse(s) overall according to: 'group.splits = 0 1'.
#>
#> analysis.number hierarchy.level dependent.var imp.var groups.divided.by group.differences.by
#> 1 0 valueTransfBista imp <NA>
#> 2 1 valueTransfBista imp country <NA>
#> 3 1 valueTransfBista imp sex sex
#>
#> Assume unnested structure with 5 imputations.
#>
#> `BIFIEsurvey::BIFIE.data.jack`(data = "datL", wgt = "wgtOne",
#> jktype = "JK_GROUP", jkzone = "idclass", jkrep = NULL, jkfac = NULL,
#> fayfac = 0.995951417004049, cdata = FALSE)
#> MI data with 5 datasets || 247 replication weights with fayfac=0.996 || 4320 cases and 11 variables
#> 'BIFIE.univar' for 'call = mean'. dependent = 'valueTransfBista'. group(s) = 'dummyGroup'.
#>
#> `BIFIEsurvey::BIFIE.data.jack`(data = "datL", wgt = "wgtOne",
#> jktype = "JK_GROUP", jkzone = "idclass", jkrep = NULL, jkfac = NULL,
#> fayfac = 0.995951417004049, cdata = FALSE)
#> MI data with 5 datasets || 247 replication weights with fayfac=0.996 || 4320 cases and 10 variables
#> 'BIFIE.univar' for 'call = mean'. dependent = 'valueTransfBista'. group(s) = 'country'.
#>
#> `BIFIEsurvey::BIFIE.data.jack`(data = "datL", wgt = "wgtOne",
#> jktype = "JK_GROUP", jkzone = "idclass", jkrep = NULL, jkfac = NULL,
#> fayfac = 0.995951417004049, cdata = FALSE)
#> MI data with 5 datasets || 247 replication weights with fayfac=0.996 || 4320 cases and 10 variables
#> 'BIFIE.univar' for 'call = mean'. dependent = 'valueTransfBista'. group(s) = 'sex'.
#>
#> Compute cross level differences using 'wec' method. Assume heteroscedastic variances.
#> 'wec' method: Assume equally weighted cases.
#> Warning: Group variable 'country' must be of class 'factor' for 'wec'. Change class of 'country' from 'character' to 'factor'.
#>
#> Trend group: '2010'
#> 1 analyse(s) overall according to: 'group.splits = 0'.
#> Assume unnested structure with 5 imputations.
#> Create 240 replicate weights according to JK1 procedure.
#>
#>
#> Trend group: '2015'
#> 1 analyse(s) overall according to: 'group.splits = 0'.
#> Assume unnested structure with 5 imputations.
#> Create 248 replicate weights according to JK1 procedure.
#>
#>
#> Trend group: '2020'
#> 1 analyse(s) overall according to: 'group.splits = 0'.
#> Assume unnested structure with 5 imputations.
#> Create 247 replicate weights according to JK1 procedure.
#>
#> Note: No linking error was defined. Linking error will be defaulted to '0'.
#> 'wec' method: Assume equally weighted cases.
#>
#> Trend group: '2010'
#> 1 analyse(s) overall according to: 'group.splits = 0'.
#> Assume unnested structure with 5 imputations.
#> Create 240 replicate weights according to JK1 procedure.
#>
#>
#> Trend group: '2015'
#> 1 analyse(s) overall according to: 'group.splits = 0'.
#> Assume unnested structure with 5 imputations.
#> Create 248 replicate weights according to JK1 procedure.
#>
#>
#> Trend group: '2020'
#> 1 analyse(s) overall according to: 'group.splits = 0'.
#> Assume unnested structure with 5 imputations.
#> Create 247 replicate weights according to JK1 procedure.
#>
#> Note: No linking error was defined. Linking error will be defaulted to '0'.
#> Warning: No linking errors for parameters 'NcasesValid', 'Ncases', 'sd'. Linking errors for these parameters will be defaulted to 0.
#> Warning: Found 11 missing linking errors for dependent variable 'valueTransfBista' and parameter(s) 'sd'. Assume linking error of 0 for these cases.
#> Warning: Found 11 missing linking errors for dependent variable 'valueTransfBista' and parameter(s) 'sd'. Assume linking error of 0 for these cases.
#> Warning: Found 11 missing linking errors for dependent variable 'valueTransfBista' and parameter(s) 'sd'. Assume linking error of 0 for these cases.
#>
#> Trend group: '2010'
#> 3 analyse(s) overall according to: 'group.splits = 0 1'.
#>
#> analysis.number hierarchy.level dependent.var imp.var groups.divided.by group.differences.by
#> 1 0 valueTransfBista imp <NA>
#> 2 1 valueTransfBista imp country <NA>
#> 3 1 valueTransfBista imp sex sex
#>
#> Assume unnested structure with 5 imputations.
#>
#> `BIFIEsurvey::BIFIE.data.jack`(data = "datL", wgt = "wgtOne",
#> jktype = "JK_GROUP", jkzone = "idclass", jkrep = NULL, jkfac = NULL,
#> fayfac = 0.995833333333333, cdata = FALSE)
#> MI data with 5 datasets || 240 replication weights with fayfac=0.996 || 4476 cases and 11 variables
#> 'BIFIE.univar' for 'call = mean'. dependent = 'valueTransfBista'. group(s) = 'dummyGroup'.
#>
#> `BIFIEsurvey::BIFIE.data.jack`(data = "datL", wgt = "wgtOne",
#> jktype = "JK_GROUP", jkzone = "idclass", jkrep = NULL, jkfac = NULL,
#> fayfac = 0.995833333333333, cdata = FALSE)
#> MI data with 5 datasets || 240 replication weights with fayfac=0.996 || 4476 cases and 10 variables
#> 'BIFIE.univar' for 'call = mean'. dependent = 'valueTransfBista'. group(s) = 'country'.
#>
#> `BIFIEsurvey::BIFIE.data.jack`(data = "datL", wgt = "wgtOne",
#> jktype = "JK_GROUP", jkzone = "idclass", jkrep = NULL, jkfac = NULL,
#> fayfac = 0.995833333333333, cdata = FALSE)
#> MI data with 5 datasets || 240 replication weights with fayfac=0.996 || 4476 cases and 10 variables
#> 'BIFIE.univar' for 'call = mean'. dependent = 'valueTransfBista'. group(s) = 'sex'.
#>
#>
#> Trend group: '2015'
#> 3 analyse(s) overall according to: 'group.splits = 0 1'.
#>
#> analysis.number hierarchy.level dependent.var imp.var groups.divided.by group.differences.by
#> 1 0 valueTransfBista imp <NA>
#> 2 1 valueTransfBista imp country <NA>
#> 3 1 valueTransfBista imp sex sex
#>
#> Assume unnested structure with 5 imputations.
#>
#> `BIFIEsurvey::BIFIE.data.jack`(data = "datL", wgt = "wgtOne",
#> jktype = "JK_GROUP", jkzone = "idclass", jkrep = NULL, jkfac = NULL,
#> fayfac = 0.996062992125984, cdata = FALSE)
#> MI data with 5 datasets || 254 replication weights with fayfac=0.996 || 4516 cases and 11 variables
#> 'BIFIE.univar' for 'call = mean'. dependent = 'valueTransfBista'. group(s) = 'dummyGroup'.
#>
#> `BIFIEsurvey::BIFIE.data.jack`(data = "datL", wgt = "wgtOne",
#> jktype = "JK_GROUP", jkzone = "idclass", jkrep = NULL, jkfac = NULL,
#> fayfac = 0.996062992125984, cdata = FALSE)
#> MI data with 5 datasets || 254 replication weights with fayfac=0.996 || 4516 cases and 10 variables
#> 'BIFIE.univar' for 'call = mean'. dependent = 'valueTransfBista'. group(s) = 'country'.
#>
#> `BIFIEsurvey::BIFIE.data.jack`(data = "datL", wgt = "wgtOne",
#> jktype = "JK_GROUP", jkzone = "idclass", jkrep = NULL, jkfac = NULL,
#> fayfac = 0.996062992125984, cdata = FALSE)
#> MI data with 5 datasets || 254 replication weights with fayfac=0.996 || 4516 cases and 10 variables
#> 'BIFIE.univar' for 'call = mean'. dependent = 'valueTransfBista'. group(s) = 'sex'.
#>
#>
#> Trend group: '2020'
#> 3 analyse(s) overall according to: 'group.splits = 0 1'.
#>
#> analysis.number hierarchy.level dependent.var imp.var groups.divided.by group.differences.by
#> 1 0 valueTransfBista imp <NA>
#> 2 1 valueTransfBista imp country <NA>
#> 3 1 valueTransfBista imp sex sex
#>
#> Assume unnested structure with 5 imputations.
#>
#> `BIFIEsurvey::BIFIE.data.jack`(data = "datL", wgt = "wgtOne",
#> jktype = "JK_GROUP", jkzone = "idclass", jkrep = NULL, jkfac = NULL,
#> fayfac = 0.99609375, cdata = FALSE)
#> MI data with 5 datasets || 256 replication weights with fayfac=0.996 || 4473 cases and 11 variables
#> 'BIFIE.univar' for 'call = mean'. dependent = 'valueTransfBista'. group(s) = 'dummyGroup'.
#>
#> `BIFIEsurvey::BIFIE.data.jack`(data = "datL", wgt = "wgtOne",
#> jktype = "JK_GROUP", jkzone = "idclass", jkrep = NULL, jkfac = NULL,
#> fayfac = 0.99609375, cdata = FALSE)
#> MI data with 5 datasets || 256 replication weights with fayfac=0.996 || 4473 cases and 10 variables
#> 'BIFIE.univar' for 'call = mean'. dependent = 'valueTransfBista'. group(s) = 'country'.
#>
#> `BIFIEsurvey::BIFIE.data.jack`(data = "datL", wgt = "wgtOne",
#> jktype = "JK_GROUP", jkzone = "idclass", jkrep = NULL, jkfac = NULL,
#> fayfac = 0.99609375, cdata = FALSE)
#> MI data with 5 datasets || 256 replication weights with fayfac=0.996 || 4473 cases and 10 variables
#> 'BIFIE.univar' for 'call = mean'. dependent = 'valueTransfBista'. group(s) = 'sex'.
#>
#> Compute cross level differences using 'wec' method. Assume heteroscedastic variances.
#> 'wec' method: Assume equally weighted cases.
#> Warning: Group variable 'country' must be of class 'factor' for 'wec'. Change class of 'country' from 'character' to 'factor'.
#>
#> Trend group: '2010'
#> 1 analyse(s) overall according to: 'group.splits = 0'.
#> Assume unnested structure with 5 imputations.
#> Create 240 replicate weights according to JK1 procedure.
#>
#>
#> Trend group: '2015'
#> 1 analyse(s) overall according to: 'group.splits = 0'.
#> Assume unnested structure with 5 imputations.
#> Create 254 replicate weights according to JK1 procedure.
#>
#>
#> Trend group: '2020'
#> 1 analyse(s) overall according to: 'group.splits = 0'.
#> Assume unnested structure with 5 imputations.
#> Create 256 replicate weights according to JK1 procedure.
#>
#> Note: No linking error was defined. Linking error will be defaulted to '0'.
#> 'wec' method: Assume equally weighted cases.
#>
#> Trend group: '2010'
#> 1 analyse(s) overall according to: 'group.splits = 0'.
#> Assume unnested structure with 5 imputations.
#> Create 240 replicate weights according to JK1 procedure.
#>
#>
#> Trend group: '2015'
#> 1 analyse(s) overall according to: 'group.splits = 0'.
#> Assume unnested structure with 5 imputations.
#> Create 254 replicate weights according to JK1 procedure.
#>
#>
#> Trend group: '2020'
#> 1 analyse(s) overall according to: 'group.splits = 0'.
#> Assume unnested structure with 5 imputations.
#> Create 256 replicate weights according to JK1 procedure.
#>
#> Note: No linking error was defined. Linking error will be defaulted to '0'.
#> Warning: No linking errors for parameters 'NcasesValid', 'Ncases', 'sd'. Linking errors for these parameters will be defaulted to 0.
#> Warning: Found 11 missing linking errors for dependent variable 'valueTransfBista' and parameter(s) 'sd'. Assume linking error of 0 for these cases.
#> Warning: Found 11 missing linking errors for dependent variable 'valueTransfBista' and parameter(s) 'sd'. Assume linking error of 0 for these cases.
#> Warning: Found 11 missing linking errors for dependent variable 'valueTransfBista' and parameter(s) 'sd'. Assume linking error of 0 for these cases.
means2<- do.call("rbind", means2)
################################################################################
### Example 6e: eatRep (repTable) with plausible values and linking errors ###
################################################################################
# Example 6e needs the objects ('pers', 'lErr') created in example 6c
# load the 'eatRep' package ... note: needs eatRep version 0.14.0 or higher
library(eatRep)
# compute frequencies for trait levels, for both domains, with trend
freqs <- by(data = pers, INDICES = pers[,"domain"], FUN = function ( dim ) {
m <- repTable(datL = dim, ID="idstud", PSU = "idclass", type = "jk1",
imp = "imp", groups = "country", dependent = "traitLevel",
trend = "year", linkErr = lErr[which(lErr[,"domain"] == dim[1,"domain"]),])
r <- report(m, add = list(domain = dim[1,"domain"]))
return(r)})
#>
#> Trend group: '2010'
#> 1 analyse(s) overall according to: 'group.splits = 1'.
#> Assume unnested structure with 5 imputations.
#> Create 240 replicate weights according to JK1 procedure.
#>
#>
#> Trend group: '2015'
#> 1 analyse(s) overall according to: 'group.splits = 1'.
#> Assume unnested structure with 5 imputations.
#> Create 248 replicate weights according to JK1 procedure.
#>
#>
#> Trend group: '2020'
#> 1 analyse(s) overall according to: 'group.splits = 1'.
#> Assume unnested structure with 5 imputations.
#> Create 247 replicate weights according to JK1 procedure.
#>
#> Warning: No linking errors for parameters 'Ncases'. Linking errors for these parameters will be defaulted to 0.
#>
#> Trend group: '2010'
#> 1 analyse(s) overall according to: 'group.splits = 1'.
#> Assume unnested structure with 5 imputations.
#> Create 240 replicate weights according to JK1 procedure.
#>
#>
#> Trend group: '2015'
#> 1 analyse(s) overall according to: 'group.splits = 1'.
#> Assume unnested structure with 5 imputations.
#> Create 254 replicate weights according to JK1 procedure.
#>
#>
#> Trend group: '2020'
#> 1 analyse(s) overall according to: 'group.splits = 1'.
#> Assume unnested structure with 5 imputations.
#> Create 256 replicate weights according to JK1 procedure.
#>
#> Warning: No linking errors for parameters 'Ncases'. Linking errors for these parameters will be defaulted to 0.
freqs <- do.call("rbind", freqs)
# additionally: sex differences in each country, using 'group.differences.by' argument
# Note: for frequency tables group differences may result in a chi square test or in
# a difference of each categories' frequency.
# first: request chi square test
freqs1<- by(data = pers, INDICES = pers[,"domain"], FUN = function ( dim ) {
m <- repTable(datL = dim, ID="idstud", PSU = "idclass", type = "jk1",
imp = "imp", groups = c("country","sex"), group.differences.by = "sex",
chiSquare = TRUE,dependent = "traitLevel", trend = "year",
linkErr = lErr[which(lErr[,"domain"] == dim[1,"domain"]),])
r <- report(m, add = list(domain = dim[1,"domain"]))
return(r)})
#>
#> Trend group: '2010'
#> 1 analyse(s) overall according to: 'group.splits = 2'.
#> Assume unnested structure with 5 imputations.
#> Create 240 replicate weights according to JK1 procedure.
#>
#>
#> Trend group: '2015'
#> 1 analyse(s) overall according to: 'group.splits = 2'.
#> Assume unnested structure with 5 imputations.
#> Create 248 replicate weights according to JK1 procedure.
#>
#>
#> Trend group: '2020'
#> 1 analyse(s) overall according to: 'group.splits = 2'.
#> Assume unnested structure with 5 imputations.
#> Create 247 replicate weights according to JK1 procedure.
#>
#> Warning: No linking errors for parameters 'chiSquareTest', 'Ncases'. Linking errors for these parameters will be defaulted to 0.
#> Chi sqare test results cannot be transferred to old report() structure and will be ignored. Please use report2() instead.
#> Trend group: '2010'
#> 1 analyse(s) overall according to: 'group.splits = 2'.
#> Assume unnested structure with 5 imputations.
#> Create 240 replicate weights according to JK1 procedure.
#>
#>
#> Trend group: '2015'
#> 1 analyse(s) overall according to: 'group.splits = 2'.
#> Assume unnested structure with 5 imputations.
#> Create 254 replicate weights according to JK1 procedure.
#>
#>
#> Trend group: '2020'
#> 1 analyse(s) overall according to: 'group.splits = 2'.
#> Assume unnested structure with 5 imputations.
#> Create 256 replicate weights according to JK1 procedure.
#>
#> Warning: No linking errors for parameters 'chiSquareTest', 'Ncases'. Linking errors for these parameters will be defaulted to 0.
#> Chi sqare test results cannot be transferred to old report() structure and will be ignored. Please use report2() instead.
freqs1<- do.call("rbind", freqs1)
# differences for each competence level (chiSquare = FALSE)
# (for faster computation, we omit jackknife procedure)
freqs2<- by(data = pers, INDICES = pers[,"domain"], FUN = function ( dim ) {
m <- repTable(datL = dim, ID="idstud", type = "none", imp = "imp",
groups = c("country","sex"), group.differences.by = "sex",
chiSquare = FALSE,dependent = "traitLevel", trend = "year",
linkErr = lErr[which(lErr[,"domain"] == dim[1,"domain"]),],
group.splits = 0:2, cross.differences = TRUE)
r <- report(m, add = list(domain = dim[1,"domain"]))
return(r)})
#> To date, only method 'old' is applicable for cross level differences in frequency tables.
#>
#> Trend group: '2010'
#> 4 analyse(s) overall according to: 'group.splits = 0 1 2'.
#>
#> analysis.number hierarchy.level dependent.var imp.var groups.divided.by group.differences.by
#> 1 0 traitLevel1 imp <NA>
#> 2 1 traitLevel1 imp country <NA>
#> 3 1 traitLevel1 imp sex sex
#> 4 2 traitLevel1 imp country + sex sex
#>
#> Assume unnested structure with 5 imputations.
#>
#>
#> Trend group: '2015'
#> 4 analyse(s) overall according to: 'group.splits = 0 1 2'.
#>
#> analysis.number hierarchy.level dependent.var imp.var groups.divided.by group.differences.by
#> 1 0 traitLevel1 imp <NA>
#> 2 1 traitLevel1 imp country <NA>
#> 3 1 traitLevel1 imp sex sex
#> 4 2 traitLevel1 imp country + sex sex
#>
#> Assume unnested structure with 5 imputations.
#>
#>
#> Trend group: '2020'
#> 4 analyse(s) overall according to: 'group.splits = 0 1 2'.
#>
#> analysis.number hierarchy.level dependent.var imp.var groups.divided.by group.differences.by
#> 1 0 traitLevel1 imp <NA>
#> 2 1 traitLevel1 imp country <NA>
#> 3 1 traitLevel1 imp sex sex
#> 4 2 traitLevel1 imp country + sex sex
#>
#> Assume unnested structure with 5 imputations.
#>
#>
#> Trend group: '2010'
#> 4 analyse(s) overall according to: 'group.splits = 0 1 2'.
#>
#> analysis.number hierarchy.level dependent.var imp.var groups.divided.by group.differences.by
#> 1 0 traitLevel2 imp <NA>
#> 2 1 traitLevel2 imp country <NA>
#> 3 1 traitLevel2 imp sex sex
#> 4 2 traitLevel2 imp country + sex sex
#>
#> Assume unnested structure with 5 imputations.
#>
#>
#> Trend group: '2015'
#> 4 analyse(s) overall according to: 'group.splits = 0 1 2'.
#>
#> analysis.number hierarchy.level dependent.var imp.var groups.divided.by group.differences.by
#> 1 0 traitLevel2 imp <NA>
#> 2 1 traitLevel2 imp country <NA>
#> 3 1 traitLevel2 imp sex sex
#> 4 2 traitLevel2 imp country + sex sex
#>
#> Assume unnested structure with 5 imputations.
#>
#>
#> Trend group: '2020'
#> 4 analyse(s) overall according to: 'group.splits = 0 1 2'.
#>
#> analysis.number hierarchy.level dependent.var imp.var groups.divided.by group.differences.by
#> 1 0 traitLevel2 imp <NA>
#> 2 1 traitLevel2 imp country <NA>
#> 3 1 traitLevel2 imp sex sex
#> 4 2 traitLevel2 imp country + sex sex
#>
#> Assume unnested structure with 5 imputations.
#>
#>
#> Trend group: '2010'
#> 4 analyse(s) overall according to: 'group.splits = 0 1 2'.
#>
#> analysis.number hierarchy.level dependent.var imp.var groups.divided.by group.differences.by
#> 1 0 traitLevel3 imp <NA>
#> 2 1 traitLevel3 imp country <NA>
#> 3 1 traitLevel3 imp sex sex
#> 4 2 traitLevel3 imp country + sex sex
#>
#> Assume unnested structure with 5 imputations.
#>
#>
#> Trend group: '2015'
#> 4 analyse(s) overall according to: 'group.splits = 0 1 2'.
#>
#> analysis.number hierarchy.level dependent.var imp.var groups.divided.by group.differences.by
#> 1 0 traitLevel3 imp <NA>
#> 2 1 traitLevel3 imp country <NA>
#> 3 1 traitLevel3 imp sex sex
#> 4 2 traitLevel3 imp country + sex sex
#>
#> Assume unnested structure with 5 imputations.
#>
#>
#> Trend group: '2020'
#> 4 analyse(s) overall according to: 'group.splits = 0 1 2'.
#>
#> analysis.number hierarchy.level dependent.var imp.var groups.divided.by group.differences.by
#> 1 0 traitLevel3 imp <NA>
#> 2 1 traitLevel3 imp country <NA>
#> 3 1 traitLevel3 imp sex sex
#> 4 2 traitLevel3 imp country + sex sex
#>
#> Assume unnested structure with 5 imputations.
#>
#>
#> Trend group: '2010'
#> 4 analyse(s) overall according to: 'group.splits = 0 1 2'.
#>
#> analysis.number hierarchy.level dependent.var imp.var groups.divided.by group.differences.by
#> 1 0 traitLevel4 imp <NA>
#> 2 1 traitLevel4 imp country <NA>
#> 3 1 traitLevel4 imp sex sex
#> 4 2 traitLevel4 imp country + sex sex
#>
#> Assume unnested structure with 5 imputations.
#>
#>
#> Trend group: '2015'
#> 4 analyse(s) overall according to: 'group.splits = 0 1 2'.
#>
#> analysis.number hierarchy.level dependent.var imp.var groups.divided.by group.differences.by
#> 1 0 traitLevel4 imp <NA>
#> 2 1 traitLevel4 imp country <NA>
#> 3 1 traitLevel4 imp sex sex
#> 4 2 traitLevel4 imp country + sex sex
#>
#> Assume unnested structure with 5 imputations.
#>
#>
#> Trend group: '2020'
#> 4 analyse(s) overall according to: 'group.splits = 0 1 2'.
#>
#> analysis.number hierarchy.level dependent.var imp.var groups.divided.by group.differences.by
#> 1 0 traitLevel4 imp <NA>
#> 2 1 traitLevel4 imp country <NA>
#> 3 1 traitLevel4 imp sex sex
#> 4 2 traitLevel4 imp country + sex sex
#>
#> Assume unnested structure with 5 imputations.
#>
#>
#> Trend group: '2010'
#> 4 analyse(s) overall according to: 'group.splits = 0 1 2'.
#>
#> analysis.number hierarchy.level dependent.var imp.var groups.divided.by group.differences.by
#> 1 0 traitLevel5 imp <NA>
#> 2 1 traitLevel5 imp country <NA>
#> 3 1 traitLevel5 imp sex sex
#> 4 2 traitLevel5 imp country + sex sex
#>
#> Assume unnested structure with 5 imputations.
#>
#>
#> Trend group: '2015'
#> 4 analyse(s) overall according to: 'group.splits = 0 1 2'.
#>
#> analysis.number hierarchy.level dependent.var imp.var groups.divided.by group.differences.by
#> 1 0 traitLevel5 imp <NA>
#> 2 1 traitLevel5 imp country <NA>
#> 3 1 traitLevel5 imp sex sex
#> 4 2 traitLevel5 imp country + sex sex
#>
#> Assume unnested structure with 5 imputations.
#>
#>
#> Trend group: '2020'
#> 4 analyse(s) overall according to: 'group.splits = 0 1 2'.
#>
#> analysis.number hierarchy.level dependent.var imp.var groups.divided.by group.differences.by
#> 1 0 traitLevel5 imp <NA>
#> 2 1 traitLevel5 imp country <NA>
#> 3 1 traitLevel5 imp sex sex
#> 4 2 traitLevel5 imp country + sex sex
#>
#> Assume unnested structure with 5 imputations.
#>
#> Warning: No standard error for parameter 'NcasesValid'. Cannot compute standard errors and p value for cross-level difference between 'wholeGroup' and 'countryA'.
#> Warning: No standard error for parameter 'NcasesValid'. Cannot compute standard errors and p value for cross-level difference between 'wholeGroup' and 'countryB'.
#> Warning: No standard error for parameter 'NcasesValid'. Cannot compute standard errors and p value for cross-level difference between 'wholeGroup' and 'countryC'.
#> Warning: No standard error for parameter 'NcasesValid'. Cannot compute standard errors and p value for cross-level difference between 'wholeGroup' and 'female'.
#> Warning: No standard error for parameter 'NcasesValid'. Cannot compute standard errors and p value for cross-level difference between 'wholeGroup' and 'male'.
#> Warning: No standard error for parameter 'NcasesValid'. Cannot compute standard errors and p value for cross-level difference between 'wholeGroup' and 'countryA_female'.
#> Warning: No standard error for parameter 'NcasesValid'. Cannot compute standard errors and p value for cross-level difference between 'wholeGroup' and 'countryA_male'.
#> Warning: No standard error for parameter 'NcasesValid'. Cannot compute standard errors and p value for cross-level difference between 'wholeGroup' and 'countryB_female'.
#> Warning: No standard error for parameter 'NcasesValid'. Cannot compute standard errors and p value for cross-level difference between 'wholeGroup' and 'countryB_male'.
#> Warning: No standard error for parameter 'NcasesValid'. Cannot compute standard errors and p value for cross-level difference between 'wholeGroup' and 'countryC_female'.
#> Warning: No standard error for parameter 'NcasesValid'. Cannot compute standard errors and p value for cross-level difference between 'wholeGroup' and 'countryC_male'.
#> Warning: No standard error for parameter 'NcasesValid'. Cannot compute standard errors and p value for cross-level difference between 'countryA' and 'countryA_female'.
#> Warning: No standard error for parameter 'NcasesValid'. Cannot compute standard errors and p value for cross-level difference between 'countryA' and 'countryA_male'.
#> Warning: No standard error for parameter 'NcasesValid'. Cannot compute standard errors and p value for cross-level difference between 'countryB' and 'countryB_female'.
#> Warning: No standard error for parameter 'NcasesValid'. Cannot compute standard errors and p value for cross-level difference between 'countryB' and 'countryB_male'.
#> Warning: No standard error for parameter 'NcasesValid'. Cannot compute standard errors and p value for cross-level difference between 'countryC' and 'countryC_female'.
#> Warning: No standard error for parameter 'NcasesValid'. Cannot compute standard errors and p value for cross-level difference between 'countryC' and 'countryC_male'.
#> Warning: No standard error for parameter 'NcasesValid'. Cannot compute standard errors and p value for cross-level difference between 'female' and 'countryA_female'.
#> Warning: No standard error for parameter 'NcasesValid'. Cannot compute standard errors and p value for cross-level difference between 'female' and 'countryB_female'.
#> Warning: No standard error for parameter 'NcasesValid'. Cannot compute standard errors and p value for cross-level difference between 'female' and 'countryC_female'.
#> Warning: No standard error for parameter 'NcasesValid'. Cannot compute standard errors and p value for cross-level difference between 'male' and 'countryA_male'.
#> Warning: No standard error for parameter 'NcasesValid'. Cannot compute standard errors and p value for cross-level difference between 'male' and 'countryB_male'.
#> Warning: No standard error for parameter 'NcasesValid'. Cannot compute standard errors and p value for cross-level difference between 'male' and 'countryC_male'.
#> Warning: No standard error for parameter 'NcasesValid'. Cannot compute standard errors and p value for cross-level difference between 'wholeGroup' and 'countryA'.
#> Warning: No standard error for parameter 'NcasesValid'. Cannot compute standard errors and p value for cross-level difference between 'wholeGroup' and 'countryB'.
#> Warning: No standard error for parameter 'NcasesValid'. Cannot compute standard errors and p value for cross-level difference between 'wholeGroup' and 'countryC'.
#> Warning: No standard error for parameter 'NcasesValid'. Cannot compute standard errors and p value for cross-level difference between 'wholeGroup' and 'female'.
#> Warning: No standard error for parameter 'NcasesValid'. Cannot compute standard errors and p value for cross-level difference between 'wholeGroup' and 'male'.
#> Warning: No standard error for parameter 'NcasesValid'. Cannot compute standard errors and p value for cross-level difference between 'wholeGroup' and 'countryA_female'.
#> Warning: No standard error for parameter 'NcasesValid'. Cannot compute standard errors and p value for cross-level difference between 'wholeGroup' and 'countryA_male'.
#> Warning: No standard error for parameter 'NcasesValid'. Cannot compute standard errors and p value for cross-level difference between 'wholeGroup' and 'countryB_female'.
#> Warning: No standard error for parameter 'NcasesValid'. Cannot compute standard errors and p value for cross-level difference between 'wholeGroup' and 'countryB_male'.
#> Warning: No standard error for parameter 'NcasesValid'. Cannot compute standard errors and p value for cross-level difference between 'wholeGroup' and 'countryC_female'.
#> Warning: No standard error for parameter 'NcasesValid'. Cannot compute standard errors and p value for cross-level difference between 'wholeGroup' and 'countryC_male'.
#> Warning: No standard error for parameter 'NcasesValid'. Cannot compute standard errors and p value for cross-level difference between 'countryA' and 'countryA_female'.
#> Warning: No standard error for parameter 'NcasesValid'. Cannot compute standard errors and p value for cross-level difference between 'countryA' and 'countryA_male'.
#> Warning: No standard error for parameter 'NcasesValid'. Cannot compute standard errors and p value for cross-level difference between 'countryB' and 'countryB_female'.
#> Warning: No standard error for parameter 'NcasesValid'. Cannot compute standard errors and p value for cross-level difference between 'countryB' and 'countryB_male'.
#> Warning: No standard error for parameter 'NcasesValid'. Cannot compute standard errors and p value for cross-level difference between 'countryC' and 'countryC_female'.
#> Warning: No standard error for parameter 'NcasesValid'. Cannot compute standard errors and p value for cross-level difference between 'countryC' and 'countryC_male'.
#> Warning: No standard error for parameter 'NcasesValid'. Cannot compute standard errors and p value for cross-level difference between 'female' and 'countryA_female'.
#> Warning: No standard error for parameter 'NcasesValid'. Cannot compute standard errors and p value for cross-level difference between 'female' and 'countryB_female'.
#> Warning: No standard error for parameter 'NcasesValid'. Cannot compute standard errors and p value for cross-level difference between 'female' and 'countryC_female'.
#> Warning: No standard error for parameter 'NcasesValid'. Cannot compute standard errors and p value for cross-level difference between 'male' and 'countryA_male'.
#> Warning: No standard error for parameter 'NcasesValid'. Cannot compute standard errors and p value for cross-level difference between 'male' and 'countryB_male'.
#> Warning: No standard error for parameter 'NcasesValid'. Cannot compute standard errors and p value for cross-level difference between 'male' and 'countryC_male'.
#> Warning: No standard error for parameter 'NcasesValid'. Cannot compute standard errors and p value for cross-level difference between 'wholeGroup' and 'countryA'.
#> Warning: No standard error for parameter 'NcasesValid'. Cannot compute standard errors and p value for cross-level difference between 'wholeGroup' and 'countryB'.
#> Warning: No standard error for parameter 'NcasesValid'. Cannot compute standard errors and p value for cross-level difference between 'wholeGroup' and 'countryC'.
#> Warning: No standard error for parameter 'NcasesValid'. Cannot compute standard errors and p value for cross-level difference between 'wholeGroup' and 'female'.
#> Warning: No standard error for parameter 'NcasesValid'. Cannot compute standard errors and p value for cross-level difference between 'wholeGroup' and 'male'.
#> Warning: No standard error for parameter 'NcasesValid'. Cannot compute standard errors and p value for cross-level difference between 'wholeGroup' and 'countryA_female'.
#> Warning: No standard error for parameter 'NcasesValid'. Cannot compute standard errors and p value for cross-level difference between 'wholeGroup' and 'countryA_male'.
#> Warning: No standard error for parameter 'NcasesValid'. Cannot compute standard errors and p value for cross-level difference between 'wholeGroup' and 'countryB_female'.
#> Warning: No standard error for parameter 'NcasesValid'. Cannot compute standard errors and p value for cross-level difference between 'wholeGroup' and 'countryB_male'.
#> Warning: No standard error for parameter 'NcasesValid'. Cannot compute standard errors and p value for cross-level difference between 'wholeGroup' and 'countryC_female'.
#> Warning: No standard error for parameter 'NcasesValid'. Cannot compute standard errors and p value for cross-level difference between 'wholeGroup' and 'countryC_male'.
#> Warning: No standard error for parameter 'NcasesValid'. Cannot compute standard errors and p value for cross-level difference between 'countryA' and 'countryA_female'.
#> Warning: No standard error for parameter 'NcasesValid'. Cannot compute standard errors and p value for cross-level difference between 'countryA' and 'countryA_male'.
#> Warning: No standard error for parameter 'NcasesValid'. Cannot compute standard errors and p value for cross-level difference between 'countryB' and 'countryB_female'.
#> Warning: No standard error for parameter 'NcasesValid'. Cannot compute standard errors and p value for cross-level difference between 'countryB' and 'countryB_male'.
#> Warning: No standard error for parameter 'NcasesValid'. Cannot compute standard errors and p value for cross-level difference between 'countryC' and 'countryC_female'.
#> Warning: No standard error for parameter 'NcasesValid'. Cannot compute standard errors and p value for cross-level difference between 'countryC' and 'countryC_male'.
#> Warning: No standard error for parameter 'NcasesValid'. Cannot compute standard errors and p value for cross-level difference between 'female' and 'countryA_female'.
#> Warning: No standard error for parameter 'NcasesValid'. Cannot compute standard errors and p value for cross-level difference between 'female' and 'countryB_female'.
#> Warning: No standard error for parameter 'NcasesValid'. Cannot compute standard errors and p value for cross-level difference between 'female' and 'countryC_female'.
#> Warning: No standard error for parameter 'NcasesValid'. Cannot compute standard errors and p value for cross-level difference between 'male' and 'countryA_male'.
#> Warning: No standard error for parameter 'NcasesValid'. Cannot compute standard errors and p value for cross-level difference between 'male' and 'countryB_male'.
#> Warning: No standard error for parameter 'NcasesValid'. Cannot compute standard errors and p value for cross-level difference between 'male' and 'countryC_male'.
#> Warning: No linking errors for parameters 'Ncases', 'NcasesValid'. Linking errors for these parameters will be defaulted to 0.
#> Warning: Found 35 missing linking errors for dependent variable 'traitLevel' and parameter(s) 'NcasesValid'. Assume linking error of 0 for these cases.
#> Warning: Found 35 missing linking errors for dependent variable 'traitLevel' and parameter(s) 'NcasesValid'. Assume linking error of 0 for these cases.
#> Warning: Found 35 missing linking errors for dependent variable 'traitLevel' and parameter(s) 'NcasesValid'. Assume linking error of 0 for these cases.
#> To date, only method 'old' is applicable for cross level differences in frequency tables.
#>
#> Trend group: '2010'
#> 4 analyse(s) overall according to: 'group.splits = 0 1 2'.
#>
#> analysis.number hierarchy.level dependent.var imp.var groups.divided.by group.differences.by
#> 1 0 traitLevel1 imp <NA>
#> 2 1 traitLevel1 imp country <NA>
#> 3 1 traitLevel1 imp sex sex
#> 4 2 traitLevel1 imp country + sex sex
#>
#> Assume unnested structure with 5 imputations.
#>
#>
#> Trend group: '2015'
#> 4 analyse(s) overall according to: 'group.splits = 0 1 2'.
#>
#> analysis.number hierarchy.level dependent.var imp.var groups.divided.by group.differences.by
#> 1 0 traitLevel1 imp <NA>
#> 2 1 traitLevel1 imp country <NA>
#> 3 1 traitLevel1 imp sex sex
#> 4 2 traitLevel1 imp country + sex sex
#>
#> Assume unnested structure with 5 imputations.
#>
#>
#> Trend group: '2020'
#> 4 analyse(s) overall according to: 'group.splits = 0 1 2'.
#>
#> analysis.number hierarchy.level dependent.var imp.var groups.divided.by group.differences.by
#> 1 0 traitLevel1 imp <NA>
#> 2 1 traitLevel1 imp country <NA>
#> 3 1 traitLevel1 imp sex sex
#> 4 2 traitLevel1 imp country + sex sex
#>
#> Assume unnested structure with 5 imputations.
#>
#>
#> Trend group: '2010'
#> 4 analyse(s) overall according to: 'group.splits = 0 1 2'.
#>
#> analysis.number hierarchy.level dependent.var imp.var groups.divided.by group.differences.by
#> 1 0 traitLevel2 imp <NA>
#> 2 1 traitLevel2 imp country <NA>
#> 3 1 traitLevel2 imp sex sex
#> 4 2 traitLevel2 imp country + sex sex
#>
#> Assume unnested structure with 5 imputations.
#>
#>
#> Trend group: '2015'
#> 4 analyse(s) overall according to: 'group.splits = 0 1 2'.
#>
#> analysis.number hierarchy.level dependent.var imp.var groups.divided.by group.differences.by
#> 1 0 traitLevel2 imp <NA>
#> 2 1 traitLevel2 imp country <NA>
#> 3 1 traitLevel2 imp sex sex
#> 4 2 traitLevel2 imp country + sex sex
#>
#> Assume unnested structure with 5 imputations.
#>
#>
#> Trend group: '2020'
#> 4 analyse(s) overall according to: 'group.splits = 0 1 2'.
#>
#> analysis.number hierarchy.level dependent.var imp.var groups.divided.by group.differences.by
#> 1 0 traitLevel2 imp <NA>
#> 2 1 traitLevel2 imp country <NA>
#> 3 1 traitLevel2 imp sex sex
#> 4 2 traitLevel2 imp country + sex sex
#>
#> Assume unnested structure with 5 imputations.
#>
#>
#> Trend group: '2010'
#> 4 analyse(s) overall according to: 'group.splits = 0 1 2'.
#>
#> analysis.number hierarchy.level dependent.var imp.var groups.divided.by group.differences.by
#> 1 0 traitLevel3 imp <NA>
#> 2 1 traitLevel3 imp country <NA>
#> 3 1 traitLevel3 imp sex sex
#> 4 2 traitLevel3 imp country + sex sex
#>
#> Assume unnested structure with 5 imputations.
#>
#>
#> Trend group: '2015'
#> 4 analyse(s) overall according to: 'group.splits = 0 1 2'.
#>
#> analysis.number hierarchy.level dependent.var imp.var groups.divided.by group.differences.by
#> 1 0 traitLevel3 imp <NA>
#> 2 1 traitLevel3 imp country <NA>
#> 3 1 traitLevel3 imp sex sex
#> 4 2 traitLevel3 imp country + sex sex
#>
#> Assume unnested structure with 5 imputations.
#>
#>
#> Trend group: '2020'
#> 4 analyse(s) overall according to: 'group.splits = 0 1 2'.
#>
#> analysis.number hierarchy.level dependent.var imp.var groups.divided.by group.differences.by
#> 1 0 traitLevel3 imp <NA>
#> 2 1 traitLevel3 imp country <NA>
#> 3 1 traitLevel3 imp sex sex
#> 4 2 traitLevel3 imp country + sex sex
#>
#> Assume unnested structure with 5 imputations.
#>
#>
#> Trend group: '2010'
#> 4 analyse(s) overall according to: 'group.splits = 0 1 2'.
#>
#> analysis.number hierarchy.level dependent.var imp.var groups.divided.by group.differences.by
#> 1 0 traitLevel4 imp <NA>
#> 2 1 traitLevel4 imp country <NA>
#> 3 1 traitLevel4 imp sex sex
#> 4 2 traitLevel4 imp country + sex sex
#>
#> Assume unnested structure with 5 imputations.
#>
#>
#> Trend group: '2015'
#> 4 analyse(s) overall according to: 'group.splits = 0 1 2'.
#>
#> analysis.number hierarchy.level dependent.var imp.var groups.divided.by group.differences.by
#> 1 0 traitLevel4 imp <NA>
#> 2 1 traitLevel4 imp country <NA>
#> 3 1 traitLevel4 imp sex sex
#> 4 2 traitLevel4 imp country + sex sex
#>
#> Assume unnested structure with 5 imputations.
#>
#>
#> Trend group: '2020'
#> 4 analyse(s) overall according to: 'group.splits = 0 1 2'.
#>
#> analysis.number hierarchy.level dependent.var imp.var groups.divided.by group.differences.by
#> 1 0 traitLevel4 imp <NA>
#> 2 1 traitLevel4 imp country <NA>
#> 3 1 traitLevel4 imp sex sex
#> 4 2 traitLevel4 imp country + sex sex
#>
#> Assume unnested structure with 5 imputations.
#>
#>
#> Trend group: '2010'
#> 4 analyse(s) overall according to: 'group.splits = 0 1 2'.
#>
#> analysis.number hierarchy.level dependent.var imp.var groups.divided.by group.differences.by
#> 1 0 traitLevel5 imp <NA>
#> 2 1 traitLevel5 imp country <NA>
#> 3 1 traitLevel5 imp sex sex
#> 4 2 traitLevel5 imp country + sex sex
#>
#> Assume unnested structure with 5 imputations.
#>
#>
#> Trend group: '2015'
#> 4 analyse(s) overall according to: 'group.splits = 0 1 2'.
#>
#> analysis.number hierarchy.level dependent.var imp.var groups.divided.by group.differences.by
#> 1 0 traitLevel5 imp <NA>
#> 2 1 traitLevel5 imp country <NA>
#> 3 1 traitLevel5 imp sex sex
#> 4 2 traitLevel5 imp country + sex sex
#>
#> Assume unnested structure with 5 imputations.
#>
#>
#> Trend group: '2020'
#> 4 analyse(s) overall according to: 'group.splits = 0 1 2'.
#>
#> analysis.number hierarchy.level dependent.var imp.var groups.divided.by group.differences.by
#> 1 0 traitLevel5 imp <NA>
#> 2 1 traitLevel5 imp country <NA>
#> 3 1 traitLevel5 imp sex sex
#> 4 2 traitLevel5 imp country + sex sex
#>
#> Assume unnested structure with 5 imputations.
#>
#> Warning: No standard error for parameter 'NcasesValid'. Cannot compute standard errors and p value for cross-level difference between 'wholeGroup' and 'countryA'.
#> Warning: No standard error for parameter 'NcasesValid'. Cannot compute standard errors and p value for cross-level difference between 'wholeGroup' and 'countryB'.
#> Warning: No standard error for parameter 'NcasesValid'. Cannot compute standard errors and p value for cross-level difference between 'wholeGroup' and 'countryC'.
#> Warning: No standard error for parameter 'NcasesValid'. Cannot compute standard errors and p value for cross-level difference between 'wholeGroup' and 'female'.
#> Warning: No standard error for parameter 'NcasesValid'. Cannot compute standard errors and p value for cross-level difference between 'wholeGroup' and 'male'.
#> Warning: No standard error for parameter 'NcasesValid'. Cannot compute standard errors and p value for cross-level difference between 'wholeGroup' and 'countryA_female'.
#> Warning: No standard error for parameter 'NcasesValid'. Cannot compute standard errors and p value for cross-level difference between 'wholeGroup' and 'countryA_male'.
#> Warning: No standard error for parameter 'NcasesValid'. Cannot compute standard errors and p value for cross-level difference between 'wholeGroup' and 'countryB_female'.
#> Warning: No standard error for parameter 'NcasesValid'. Cannot compute standard errors and p value for cross-level difference between 'wholeGroup' and 'countryB_male'.
#> Warning: No standard error for parameter 'NcasesValid'. Cannot compute standard errors and p value for cross-level difference between 'wholeGroup' and 'countryC_female'.
#> Warning: No standard error for parameter 'NcasesValid'. Cannot compute standard errors and p value for cross-level difference between 'wholeGroup' and 'countryC_male'.
#> Warning: No standard error for parameter 'NcasesValid'. Cannot compute standard errors and p value for cross-level difference between 'countryA' and 'countryA_female'.
#> Warning: No standard error for parameter 'NcasesValid'. Cannot compute standard errors and p value for cross-level difference between 'countryA' and 'countryA_male'.
#> Warning: No standard error for parameter 'NcasesValid'. Cannot compute standard errors and p value for cross-level difference between 'countryB' and 'countryB_female'.
#> Warning: No standard error for parameter 'NcasesValid'. Cannot compute standard errors and p value for cross-level difference between 'countryB' and 'countryB_male'.
#> Warning: No standard error for parameter 'NcasesValid'. Cannot compute standard errors and p value for cross-level difference between 'countryC' and 'countryC_female'.
#> Warning: No standard error for parameter 'NcasesValid'. Cannot compute standard errors and p value for cross-level difference between 'countryC' and 'countryC_male'.
#> Warning: No standard error for parameter 'NcasesValid'. Cannot compute standard errors and p value for cross-level difference between 'female' and 'countryA_female'.
#> Warning: No standard error for parameter 'NcasesValid'. Cannot compute standard errors and p value for cross-level difference between 'female' and 'countryB_female'.
#> Warning: No standard error for parameter 'NcasesValid'. Cannot compute standard errors and p value for cross-level difference between 'female' and 'countryC_female'.
#> Warning: No standard error for parameter 'NcasesValid'. Cannot compute standard errors and p value for cross-level difference between 'male' and 'countryA_male'.
#> Warning: No standard error for parameter 'NcasesValid'. Cannot compute standard errors and p value for cross-level difference between 'male' and 'countryB_male'.
#> Warning: No standard error for parameter 'NcasesValid'. Cannot compute standard errors and p value for cross-level difference between 'male' and 'countryC_male'.
#> Warning: No standard error for parameter 'NcasesValid'. Cannot compute standard errors and p value for cross-level difference between 'wholeGroup' and 'countryA'.
#> Warning: No standard error for parameter 'NcasesValid'. Cannot compute standard errors and p value for cross-level difference between 'wholeGroup' and 'countryB'.
#> Warning: No standard error for parameter 'NcasesValid'. Cannot compute standard errors and p value for cross-level difference between 'wholeGroup' and 'countryC'.
#> Warning: No standard error for parameter 'NcasesValid'. Cannot compute standard errors and p value for cross-level difference between 'wholeGroup' and 'female'.
#> Warning: No standard error for parameter 'NcasesValid'. Cannot compute standard errors and p value for cross-level difference between 'wholeGroup' and 'male'.
#> Warning: No standard error for parameter 'NcasesValid'. Cannot compute standard errors and p value for cross-level difference between 'wholeGroup' and 'countryA_female'.
#> Warning: No standard error for parameter 'NcasesValid'. Cannot compute standard errors and p value for cross-level difference between 'wholeGroup' and 'countryA_male'.
#> Warning: No standard error for parameter 'NcasesValid'. Cannot compute standard errors and p value for cross-level difference between 'wholeGroup' and 'countryB_female'.
#> Warning: No standard error for parameter 'NcasesValid'. Cannot compute standard errors and p value for cross-level difference between 'wholeGroup' and 'countryB_male'.
#> Warning: No standard error for parameter 'NcasesValid'. Cannot compute standard errors and p value for cross-level difference between 'wholeGroup' and 'countryC_female'.
#> Warning: No standard error for parameter 'NcasesValid'. Cannot compute standard errors and p value for cross-level difference between 'wholeGroup' and 'countryC_male'.
#> Warning: No standard error for parameter 'NcasesValid'. Cannot compute standard errors and p value for cross-level difference between 'countryA' and 'countryA_female'.
#> Warning: No standard error for parameter 'NcasesValid'. Cannot compute standard errors and p value for cross-level difference between 'countryA' and 'countryA_male'.
#> Warning: No standard error for parameter 'NcasesValid'. Cannot compute standard errors and p value for cross-level difference between 'countryB' and 'countryB_female'.
#> Warning: No standard error for parameter 'NcasesValid'. Cannot compute standard errors and p value for cross-level difference between 'countryB' and 'countryB_male'.
#> Warning: No standard error for parameter 'NcasesValid'. Cannot compute standard errors and p value for cross-level difference between 'countryC' and 'countryC_female'.
#> Warning: No standard error for parameter 'NcasesValid'. Cannot compute standard errors and p value for cross-level difference between 'countryC' and 'countryC_male'.
#> Warning: No standard error for parameter 'NcasesValid'. Cannot compute standard errors and p value for cross-level difference between 'female' and 'countryA_female'.
#> Warning: No standard error for parameter 'NcasesValid'. Cannot compute standard errors and p value for cross-level difference between 'female' and 'countryB_female'.
#> Warning: No standard error for parameter 'NcasesValid'. Cannot compute standard errors and p value for cross-level difference between 'female' and 'countryC_female'.
#> Warning: No standard error for parameter 'NcasesValid'. Cannot compute standard errors and p value for cross-level difference between 'male' and 'countryA_male'.
#> Warning: No standard error for parameter 'NcasesValid'. Cannot compute standard errors and p value for cross-level difference between 'male' and 'countryB_male'.
#> Warning: No standard error for parameter 'NcasesValid'. Cannot compute standard errors and p value for cross-level difference between 'male' and 'countryC_male'.
#> Warning: No standard error for parameter 'NcasesValid'. Cannot compute standard errors and p value for cross-level difference between 'wholeGroup' and 'countryA'.
#> Warning: No standard error for parameter 'NcasesValid'. Cannot compute standard errors and p value for cross-level difference between 'wholeGroup' and 'countryB'.
#> Warning: No standard error for parameter 'NcasesValid'. Cannot compute standard errors and p value for cross-level difference between 'wholeGroup' and 'countryC'.
#> Warning: No standard error for parameter 'NcasesValid'. Cannot compute standard errors and p value for cross-level difference between 'wholeGroup' and 'female'.
#> Warning: No standard error for parameter 'NcasesValid'. Cannot compute standard errors and p value for cross-level difference between 'wholeGroup' and 'male'.
#> Warning: No standard error for parameter 'NcasesValid'. Cannot compute standard errors and p value for cross-level difference between 'wholeGroup' and 'countryA_female'.
#> Warning: No standard error for parameter 'NcasesValid'. Cannot compute standard errors and p value for cross-level difference between 'wholeGroup' and 'countryA_male'.
#> Warning: No standard error for parameter 'NcasesValid'. Cannot compute standard errors and p value for cross-level difference between 'wholeGroup' and 'countryB_female'.
#> Warning: No standard error for parameter 'NcasesValid'. Cannot compute standard errors and p value for cross-level difference between 'wholeGroup' and 'countryB_male'.
#> Warning: No standard error for parameter 'NcasesValid'. Cannot compute standard errors and p value for cross-level difference between 'wholeGroup' and 'countryC_female'.
#> Warning: No standard error for parameter 'NcasesValid'. Cannot compute standard errors and p value for cross-level difference between 'wholeGroup' and 'countryC_male'.
#> Warning: No standard error for parameter 'NcasesValid'. Cannot compute standard errors and p value for cross-level difference between 'countryA' and 'countryA_female'.
#> Warning: No standard error for parameter 'NcasesValid'. Cannot compute standard errors and p value for cross-level difference between 'countryA' and 'countryA_male'.
#> Warning: No standard error for parameter 'NcasesValid'. Cannot compute standard errors and p value for cross-level difference between 'countryB' and 'countryB_female'.
#> Warning: No standard error for parameter 'NcasesValid'. Cannot compute standard errors and p value for cross-level difference between 'countryB' and 'countryB_male'.
#> Warning: No standard error for parameter 'NcasesValid'. Cannot compute standard errors and p value for cross-level difference between 'countryC' and 'countryC_female'.
#> Warning: No standard error for parameter 'NcasesValid'. Cannot compute standard errors and p value for cross-level difference between 'countryC' and 'countryC_male'.
#> Warning: No standard error for parameter 'NcasesValid'. Cannot compute standard errors and p value for cross-level difference between 'female' and 'countryA_female'.
#> Warning: No standard error for parameter 'NcasesValid'. Cannot compute standard errors and p value for cross-level difference between 'female' and 'countryB_female'.
#> Warning: No standard error for parameter 'NcasesValid'. Cannot compute standard errors and p value for cross-level difference between 'female' and 'countryC_female'.
#> Warning: No standard error for parameter 'NcasesValid'. Cannot compute standard errors and p value for cross-level difference between 'male' and 'countryA_male'.
#> Warning: No standard error for parameter 'NcasesValid'. Cannot compute standard errors and p value for cross-level difference between 'male' and 'countryB_male'.
#> Warning: No standard error for parameter 'NcasesValid'. Cannot compute standard errors and p value for cross-level difference between 'male' and 'countryC_male'.
#> Warning: No linking errors for parameters 'Ncases', 'NcasesValid'. Linking errors for these parameters will be defaulted to 0.
#> Warning: Found 35 missing linking errors for dependent variable 'traitLevel' and parameter(s) 'NcasesValid'. Assume linking error of 0 for these cases.
#> Warning: Found 35 missing linking errors for dependent variable 'traitLevel' and parameter(s) 'NcasesValid'. Assume linking error of 0 for these cases.
#> Warning: Found 35 missing linking errors for dependent variable 'traitLevel' and parameter(s) 'NcasesValid'. Assume linking error of 0 for these cases.
freqs2<- do.call("rbind", freqs2)
################################################################################
### Example 7: Linking and equating for multiple models (III) ###
################################################################################
# For the purpose of illustration, this example repeats analyses of example 6,
# using a 2pl estimation now. Example 7 demonstrates routines without trend
# estimation.
# Preparation: assume time of measurement 't1' corresponds to the year 2003.
datT1<- reshape2::dcast(subset ( trends, year == 2010),
formula = idstud+sex+country+language+ses~item, value.var="value")
# First step: item calibration in separate unidimensional models for each domain
# split 2 models. Note: not all items of the Q matrix are present in the data.
# Items which occur only in the Q matrix will be ignored.
modsT1<- splitModels ( qMatrix = qMat, nCores = 1)
#> --------------------------------
#> splitModels: generating 2 models
#> ..
#> see <returned>$models
#> number of cores: 1
#> --------------------------------
# lets specify a 2pl model with constraints: a common discrimination for all
# items belonging to the same task
slopes<- data.frame ( variable = qMat[,"item"],
slope = as.numeric(as.factor(substr(qMat[,"item"],1,3))))
# prepare 2pl model
defT1 <- defineModel(dat = datT1, id = "idstud", check.for.linking = TRUE,
splittedModels = modsT1, irtmodel = "2PL.groups", est.slopegroups = slopes,
software = "tam")
#>
#> Specification of 'qMatrix' and 'person.groups' results in 2 model(s).
#> Warning: Model split preparation for model No. 1, model name domainlistening: 83 items from 134 items listed the Q matrix not found in data:
#> ℹ 'T12_11', 'T15_17', 'T15_18', 'T13_18', 'T19_06', 'T19_08', 'T19_05', 'T19_11', 'T19_01', 'T19_02', 'T19_30', 'T19_03', 'T19_07', 'T19_04', 'T19_09', 'T21_11', 'T18_01', 'T21_01', 'T18_05', 'T21_12', 'T18_13', 'T21_10', 'T18_11', 'T21_05', 'T21_13', 'T18_08', 'T18_07', 'T21_09', 'T18_04', 'T18_02', 'T21_02', 'T21_06', 'T18_09', 'T21_04', 'T18_12', 'T21_14', 'T21_03', 'T18_10', 'T30_06', 'T29_04', 'T29_02', 'T30_02', 'T30_03', 'T30_01', 'T29_05', 'T29_03', 'T28_08', 'T28_09', 'T28_02', 'T28_03', 'T28_04', 'T28_01', 'T28_10', 'T28_06', 'T20_05', 'T20_09', 'T20_30', 'T20_08', 'T20_12', 'T20_06', 'T20_07', 'T20_02', 'T20_13', 'T20_03', 'T20_01', 'T20_04', 'T20_11', 'T20_10', 'T32_04', 'T31_03', 'T31_01', 'T31_04', 'T32_03', 'T32_02', 'T32_01', 'T17_02', 'T17_09', 'T17_10', 'T17_04', 'T17_03', 'T17_08', 'T17_06', 'T17_01'
#> Warning: Model split preparation for model No. 2, model name domainreading: 69 items from 149 items listed the Q matrix not found in data:
#> ℹ 'T09_12', 'T01_08', 'T10_10', 'T04_09', 'T04_08', 'T11_0X', 'T08_09', 'T08_08', 'T03_09', 'T06_1X', 'T26_03', 'T26_05', 'T26_07', 'T26_09', 'T26_06', 'T26_10', 'T26_08', 'T26_02', 'T26_04', 'T27_01', 'T27_05', 'T27_07', 'T27_06', 'T27_04', 'T27_02', 'T27_03', 'T24_05', 'T24_08', 'T24_02', 'T24_10', 'T24_06', 'T24_24', 'T24_25', 'T24_09', 'T24_21', 'T24_23', 'T23_10', 'T23_13', 'T23_09', 'T23_15', 'T23_03', 'T23_04', 'T23_14', 'T23_07', 'T23_08', 'T23_02', 'T23_06', 'T23_05', 'T25_05', 'T25_22', 'T22_03', 'T25_06', 'T22_12', 'T25_09', 'T25_10', 'T25_08', 'T22_11', 'T22_02', 'T25_11', 'T25_07', 'T25_14', 'T25_12', 'T25_04', 'T22_08', 'T25_23', 'T22_10', 'T22_06', 'T25_13', 'T22_01'
#>
#>
#> =========================================
#> Model No. 1
#> Model name: domainlistening
#> Number of items: 51
#> Number of persons: 4476
#> Number of dimensions: 1
#> =========================================
#>
#> Following 83 item(s) missed in data frame will be removed from Q matrix:
#> T12_11, T15_17, T15_18, T13_18, T19_06, T19_08, T19_05, T19_11, T19_01, T19_02, T19_30, T19_03, T19_07, T19_04, T19_09, T21_11, T18_01, T21_01, T18_05, T21_12, T18_13, T21_10, T18_11, T21_05, T21_13, T18_08, T18_07, T21_09, T18_04, T18_02, T21_02, T21_06, T18_09, T21_04, T18_12, T21_14, T21_03, T18_10, T30_06, T29_04, T29_02, T30_02, T30_03, T30_01, T29_05, T29_03, T28_08, T28_09, T28_02, T28_03, T28_04, T28_01, T28_10, T28_06, T20_05, T20_09, T20_30, T20_08, T20_12, T20_06, T20_07, T20_02, T20_13, T20_03, T20_01, T20_04, T20_11, T20_10, T32_04, T31_03, T31_01, T31_04, T32_03, T32_02, T32_01, T17_02, T17_09, T17_10, T17_04, T17_03, T17_08, T17_06, T17_01
#> 76 subject(s) do not solve any item:
#> P00009 (6 false), P00864 (6 false), P02759 (21 false) ...
#> 55 subject(s) solved each item: P00099 (10 correct), P00404 (10 correct), P02659 (11 correct) ...
#> Dataset is completely linked.
#> 'gauss' has been chosen for estimation method. Number of nodes was not explicitly specified. Set nodes to 20.
#> Q matrix specifies 1 dimension(s).
#> Following 232 Items in design matrix for item groups with common slopes ('est.slopegroups') which are not in dataset:
#> T07_08, T02_02, T07_01, T02_06, T07_05, T07_06, T02_07, T07_02, T07_04, T02_03, T07_09, T02_01, T02_04, T07_03, T02_05, T07_07, T07_10, T09_03, T09_05, T01_03, T09_02, T01_06, T01_01, T09_10, T01_02, T09_06, T01_04, T09_09, T09_11, T01_05, T09_08, T09_07, T09_04, T01_07, T12_11, T09_12, T01_08, T03_07, T03_01, T03_08, T03_05, T03_06, T03_03, T10_08, T11_06, T11_03, T11_01, T04_02, T04_01, T04_06, T04_04, T11_04, T10_01, T11_08, T10_02, T10_09, T10_07, T11_02, T04_05, T04_03, T11_05, T10_06, T11_10, T11_09, T04_07, T15_17, T15_18, T13_18, T10_10, T04_09, T04_08, T11_0X, T06_05, T05_03, T05_06, T05_05, T06_02, T06_04, T06_03, T05_04, T05_02, T05_01, T06_01, T08_04, T08_06, T08_03, T08_07, T08_02, T08_05, T08_01, T08_09, T08_08, T03_09, T06_1X, T19_06, T19_08, T19_05, T19_11, T19_01, T19_02, T19_30, T19_03, T19_07, T19_04, T19_09, T26_03, T26_05, T26_07, T26_09, T26_06, T26_10, T26_08, T26_02, T26_04, T21_11, T18_01, T21_01, T18_05, T21_12, T18_13, T21_10, T18_11, T21_05, T21_13, T18_08, T18_07, T21_09, T18_04, T18_02, T21_02, T21_06, T18_09, T21_04, T18_12, T21_14, T21_03, T18_10, T27_01, T27_05, T27_07, T27_06, T27_04, T27_02, T27_03, T30_06, T29_04, T29_02, T30_02, T30_03, T30_01, T29_05, T29_03, T24_05, T24_08, T24_02, T24_10, T24_06, T24_24, T24_25, T24_09, T24_21, T24_23, T28_08, T28_09, T28_02, T28_03, T28_04, T28_01, T28_10, T28_06, T23_10, T23_13, T23_09, T23_15, T23_03, T23_04, T23_14, T23_07, T23_08, T23_02, T23_06, T23_05, T25_05, T25_22, T22_03, T25_06, T22_12, T25_09, T25_10, T25_08, T22_11, T22_02, T25_11, T25_07, T25_14, T25_12, T25_04, T22_08, T25_23, T22_10, T22_06, T25_13, T22_01, T20_05, T20_09, T20_30, T20_08, T20_12, T20_06, T20_07, T20_02, T20_13, T20_03, T20_01, T20_04, T20_11, T20_10, T32_04, T31_03, T31_01, T31_04, T32_03, T32_02, T32_01, T17_02, T17_09, T17_10, T17_04, T17_03, T17_08, T17_06, T17_01
#> Remove these item(s) from design matrix.
#>
#>
#> =======================================
#> Model No. 2
#> Model name: domainreading
#> Number of items: 80
#> Number of persons: 4476
#> Number of dimensions: 1
#> =======================================
#>
#> Following 69 item(s) missed in data frame will be removed from Q matrix:
#> T09_12, T01_08, T10_10, T04_09, T04_08, T11_0X, T08_09, T08_08, T03_09, T06_1X, T26_03, T26_05, T26_07, T26_09, T26_06, T26_10, T26_08, T26_02, T26_04, T27_01, T27_05, T27_07, T27_06, T27_04, T27_02, T27_03, T24_05, T24_08, T24_02, T24_10, T24_06, T24_24, T24_25, T24_09, T24_21, T24_23, T23_10, T23_13, T23_09, T23_15, T23_03, T23_04, T23_14, T23_07, T23_08, T23_02, T23_06, T23_05, T25_05, T25_22, T22_03, T25_06, T22_12, T25_09, T25_10, T25_08, T22_11, T22_02, T25_11, T25_07, T25_14, T25_12, T25_04, T22_08, T25_23, T22_10, T22_06, T25_13, T22_01
#> 17 subject(s) do not solve any item:
#> P00173 (6 false), P01137 (7 false), P02863 (23 false) ...
#> 97 subject(s) solved each item: P00078 (6 correct), P00389 (10 correct), P03338 (27 correct) ...
#> Dataset is completely linked.
#> 'gauss' has been chosen for estimation method. Number of nodes was not explicitly specified. Set nodes to 20.
#> Q matrix specifies 1 dimension(s).
#> Following 203 Items in design matrix for item groups with common slopes ('est.slopegroups') which are not in dataset:
#> T12_06, T12_07, T14_03, T14_06, T12_01, T12_03, T12_08, T12_10, T14_02, T14_05, T12_02, T12_04, T14_04, T12_05, T14_01, T12_09, T12_11, T09_12, T01_08, T13_15, T15_03, T13_04, T13_07, T13_16, T13_01, T15_02, T15_01, T15_15, T15_16, T15_09, T13_12, T13_13, T15_14, T13_08, T13_06, T15_12, T13_10, T13_17, T15_11, T13_05, T13_09, T15_10, T15_13, T15_17, T15_18, T13_18, T10_10, T04_09, T04_08, T11_0X, T16_08, T16_09, T16_11, T16_13, T16_01, T16_04, T16_10, T16_02, T16_12, T16_06, T16_07, T08_09, T08_08, T03_09, T06_1X, T19_06, T19_08, T19_05, T19_11, T19_01, T19_02, T19_30, T19_03, T19_07, T19_04, T19_09, T26_03, T26_05, T26_07, T26_09, T26_06, T26_10, T26_08, T26_02, T26_04, T21_11, T18_01, T21_01, T18_05, T21_12, T18_13, T21_10, T18_11, T21_05, T21_13, T18_08, T18_07, T21_09, T18_04, T18_02, T21_02, T21_06, T18_09, T21_04, T18_12, T21_14, T21_03, T18_10, T27_01, T27_05, T27_07, T27_06, T27_04, T27_02, T27_03, T30_06, T29_04, T29_02, T30_02, T30_03, T30_01, T29_05, T29_03, T24_05, T24_08, T24_02, T24_10, T24_06, T24_24, T24_25, T24_09, T24_21, T24_23, T28_08, T28_09, T28_02, T28_03, T28_04, T28_01, T28_10, T28_06, T23_10, T23_13, T23_09, T23_15, T23_03, T23_04, T23_14, T23_07, T23_08, T23_02, T23_06, T23_05, T25_05, T25_22, T22_03, T25_06, T22_12, T25_09, T25_10, T25_08, T22_11, T22_02, T25_11, T25_07, T25_14, T25_12, T25_04, T22_08, T25_23, T22_10, T22_06, T25_13, T22_01, T20_05, T20_09, T20_30, T20_08, T20_12, T20_06, T20_07, T20_02, T20_13, T20_03, T20_01, T20_04, T20_11, T20_10, T32_04, T31_03, T31_01, T31_04, T32_03, T32_02, T32_01, T17_02, T17_09, T17_10, T17_04, T17_03, T17_08, T17_06, T17_01
#> Remove these item(s) from design matrix.
# run 2 models
runT1 <- runModel(defT1)
# get the results
resT1 <- getResults(runT1)
#> Getting standard errors with the tam.se function: 0.3 secs
#> Getting WLEs calling tam.wle from getTamWles: 0.4 secs
#> |*****|
#> |-----|
#> Getting PVs calling tam.pv from getTamPVs: 0.9 secs
#> Getting Q3 statistic calling tam.modelfit from getTamQ3: 0.8 secs
#> Getting standard errors with the tam.se function: 0.6 secs
#> Getting WLEs calling tam.wle from getTamWles: 0.9 secs
#> |*****|
#> |-----|
#> Getting PVs calling tam.pv from getTamPVs: 0.9 secs
#> Getting Q3 statistic calling tam.modelfit from getTamQ3: 1.9 secs
# extract item parameters from the 'results' object
itemT1<- itemFromRes(resT1)
# Second step: drawing plausible values. Two-dimensional model is specified for
# each person group with fixed item parameters. Moreover, a latent regression
# model is used (in the actual 'Laendervergleich', regressors are principal
# components).
# define person grouping
# Running second step: split models according to person groups
# ('all.persons' must be FALSE, otherwise the whole group would be treated as
# a separate distinct group.)
modT1P<- splitModels ( person.groups = datT1[,c("idstud", "country")], all.persons = FALSE, nCores = 1)
#> --------------------------------
#> splitModels: generating 3 models
#> ...
#> see <returned>$models
#> number of cores: 1
#> --------------------------------
# define the 3 country-specific 2-dimensional models, specifying latent regression
# model and fixed item and fixed slope parameters.
defT1P<- defineModel(dat = datT1, items = itemT1[,"item"], id = "idstud", irtmodel = "2PL",
check.for.linking = TRUE, splittedModels = modT1P, qMatrix = qMat,
anchor = itemT1[,c("item", "est")], fixSlopeMat = itemT1[,c("item", "estSlope")],
HG.var = c("ses", "sex", "language"), software = "tam")
#>
#> Specification of 'qMatrix' and 'person.groups' results in 3 model(s).
#>
#>
#> ==========================================
#> Model No. 1
#> Model name: country.countryC
#> Number of items: 131
#> Number of persons: 1569
#> Number of dimensions: 2
#> ==========================================
#>
#> Following 152 item(s) missed in data frame will be removed from Q matrix:
#> T12_11, T09_12, T01_08, T15_17, T15_18, T13_18, T10_10, T04_09, T04_08, T11_0X, T08_09, T08_08, T03_09, T06_1X, T19_06, T19_08, T19_05, T19_11, T19_01, T19_02, T19_30, T19_03, T19_07, T19_04, T19_09, T26_03, T26_05, T26_07, T26_09, T26_06, T26_10, T26_08, T26_02, T26_04, T21_11, T18_01, T21_01, T18_05, T21_12, T18_13, T21_10, T18_11, T21_05, T21_13, T18_08, T18_07, T21_09, T18_04, T18_02, T21_02, T21_06, T18_09, T21_04, T18_12, T21_14, T21_03, T18_10, T27_01, T27_05, T27_07, T27_06, T27_04, T27_02, T27_03, T30_06, T29_04, T29_02, T30_02, T30_03, T30_01, T29_05, T29_03, T24_05, T24_08, T24_02, T24_10, T24_06, T24_24, T24_25, T24_09, T24_21, T24_23, T28_08, T28_09, T28_02, T28_03, T28_04, T28_01, T28_10, T28_06, T23_10, T23_13, T23_09, T23_15, T23_03, T23_04, T23_14, T23_07, T23_08, T23_02, T23_06, T23_05, T25_05, T25_22, T22_03, T25_06, T22_12, T25_09, T25_10, T25_08, T22_11, T22_02, T25_11, T25_07, T25_14, T25_12, T25_04, T22_08, T25_23, T22_10, T22_06, T25_13, T22_01, T20_05, T20_09, T20_30, T20_08, T20_12, T20_06, T20_07, T20_02, T20_13, T20_03, T20_01, T20_04, T20_11, T20_10, T32_04, T31_03, T31_01, T31_04, T32_03, T32_02, T32_01, T17_02, T17_09, T17_10, T17_04, T17_03, T17_08, T17_06, T17_01
#> Background variable(s) 'sex', 'language' of class
#> 'factor', 'factor' will be converted to indicator variables.
#> Variable 'sex' was converted to 1 indicator(s) with name(s) 'sexmale'.
#> Variable 'language' was converted to 2 indicator(s) with name(s) 'languagenativeAndOther', 'languageother'.
#> 5 subject(s) solved each item: P00106 (17 correct), P00939 (17 correct), P00393 (20 correct) ...
#> Dataset is completely linked.
#> 'gauss' has been chosen for estimation method. Number of nodes was not explicitly specified. Set nodes to 20.
#> 131 common items found in 'anchor' list and data frame.
#> Q matrix specifies 2 dimension(s).
#> Warning: To date, fixing slopes only works for dichotomous unidimensional or between-item multidimensional models.
#>
#>
#> ==========================================
#> Model No. 2
#> Model name: country.countryA
#> Number of items: 131
#> Number of persons: 1598
#> Number of dimensions: 2
#> ==========================================
#>
#> Following 152 item(s) missed in data frame will be removed from Q matrix:
#> T12_11, T09_12, T01_08, T15_17, T15_18, T13_18, T10_10, T04_09, T04_08, T11_0X, T08_09, T08_08, T03_09, T06_1X, T19_06, T19_08, T19_05, T19_11, T19_01, T19_02, T19_30, T19_03, T19_07, T19_04, T19_09, T26_03, T26_05, T26_07, T26_09, T26_06, T26_10, T26_08, T26_02, T26_04, T21_11, T18_01, T21_01, T18_05, T21_12, T18_13, T21_10, T18_11, T21_05, T21_13, T18_08, T18_07, T21_09, T18_04, T18_02, T21_02, T21_06, T18_09, T21_04, T18_12, T21_14, T21_03, T18_10, T27_01, T27_05, T27_07, T27_06, T27_04, T27_02, T27_03, T30_06, T29_04, T29_02, T30_02, T30_03, T30_01, T29_05, T29_03, T24_05, T24_08, T24_02, T24_10, T24_06, T24_24, T24_25, T24_09, T24_21, T24_23, T28_08, T28_09, T28_02, T28_03, T28_04, T28_01, T28_10, T28_06, T23_10, T23_13, T23_09, T23_15, T23_03, T23_04, T23_14, T23_07, T23_08, T23_02, T23_06, T23_05, T25_05, T25_22, T22_03, T25_06, T22_12, T25_09, T25_10, T25_08, T22_11, T22_02, T25_11, T25_07, T25_14, T25_12, T25_04, T22_08, T25_23, T22_10, T22_06, T25_13, T22_01, T20_05, T20_09, T20_30, T20_08, T20_12, T20_06, T20_07, T20_02, T20_13, T20_03, T20_01, T20_04, T20_11, T20_10, T32_04, T31_03, T31_01, T31_04, T32_03, T32_02, T32_01, T17_02, T17_09, T17_10, T17_04, T17_03, T17_08, T17_06, T17_01
#> Background variable(s) 'sex', 'language' of class
#> 'factor', 'factor' will be converted to indicator variables.
#> Variable 'sex' was converted to 1 indicator(s) with name(s) 'sexmale'.
#> Variable 'language' was converted to 2 indicator(s) with name(s) 'languagenativeAndOther', 'languageother'.
#> Dataset is completely linked.
#> 'gauss' has been chosen for estimation method. Number of nodes was not explicitly specified. Set nodes to 20.
#> 131 common items found in 'anchor' list and data frame.
#> Q matrix specifies 2 dimension(s).
#> Warning: To date, fixing slopes only works for dichotomous unidimensional or between-item multidimensional models.
#>
#>
#> ==========================================
#> Model No. 3
#> Model name: country.countryB
#> Number of items: 131
#> Number of persons: 1309
#> Number of dimensions: 2
#> ==========================================
#>
#> Following 152 item(s) missed in data frame will be removed from Q matrix:
#> T12_11, T09_12, T01_08, T15_17, T15_18, T13_18, T10_10, T04_09, T04_08, T11_0X, T08_09, T08_08, T03_09, T06_1X, T19_06, T19_08, T19_05, T19_11, T19_01, T19_02, T19_30, T19_03, T19_07, T19_04, T19_09, T26_03, T26_05, T26_07, T26_09, T26_06, T26_10, T26_08, T26_02, T26_04, T21_11, T18_01, T21_01, T18_05, T21_12, T18_13, T21_10, T18_11, T21_05, T21_13, T18_08, T18_07, T21_09, T18_04, T18_02, T21_02, T21_06, T18_09, T21_04, T18_12, T21_14, T21_03, T18_10, T27_01, T27_05, T27_07, T27_06, T27_04, T27_02, T27_03, T30_06, T29_04, T29_02, T30_02, T30_03, T30_01, T29_05, T29_03, T24_05, T24_08, T24_02, T24_10, T24_06, T24_24, T24_25, T24_09, T24_21, T24_23, T28_08, T28_09, T28_02, T28_03, T28_04, T28_01, T28_10, T28_06, T23_10, T23_13, T23_09, T23_15, T23_03, T23_04, T23_14, T23_07, T23_08, T23_02, T23_06, T23_05, T25_05, T25_22, T22_03, T25_06, T22_12, T25_09, T25_10, T25_08, T22_11, T22_02, T25_11, T25_07, T25_14, T25_12, T25_04, T22_08, T25_23, T22_10, T22_06, T25_13, T22_01, T20_05, T20_09, T20_30, T20_08, T20_12, T20_06, T20_07, T20_02, T20_13, T20_03, T20_01, T20_04, T20_11, T20_10, T32_04, T31_03, T31_01, T31_04, T32_03, T32_02, T32_01, T17_02, T17_09, T17_10, T17_04, T17_03, T17_08, T17_06, T17_01
#> Background variable(s) 'sex', 'language' of class
#> 'factor', 'factor' will be converted to indicator variables.
#> Variable 'sex' was converted to 1 indicator(s) with name(s) 'sexmale'.
#> Variable 'language' was converted to 2 indicator(s) with name(s) 'languagenativeAndOther', 'languageother'.
#> Dataset is completely linked.
#> 'gauss' has been chosen for estimation method. Number of nodes was not explicitly specified. Set nodes to 20.
#> 131 common items found in 'anchor' list and data frame.
#> Q matrix specifies 2 dimension(s).
#> Warning: To date, fixing slopes only works for dichotomous unidimensional or between-item multidimensional models.
# run the 3 models
runT1P<- runModel(defT1P)
# get the results
resT1P<- getResults(runT1P, Q3 = FALSE)
#> Getting standard errors with the tam.se function: 33.7 secs
#> Getting infit parameters calling tam.fit from getTamInfit: 2 secs
#> Getting WLEs calling tam.wle from getTamWles: 0.8 secs
#> |*****|
#> |-----|
#> Getting PVs calling tam.pv from getTamPVs: 0.4 secs
#> Getting standard errors with the tam.se function: 39.4 secs
#> Getting infit parameters calling tam.fit from getTamInfit: 2.5 secs
#> Getting WLEs calling tam.wle from getTamWles: 0.5 secs
#> |*****|
#> |-----|
#> Getting PVs calling tam.pv from getTamPVs: 0.5 secs
#> Getting standard errors with the tam.se function: 33.2 secs
#> Getting infit parameters calling tam.fit from getTamInfit: 2 secs
#> Getting WLEs calling tam.wle from getTamWles: 0.4 secs
#> |*****|
#> |-----|
#> Getting PVs calling tam.pv from getTamPVs: 0.4 secs
################################################################################
### Example 8: anchored partial credit model excluding linking DIF (TAM) ###
################################################################################
# load partial credit long format data
data(reading)
# transform into wide format
datW <- reshape2::dcast(reading[which(reading[,"type"] != "iglu"),],idstud+sex+language+country~item, value.var = "valueSum")
# only some items are partial credit, which ones?
pc.it<- reading[which(reading[,"valueSum"] > 1),"item"] |> unique()
# give values of partial credit items
# few observation of 1-category for item D223143 ... may cause convergence trouble
lapply(datW[,pc.it], FUN = table)
#> $D204013
#>
#> 0 1 2 3 4
#> 24 42 101 263 324
#>
#> $D205143
#>
#> 0 1 2 3 4 5
#> 7 4 31 53 171 466
#>
#> $D201113
#>
#> 0 1 2 3 4
#> 15 113 141 170 104
#>
#> $D225113
#>
#> 0 1 2 3 4
#> 35 75 153 207 78
#>
#> $D224053
#>
#> 0 1 2 3 4
#> 15 25 94 172 225
#>
#> $D025033
#>
#> 0 1 2 3 4
#> 13 39 74 178 233
#>
#> $D223143
#>
#> 0 1 2 3 4 5
#> 23 6 41 100 106 254
#>
# combine the two lowest categories to avoid categories with too few observations
datW[,"D205143"] <- car::recode(datW[,"D205143"], "1=0; 2=1; 3=2; 4=3; 5=4")
# partial credit model ... we consider the female subgroup to be the reference population
# (norm population) that defines the scale and reference item parameters
dFema<- datW[which(datW[,"sex"] == "female"),] ### data for females
lapply(dFema[,pc.it], FUN = table) ### very few observations for some categories ... may cause convergence trouble
#> $D204013
#>
#> 0 1 2 3 4
#> 6 15 42 127 165
#>
#> $D205143
#>
#> 0 1 2 3 4
#> 3 17 20 97 253
#>
#> $D201113
#>
#> 0 1 2 3 4
#> 4 62 66 85 56
#>
#> $D225113
#>
#> 0 1 2 3 4
#> 14 43 90 107 43
#>
#> $D224053
#>
#> 0 1 2 3 4
#> 4 7 34 90 119
#>
#> $D025033
#>
#> 0 1 2 3 4
#> 3 19 39 100 120
#>
#> $D223143
#>
#> 0 1 2 3 4 5
#> 12 2 21 56 68 159
#>
def1 <- defineModel(dat=dFema, items = -c(1:4), id=1, irtmodel = "PCM", software="tam", nodes = 21)
#> 7 variable(s) are not strictly dichotomous with 0/1 ... Expect a rating scale model or partial credit model.
#> Items(s) 'D025033', 'D201113', 'D204013', 'D205143', 'D224053', 'D225113': 0, 1, 2, 3, 4
#> Items(s) 'D223143': 0, 1, 2, 3, 4, 5
#> 2 subject(s) do not solve any item:
#> P1296 (24 false), P1299 (24 false), P1299 (24 false) ...
#> 12 subject(s) solved each item: P0081 (12 correct), P0869 (13 correct), P1047 (29 correct) ...
#> Dataset is completely linked.
#> Q matrix specifies 1 dimension(s).
run1 <- runModel(def1)
# plot polytomous item "D205143": category 3 isn't necessary
ind1 <- grep("D205143", run1$item$item) ### where is item "D205143"
foo1 <- capture.output(plot(run1, items = ind1, type="items", export=FALSE, low=-6, high=6))
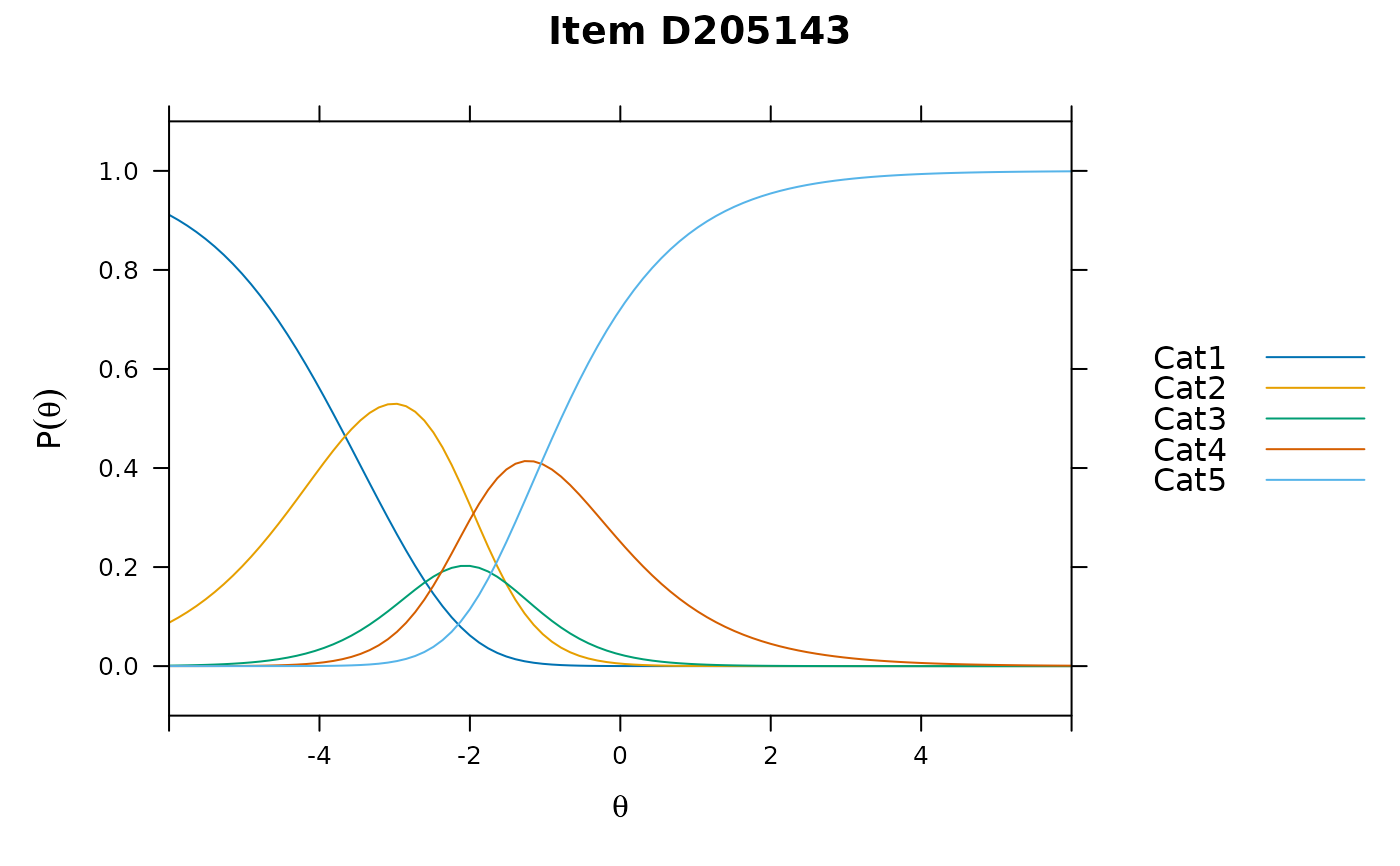 res1 <- getResults(run1)
#> Getting standard errors with the tam.se function: 0.7 secs
#> Getting infit parameters calling tam.fit from getTamInfit: 0.2 secs
#> Getting WLEs calling tam.wle from getTamWles: 0.3 secs
#> |*****|
#> |-----|
#> Getting PVs calling tam.pv from getTamPVs: 0.3 secs
#> Getting Q3 statistic calling tam.modelfit from getTamQ3: 0.9 secs
it1 <- itemFromRes(res1)
# males are focus group: initial free estimation of item parameters
def2 <- defineModel(dat=datW[which(datW[,"sex"] == "male"),], items = -c(1:4), id=1, irtmodel = "PCM",software="tam")
#> 7 variable(s) are not strictly dichotomous with 0/1 ... Expect a rating scale model or partial credit model.
#> Items(s) 'D025033', 'D201113', 'D204013', 'D205143', 'D224053', 'D225113': 0, 1, 2, 3, 4
#> Items(s) 'D223143': 0, 1, 2, 3, 4, 5
#> 4 subject(s) do not solve any item:
#> P0007 (12 false), P0107 (12 false), P0837 (13 false) ...
#> 8 subject(s) solved each item: P0968 (11 correct), P0839 (13 correct), P1233 (24 correct) ...
#> Dataset is completely linked.
#> 'gauss' has been chosen for estimation method. Number of nodes was not explicitly specified. Set nodes to 20.
#> Q matrix specifies 1 dimension(s).
run2 <- runModel(def2)
res2 <- getResults(run2)
#> Getting standard errors with the tam.se function: 0.7 secs
#> Getting infit parameters calling tam.fit from getTamInfit: 0.2 secs
#> Getting WLEs calling tam.wle from getTamWles: 0.3 secs
#> |*****|
#> |-----|
#> Getting PVs calling tam.pv from getTamPVs: 0.3 secs
#> Getting Q3 statistic calling tam.modelfit from getTamQ3: 0.9 secs
it2 <- itemFromRes(res2)
# link males to females ... males perform worse
# 10 items with linking dif identified
eq <- equat1pl(results = res2, prmNorm = it1, item = "item", cat="category", value = "est", difBound=.64, iterativ = TRUE)
#> Found 1 model(s).
#> Equating is executed for each dimension in each model separately.
#>
#> ====================================================================================================
#>
#> Model No. 1
#> Model name: not_specified
#> Number of dimension(s): 1
#> Name(s) of dimension(s): Dim1
#> Number of linking items: 128
#> Linking method: Mean.Mean
#>
#> Dimension 'Dim1': 10 of 128 items with linking |DIF| > 0.64 identified.
#> Iteration 1: Exclude item 'D205143_Cat1'.
#> Iteration 2: Exclude item 'D223143_Cat1'.
#> Iteration 3: Exclude item 'D204043_Cat1'.
#> Iteration 4: Exclude item 'D025033_Cat1'.
#> Iteration 5: Exclude item 'D201113_Cat1'.
#> Iteration 6: Exclude item 'D034053_Cat1'.
#> Iteration 7: Exclude item 'D224023_Cat1'.
#> Iteration 8: Exclude item 'D205143_Cat3'.
#> Iteration 9: Exclude item 'D221093_Cat1'.
#> Iteration 10: Exclude item 'D204033_Cat1'.
#> Iteration 11: Exclude item 'D224053_Cat3'.
#>
#> method iter itemExcluded DIF.excluded linking.constant linkerror
#> 1 iterativ 0 -0.221 0.034
#> 2 iterativ 1 D205143_Cat1 1.231 -0.211 0.033
#> 3 iterativ 2 D223143_Cat1 -1.055 -0.220 0.032
#> 4 iterativ 3 D204043_Cat1 1 -0.212 0.031
#> 5 iterativ 4 D025033_Cat1 0.996 -0.204 0.030
#> 6 iterativ 5 D201113_Cat1 0.898 -0.197 0.030
#> 7 iterativ 6 D034053_Cat1 0.849 -0.190 0.029
#> 8 iterativ 7 D224023_Cat1 0.807 -0.183 0.028
#> 9 iterativ 8 D205143_Cat3 0.795 -0.177 0.028
#> 10 iterativ 9 D221093_Cat1 -0.696 -0.183 0.028
#> 11 iterativ 10 D204033_Cat1 -0.668 -0.188 0.027
#> 12 iterativ 11 D224053_Cat3 0.658 -0.183 0.027
#>
# re-calibrate males with anchoring: use the DIF-cleaned set of anchor parameters for calibrating males on the reference scale
# variant 1: use the original item parameters for females and exclude linking dif items (it1_cleaned)
weg <- eq[["items"]][["not_specified"]][["Dim1"]][["info"]][-1,"itemExcluded"]
weg <- eatTools::whereAre(weg, paste(it1[,"item"], it1[,"category"], sep="_"))
#> Found 11 elements.
it1C <- it1[-weg,]
def3A<- defineModel(dat=datW[which(datW[,"sex"] == "male"),], items = -c(1:4), id=1, irtmodel = "PCM",
anchor = it1C, itemCol = "item", valueCol = "est", catCol = "category", software="tam")
#> 7 variable(s) are not strictly dichotomous with 0/1 ... Expect a rating scale model or partial credit model.
#> Items(s) 'D025033', 'D201113', 'D204013', 'D205143', 'D224053', 'D225113': 0, 1, 2, 3, 4
#> Items(s) 'D223143': 0, 1, 2, 3, 4, 5
#> 4 subject(s) do not solve any item:
#> P0007 (12 false), P0107 (12 false), P0837 (13 false) ...
#> 8 subject(s) solved each item: P0968 (11 correct), P0839 (13 correct), P1233 (24 correct) ...
#> Dataset is completely linked.
#> 'gauss' has been chosen for estimation method. Number of nodes was not explicitly specified. Set nodes to 20.
#> 101 common items found in 'anchor' list and data frame.
#> 5 of 106 item(s) of merging variable 'item' from data set 'item response data' not included in data set 'anchor list'.
#> Merging levels are not unique in data set 'anchor list'.
#> Q matrix specifies 1 dimension(s).
run3A<- runModel(def3A)
res3A<- getResults(run3A)
#> Getting standard errors with the tam.se function: 0.7 secs
#> Getting infit parameters calling tam.fit from getTamInfit: 0.2 secs
#> Getting WLEs calling tam.wle from getTamWles: 0.3 secs
#> |*****|
#> |-----|
#> Getting PVs calling tam.pv from getTamPVs: 0.3 secs
#> Getting Q3 statistic calling tam.modelfit from getTamQ3: 0.9 secs
it3A <- itemFromRes(res3A) ### all items except the ones with linking dif with equal item parameters? check
comp <- merge(it1[,c("item", "category", "est")], it3A[,c("item", "category", "est", "offset")], by=c("item", "category"), suffixes = c("_ref", "_foc"))
equal<- na.omit(comp[,c("est_ref", "offset")])
stopifnot(all(equal[,1] == equal[,2])) ### all item parameters without linking dif should be equal
lDif <- subset(comp, !is.na(est_foc)) ### all items with specific focus parameter must be included in linking DIF exclusion list
stopifnot(all(paste(lDif[,"item"], lDif[,"category"], sep="_") %in% eq$items[["not_specified"]][["Dim1"]][["info"]][,"itemExcluded"]))
# variant 2: use the item parameters for males (linking dif items excluded), transformed to the metric of females
def3B<- defineModel(dat=datW[which(datW[,"sex"] == "male"),], items = -c(1:4), id=1, irtmodel = "PCM",
anchor = eq[["items"]][["not_specified"]][["Dim1"]][["cleanedLinkItemPars"]],
itemCol = "item", valueCol = "est", catCol = "category", software="tam")
#> 7 variable(s) are not strictly dichotomous with 0/1 ... Expect a rating scale model or partial credit model.
#> Items(s) 'D025033', 'D201113', 'D204013', 'D205143', 'D224053', 'D225113': 0, 1, 2, 3, 4
#> Items(s) 'D223143': 0, 1, 2, 3, 4, 5
#> 4 subject(s) do not solve any item:
#> P0007 (12 false), P0107 (12 false), P0837 (13 false) ...
#> 8 subject(s) solved each item: P0968 (11 correct), P0839 (13 correct), P1233 (24 correct) ...
#> Dataset is completely linked.
#> 'gauss' has been chosen for estimation method. Number of nodes was not explicitly specified. Set nodes to 20.
#> 101 common items found in 'anchor' list and data frame.
#> 5 of 106 item(s) of merging variable 'item' from data set 'item response data' not included in data set 'anchor list'.
#> Merging levels are not unique in data set 'anchor list'.
#> Q matrix specifies 1 dimension(s).
run3B<- runModel(def3B)
res3B<- getResults(run3B)
#> Getting standard errors with the tam.se function: 0.7 secs
#> Getting WLEs calling tam.wle from getTamWles: 0.3 secs
#> |*****|
#> |-----|
#> Getting PVs calling tam.pv from getTamPVs: 0.3 secs
#> Getting Q3 statistic calling tam.modelfit from getTamQ3: 0.9 secs
it3B <- itemFromRes(res3B) ### all items except the ones with linking dif with equal item parameters? check
link <- eq[["items"]][["not_specified"]][["Dim1"]][["cleanedLinkItemPars"]][,c("item", "category", "est")]
comp <- merge(link, it3B[,c("item", "category", "est", "offset")], by=c("item", "category"), suffixes = c("_ref", "_foc"))
equal<- na.omit(comp[,c("est_ref", "offset")])
stopifnot(all(equal[,1] == equal[,2])) ### all item parameters without linking dif should be equal
lDif <- subset(comp, !is.na(est_foc)) ### all items with specific focus paraeter must be included in linking DIF exclusion list
stopifnot(all(paste(lDif[,"item"], lDif[,"category"], sep="_") %in% eq$items[["not_specified"]][["Dim1"]][["info"]][,"itemExcluded"]))
# transform to Bista metric
eq4 <- equat1pl(results = res3B) ### reference population mean and SD
#> Found 1 model(s).
#> Equating is executed for each dimension in each model separately.
#> No norm parameter defined ('prmNorm' is missing). Treat current sample as drawn from the reference population.
refP <- data.frame(domain = "Dim1", m = 0.0389, sd = 1.07108, stringsAsFactors = FALSE)
cuts <- list ( Dim1 = list(values = 390+0:3*75))
tf4 <- transformToBista(equatingList=eq4, refPop=refP, cuts=cuts, vera = FALSE)
#> The 'refPop' data.frame does not include information about reference population mean/SD on Bista metric. Values will be defaulted to mean = 500 and SD = 100.
#> Warning: Skip check whether all competence levels are occupied (due to bayesian plausible values imputation).
################################################################################
### Example 8.1: same like example 8 (anchored 1pl PCM), now using Conquest ###
################################################################################
# load partial credit long format data
data(reading)
# transform into wide format
datW <- reshape2::dcast(reading[which(reading[,"type"] != "iglu"),],idstud+sex+language+country~item, value.var = "valueSum")
# only some items are partial credit, which ones?
pc.it<- reading[which(reading[,"valueSum"] > 1),"item"] |> unique()
# give values of partial credit items
# few observation of 1-category for item D223143 ... may cause convergence trouble
lapply(datW[,pc.it], FUN = table)
#> $D204013
#>
#> 0 1 2 3 4
#> 24 42 101 263 324
#>
#> $D205143
#>
#> 0 1 2 3 4 5
#> 7 4 31 53 171 466
#>
#> $D201113
#>
#> 0 1 2 3 4
#> 15 113 141 170 104
#>
#> $D225113
#>
#> 0 1 2 3 4
#> 35 75 153 207 78
#>
#> $D224053
#>
#> 0 1 2 3 4
#> 15 25 94 172 225
#>
#> $D025033
#>
#> 0 1 2 3 4
#> 13 39 74 178 233
#>
#> $D223143
#>
#> 0 1 2 3 4 5
#> 23 6 41 100 106 254
#>
# combine the two lowest categories to avoid categories with too few observations
datW[,"D205143"] <- car::recode(datW[,"D205143"], "1=0; 2=1; 3=2; 4=3; 5=4")
# partial credit model ... we consider the female subgroup to be the reference population
# (norm population) that defines the scale and reference item parameters
dFema<- datW[which(datW[,"sex"] == "female"),] ### data for females
lapply(dFema[,pc.it], FUN = table) ### very few observations for some categories ... may cause convergence trouble
#> $D204013
#>
#> 0 1 2 3 4
#> 6 15 42 127 165
#>
#> $D205143
#>
#> 0 1 2 3 4
#> 3 17 20 97 253
#>
#> $D201113
#>
#> 0 1 2 3 4
#> 4 62 66 85 56
#>
#> $D225113
#>
#> 0 1 2 3 4
#> 14 43 90 107 43
#>
#> $D224053
#>
#> 0 1 2 3 4
#> 4 7 34 90 119
#>
#> $D025033
#>
#> 0 1 2 3 4
#> 3 19 39 100 120
#>
#> $D223143
#>
#> 0 1 2 3 4 5
#> 12 2 21 56 68 159
#>
def1 <- defineModel(dat=dFema, items = -c(1:4), id=1, model.statement = "item+item*step", nodes = 21, analysis.name = "pcm_conquest_females", dir=tempdir())
#> Cannot find conquest 2007 executable file. Please choose manually.
#> Error in file.choose(): file choice cancelled
run1 <- runModel(def1)
res1 <- getResults(run1)
#> Getting standard errors with the tam.se function: 0.7 secs
#> Getting infit parameters calling tam.fit from getTamInfit: 0.2 secs
#> Getting WLEs calling tam.wle from getTamWles: 0.6 secs
#> |*****|
#> |-----|
#> Getting PVs calling tam.pv from getTamPVs: 0.3 secs
#> Getting Q3 statistic calling tam.modelfit from getTamQ3: 0.9 secs
it1 <- itemFromRes(res1)
# males are focus group: initial free estimation of item parameters
def2 <- defineModel(dat=datW[which(datW[,"sex"] == "male"),], items = -c(1:4), id=1, model.statement = "item+item*step", nodes = 21, analysis.name = "pcm_conquest_males", dir=tempdir())
#> Cannot find conquest 2007 executable file. Please choose manually.
#> Error in file.choose(): file choice cancelled
run2 <- runModel(def2)
res2 <- getResults(run2)
#> Getting standard errors with the tam.se function: 0.7 secs
#> Getting infit parameters calling tam.fit from getTamInfit: 0.2 secs
#> Getting WLEs calling tam.wle from getTamWles: 0.3 secs
#> |*****|
#> |-----|
#> Getting PVs calling tam.pv from getTamPVs: 0.3 secs
#> Getting Q3 statistic calling tam.modelfit from getTamQ3: 0.9 secs
it2 <- itemFromRes(res2)
# link males to females ... males perform worse
# 11 items with linking dif identified
eq <- equat1pl(results = res2, prmNorm = it1, item = "item", cat="category", value = "est", difBound=.64, iterativ = TRUE)
#> Found 1 model(s).
#> Equating is executed for each dimension in each model separately.
#>
#> ====================================================================================================
#>
#> Model No. 1
#> Model name: not_specified
#> Number of dimension(s): 1
#> Name(s) of dimension(s): Dim1
#> Number of linking items: 128
#> Linking method: Mean.Mean
#>
#> Dimension 'Dim1': 10 of 128 items with linking |DIF| > 0.64 identified.
#> Iteration 1: Exclude item 'D205143_Cat1'.
#> Iteration 2: Exclude item 'D223143_Cat1'.
#> Iteration 3: Exclude item 'D204043_Cat1'.
#> Iteration 4: Exclude item 'D025033_Cat1'.
#> Iteration 5: Exclude item 'D201113_Cat1'.
#> Iteration 6: Exclude item 'D034053_Cat1'.
#> Iteration 7: Exclude item 'D224023_Cat1'.
#> Iteration 8: Exclude item 'D205143_Cat3'.
#> Iteration 9: Exclude item 'D221093_Cat1'.
#> Iteration 10: Exclude item 'D204033_Cat1'.
#> Iteration 11: Exclude item 'D224053_Cat3'.
#>
#> method iter itemExcluded DIF.excluded linking.constant linkerror
#> 1 iterativ 0 -0.221 0.034
#> 2 iterativ 1 D205143_Cat1 1.231 -0.211 0.033
#> 3 iterativ 2 D223143_Cat1 -1.055 -0.220 0.032
#> 4 iterativ 3 D204043_Cat1 1 -0.212 0.031
#> 5 iterativ 4 D025033_Cat1 0.996 -0.204 0.030
#> 6 iterativ 5 D201113_Cat1 0.898 -0.197 0.030
#> 7 iterativ 6 D034053_Cat1 0.849 -0.190 0.029
#> 8 iterativ 7 D224023_Cat1 0.807 -0.183 0.028
#> 9 iterativ 8 D205143_Cat3 0.795 -0.177 0.028
#> 10 iterativ 9 D221093_Cat1 -0.696 -0.183 0.028
#> 11 iterativ 10 D204033_Cat1 -0.668 -0.188 0.027
#> 12 iterativ 11 D224053_Cat3 0.658 -0.183 0.027
#>
# re-calibrate males with anchoring: use the DIF-cleaned set of anchor parameters for calibrating males on the reference scale
# variant 1: use the original item parameters for females and exclude linking dif items (it1_cleaned)
# please note that to date in Conquest only dichotomous items can be used for anchoring
weg <- eq[["items"]][["pcm_conquest_males"]][["Dim1"]][["info"]][-1,"itemExcluded"]
weg <- eatTools::whereAre(weg, paste(it1[,"item"], it1[,"category"], sep="_"))
#> Error in eatTools::whereAre(weg, paste(it1[, "item"], it1[, "category"], sep = "_")): Assertion on 'a' failed: Must be of type 'vector', not 'NULL'.
it1C <- it1[-weg,]
#> Error in -weg: invalid argument to unary operator
pcmIt<- unique(subset(it1C, category == "Cat2")[,"item"]) ### additionally exclude pcm items
it1C <- it1C[-eatTools::whereAre(pcmIt, it1C[,"item"]),]
#> Found 23 elements.
def3A<- defineModel(dat=datW[which(datW[,"sex"] == "male"),], items = -c(1:4), id=1,
model.statement = "item+item*step", nodes = 21, analysis.name = "pcm_conquest_males_anchored1", dir=tempdir(),
anchor = it1C[,c("item", "est")])
#> Cannot find conquest 2007 executable file. Please choose manually.
#> Error in file.choose(): file choice cancelled
run3A<- runModel(def3A)
res3A<- getResults(run3A)
#> Getting standard errors with the tam.se function: 0.7 secs
#> Getting WLEs calling tam.wle from getTamWles: 0.3 secs
#> |*****|
#> |-----|
#> Getting PVs calling tam.pv from getTamPVs: 0.3 secs
#> Getting Q3 statistic calling tam.modelfit from getTamQ3: 0.9 secs
it3A <- itemFromRes(res3A) ### all dichotomous items except the ones with linking dif with equal item parameters? check
comp <- merge(it1[,c("item", "category", "est")], it3A[,c("item", "category", "est", "offset")], by=c("item", "category"), suffixes = c("_ref", "_foc"))
equal<- na.omit(comp[,c("est_ref", "offset")])
stopifnot(all(equal[,1] == equal[,2])) ### all item parameters without linking dif should be equal
### TO DO: all items with specific focus parameter must be included in linking DIF exclusion list or must be pcm items
# variant 2: use the item parameters for males (linking dif items excluded), transformed to the metric of females
ank3B<- eq[["items"]][["pcm_conquest_males"]][["Dim1"]][["cleanedLinkItemPars"]]
pcmIt<- unique(subset(ank3B, category == "Cat2")[,"item"]) ### additionally exclude pcm items
#> Error: object 'category' not found
ank3B<- ank3B[-eatTools::whereAre(pcmIt, ank3B[,"item"]),]
#> Error in eatTools::whereAre(pcmIt, ank3B[, "item"]): Assertion on 'b' failed: Must be of type 'vector', not 'NULL'.
def3B<- defineModel(dat=datW[which(datW[,"sex"] == "male"),], items = -c(1:4), id=1,
model.statement = "item+item*step", analysis.name = "pcm_conquest_males_anchored2",
dir=tempdir(), anchor = ank3B[,c("item", "est")])
#> Cannot find conquest 2007 executable file. Please choose manually.
#> Error in file.choose(): file choice cancelled
run3B<- runModel(def3B)
res3B<- getResults(run3B)
#> Getting standard errors with the tam.se function: 0.7 secs
#> Getting WLEs calling tam.wle from getTamWles: 0.3 secs
#> |*****|
#> |-----|
#> Getting PVs calling tam.pv from getTamPVs: 0.3 secs
#> Getting Q3 statistic calling tam.modelfit from getTamQ3: 0.9 secs
it3B <- itemFromRes(res3B) ### all items except the ones with linking dif with equal item parameters? check
link <- eq[["items"]][["pcm_conquest_males"]][["Dim1"]][["cleanedLinkItemPars"]][,c("item", "category", "est")]
comp <- merge(link, it3B[,c("item", "category", "est", "offset")], by=c("item", "category"), suffixes = c("_ref", "_foc"))
#> Error in fix.by(by.x, x): 'by' must specify uniquely valid columns
equal<- na.omit(comp[,c("est_ref", "offset")])
stopifnot(all(equal[,1] == equal[,2])) ### all item parameters without linking dif should be equal
### TO DO: all items with specific focus parameter must be included in linking DIF exclusion list or must be pcm items
# transform to Bista metric
eq4 <- equat1pl(results = res3B) ### reference population mean and SD
#> Found 1 model(s).
#> Equating is executed for each dimension in each model separately.
#> No norm parameter defined ('prmNorm' is missing). Treat current sample as drawn from the reference population.
refP <- data.frame(domain = "Dim1", m = 0.0389, sd = 1.07108, stringsAsFactors = FALSE)
cuts <- list ( Dim1 = list(values = 390+0:3*75))
tf4 <- transformToBista(equatingList=eq4, refPop=refP, cuts=cuts, vera = FALSE)
#> The 'refPop' data.frame does not include information about reference population mean/SD on Bista metric. Values will be defaulted to mean = 500 and SD = 100.
#> Warning: Skip check whether all competence levels are occupied (due to bayesian plausible values imputation).
################################################################################
### Example 9: (restricted) generalized partial credit model with anchoring ###
################################################################################
# load partial credit long format data
data(reading)
# transform into wide format
datW <- reshape2::dcast(reading[which(reading[,"type"] != "iglu"),],idstud+sex+language+country~item, value.var = "valueSum")
# combine the two lowest categories to avoid categories with too few observations
datW[,"D205143"] <- car::recode(datW[,"D205143"], "1=0; 2=1; 3=2; 4=3; 5=4")
# restricted generalized partial credit model ... we consider the female subgroup to be the reference population
# we assume that task D223 and D224 are created for students with special educational need and therefore have
# different slope. Non-SPF items should have slope = 1
dFema<- datW[which(datW[,"sex"] == "female"),] ### data for females
items<- colnames(dFema)[-c(1:4)]
items<- data.frame(item = items, task = substr(items,1,4), slopeGrp = as.numeric(substr(items,1,4) %in% c("D223", "D224")), stringsAsFactors = FALSE) |> dplyr::mutate(slope = car::recode(slopeGrp, "0=1; else=NA"))
items[,"dim"] <- car::recode(substr(items[,"item"], 1,2),"'D0'='norm'; 'D2'='pilot'")
items<- data.frame ( items, model.matrix(~dim-1, data = items), stringsAsFactors = FALSE)
def1T<- defineModel(dat=dFema, items = items[,"item"], id=1, irtmodel = "GPCM.groups", software="tam", fixSlopeMat = na.omit(items[,c("item", "slope")]), est.slopegroups = items[,c("item", "slopeGrp")], nodes = 21, fac.oldxsi =0.6, increment.factor=1.05)
#> 7 variable(s) are not strictly dichotomous with 0/1 ... Expect a rating scale model or partial credit model.
#> Items(s) 'D025033', 'D201113', 'D204013', 'D205143', 'D224053', 'D225113': 0, 1, 2, 3, 4
#> Items(s) 'D223143': 0, 1, 2, 3, 4, 5
#> 2 subject(s) do not solve any item:
#> P1296 (24 false), P1299 (24 false), P1299 (24 false) ...
#> 12 subject(s) solved each item: P0081 (12 correct), P0869 (13 correct), P1047 (29 correct) ...
#> Dataset is completely linked.
#> Q matrix specifies 1 dimension(s).
#> Warning: To date, fixing slopes only works for dichotomous unidimensional or between-item multidimensional models.
#> Following 22 items in dataset without fixed slopes in 'fixSlopeMat'. Slope(s) will be estimated freely.
#> D223013, D223023, D223033, D223043, D223063, D223073, D223083, D223103, D223113, D223123, D223133, D223143, D224013, D224023, D224033, D224043, D224053, D224063, D224083, D224093, D224103, D224113
run1T<- runModel(def1T)
res1T<- getResults(run1T)
#> Getting standard errors with the tam.se function: 1 secs
#> Getting infit parameters calling tam.fit from getTamInfit: 0.2 secs
#> Getting WLEs calling tam.wle from getTamWles: 0.3 secs
#> |*****|
#> |-----|
#> Getting PVs calling tam.pv from getTamPVs: 0.3 secs
#> Getting Q3 statistic calling tam.modelfit from getTamQ3: 0.9 secs
it1T <- itemFromRes(res1T)
# males are focus group: initial free estimation of item parameters
def2T<- defineModel(dat=datW[which(datW[,"sex"] == "male"),], items = -c(1:4), id=1, irtmodel = "GPCM.groups",software="tam", fixSlopeMat = na.omit(items[,c("item", "slope")]), est.slopegroups = items[,c("item", "slopeGrp")], nodes = 21)
#> 7 variable(s) are not strictly dichotomous with 0/1 ... Expect a rating scale model or partial credit model.
#> Items(s) 'D025033', 'D201113', 'D204013', 'D205143', 'D224053', 'D225113': 0, 1, 2, 3, 4
#> Items(s) 'D223143': 0, 1, 2, 3, 4, 5
#> 4 subject(s) do not solve any item:
#> P0007 (12 false), P0107 (12 false), P0837 (13 false) ...
#> 8 subject(s) solved each item: P0968 (11 correct), P0839 (13 correct), P1233 (24 correct) ...
#> Dataset is completely linked.
#> Q matrix specifies 1 dimension(s).
#> Warning: To date, fixing slopes only works for dichotomous unidimensional or between-item multidimensional models.
#> Following 22 items in dataset without fixed slopes in 'fixSlopeMat'. Slope(s) will be estimated freely.
#> D223013, D223023, D223033, D223043, D223063, D223073, D223083, D223103, D223113, D223123, D223133, D223143, D224013, D224023, D224033, D224043, D224053, D224063, D224083, D224093, D224103, D224113
run2T<- runModel(def2T)
res2T<- getResults(run2T)
#> Getting standard errors with the tam.se function: 0.7 secs
#> Getting infit parameters calling tam.fit from getTamInfit: 0.2 secs
#> Getting WLEs calling tam.wle from getTamWles: 0.3 secs
#> |*****|
#> |-----|
#> Getting PVs calling tam.pv from getTamPVs: 0.3 secs
#> Getting Q3 statistic calling tam.modelfit from getTamQ3: 0.9 secs
it2T <- itemFromRes(res2T)
unique(round(it1T[,"estSlope"],3)); unique(round(it2T[,"estSlope"],3)) ### average discrimination (reg vs. spf) differs for males, but not for females
#> [1] 1.000 1.001
#> [1] 1.000 0.858
# use only slope=1 items for 1pl linking
eq <- equat1pl(results = res2T, prmNorm = it1T[which(it1T[,"estSlope"] ==1),], item = "item", cat="category", value = "est", difBound=.64, iterativ = TRUE)
#> Found 1 model(s).
#> Equating is executed for each dimension in each model separately.
#>
#> ====================================================================================================
#>
#> Model No. 1
#> Model name: not_specified
#> Number of dimension(s): 1
#> Name(s) of dimension(s): Dim1
#> Number of linking items: 99
#> Linking method: Mean.Mean
#>
#> Dimension 'Dim1': 8 of 99 items with linking |DIF| > 0.64 identified.
#> Iteration 1: Exclude item 'D205143_Cat1'.
#> Iteration 2: Exclude item 'D025033_Cat1'.
#> Iteration 3: Exclude item 'D204043_Cat1'.
#> Iteration 4: Exclude item 'D201113_Cat1'.
#> Iteration 5: Exclude item 'D034053_Cat1'.
#> Iteration 6: Exclude item 'D221093_Cat1'.
#> Iteration 7: Exclude item 'D205143_Cat3'.
#> Iteration 8: Exclude item 'D204033_Cat1'.
#>
#> method iter itemExcluded DIF.excluded linking.constant linkerror
#> 1 iterativ 0 -0.239 0.039
#> 2 iterativ 1 D205143_Cat1 1.3 -0.226 0.037
#> 3 iterativ 2 D025033_Cat1 1.05 -0.215 0.036
#> 4 iterativ 3 D204043_Cat1 1.01 -0.205 0.034
#> 5 iterativ 4 D201113_Cat1 0.874 -0.196 0.033
#> 6 iterativ 5 D034053_Cat1 0.851 -0.187 0.033
#> 7 iterativ 6 D221093_Cat1 -0.687 -0.194 0.032
#> 8 iterativ 7 D205143_Cat3 0.676 -0.187 0.032
#> 9 iterativ 8 D204033_Cat1 -0.666 -0.194 0.031
#>
# variant 1
# use the DIF-cleaned set of original anchor parameters for calibrating males on the reference scale
weg <- eq[["items"]][["not_specified"]][["Dim1"]][["info"]][-1,"itemExcluded"]
weg <- eatTools::whereAre(weg, paste(it1T[,"item"], it1T[,"category"], sep="_"))
#> Found 8 elements.
it1C <- subset(it1T[-weg,], estSlope == 1)
def3T<- defineModel(eatTools::na_omit_selection(dat=datW[which(datW[,"sex"] == "male"),],"language"),
items = -c(1:4), id=1, irtmodel = "GPCM.groups", anchor = it1C, HG.var = c("language", "country"),
itemCol = "item", valueCol = "est", catCol = "category", fixSlopeMat = na.omit(items[,c("item", "slope")]),
qMatrix = items[,c("item", "dimnorm", "dimpilot")], est.slopegroups = items[,c("item", "slopeGrp")], nodes = 21, software="tam")
#> 7 variable(s) are not strictly dichotomous with 0/1 ... Expect a rating scale model or partial credit model.
#> Items(s) 'D025033', 'D201113', 'D204013', 'D205143', 'D224053', 'D225113': 0, 1, 2, 3, 4
#> Items(s) 'D223143': 0, 1, 2, 3, 4, 5
#> Background variable(s) 'language', 'country' of class
#> 'factor', 'factor' will be converted to indicator variables.
#> Variable 'language' was converted to 2 indicator(s) with name(s) 'languagenativeAndOther', 'languageother'.
#> Variable 'country' was converted to 8 indicator(s) with name(s) 'countryCaladan', 'countryDisneyland', 'countryGreenland', 'countryMordor', 'countryRomulus', 'countryTatooine', 'countrytheShire', 'countryVulcan'.
#> 4 subject(s) do not solve any item:
#> P0007 (12 false), P0107 (12 false), P0837 (13 false) ...
#> 8 subject(s) solved each item: P0968 (11 correct), P0839 (13 correct), P1233 (24 correct) ...
#> Dataset is completely linked.
#> 80 common items found in 'anchor' list and data frame.
#> 26 of 106 item(s) of merging variable 'item' from data set 'item response data' not included in data set 'anchor list'.
#> Merging levels are not unique in data set 'anchor list'.
#> Q matrix specifies 2 dimension(s).
#> Warning: To date, fixing slopes only works for dichotomous unidimensional or between-item multidimensional models.
#> Following 22 items in dataset without fixed slopes in 'fixSlopeMat'. Slope(s) will be estimated freely.
#> D223013, D223023, D223033, D223043, D223063, D223073, D223083, D223103, D223113, D223123, D223133, D223143, D224013, D224023, D224033, D224043, D224053, D224063, D224083, D224093, D224103, D224113
run3T<- runModel(def3T)
#> Error: 'timeFormat' is not an exported object from 'namespace:eatTools'
res3T<- getResults(run3T)
#> Error: object 'run3T' not found
it3T <- itemFromRes(res3T) ### all items except the ones with linking dif with equal item parameters? check
#> Error: object 'res3T' not found
comp <- merge(subset(it1T[,c("item", "category", "est", "estSlope")],estSlope == 1), it3T[,c("item", "category", "est", "offset")], by=c("item", "category"), all=TRUE, suffixes = c("_ref", "_foc"))
#> Error: object 'it3T' not found
equal<- na.omit(comp[,c("est_ref", "offset")])
stopifnot(all(equal[,1] == equal[,2])) ### all item parameters without linking dif should be equal
lDif <- subset(comp, !is.na(est_foc))
stopifnot(all(paste(subset(lDif, !is.na(est_ref))[,"item"], subset(lDif, !is.na(est_ref))[,"category"], sep="_") %in% eq$items[["not_specified"]][["Dim1"]][["info"]][,"itemExcluded"]))
#> Error: all(paste(subset(lDif, !is.na(est_ref))[, "item"], subset(lDif, .... is not TRUE
# variant 2: use the 1pl item parameters for males (linking dif items excluded), transformed to the metric of females
def4T<- defineModel(eatTools::na_omit_selection(dat=datW[which(datW[,"sex"] == "male"),],"language"),
items = -c(1:4), id=1, irtmodel = "GPCM.groups", anchor = eq[["items"]][["not_specified"]][["Dim1"]][["cleanedLinkItemPars"]],
HG.var = c("language", "country"), itemCol = "item", valueCol = "est", catCol = "category", fixSlopeMat = na.omit(items[,c("item", "slope")]),
qMatrix = items[,c("item", "dimnorm", "dimpilot")], est.slopegroups = items[,c("item", "slopeGrp")], nodes = 21, software="tam")
#> 7 variable(s) are not strictly dichotomous with 0/1 ... Expect a rating scale model or partial credit model.
#> Items(s) 'D025033', 'D201113', 'D204013', 'D205143', 'D224053', 'D225113': 0, 1, 2, 3, 4
#> Items(s) 'D223143': 0, 1, 2, 3, 4, 5
#> Background variable(s) 'language', 'country' of class
#> 'factor', 'factor' will be converted to indicator variables.
#> Variable 'language' was converted to 2 indicator(s) with name(s) 'languagenativeAndOther', 'languageother'.
#> Variable 'country' was converted to 8 indicator(s) with name(s) 'countryCaladan', 'countryDisneyland', 'countryGreenland', 'countryMordor', 'countryRomulus', 'countryTatooine', 'countrytheShire', 'countryVulcan'.
#> 4 subject(s) do not solve any item:
#> P0007 (12 false), P0107 (12 false), P0837 (13 false) ...
#> 8 subject(s) solved each item: P0968 (11 correct), P0839 (13 correct), P1233 (24 correct) ...
#> Dataset is completely linked.
#> 80 common items found in 'anchor' list and data frame.
#> 26 of 106 item(s) of merging variable 'item' from data set 'item response data' not included in data set 'anchor list'.
#> Merging levels are not unique in data set 'anchor list'.
#> Q matrix specifies 2 dimension(s).
#> Warning: To date, fixing slopes only works for dichotomous unidimensional or between-item multidimensional models.
#> Following 22 items in dataset without fixed slopes in 'fixSlopeMat'. Slope(s) will be estimated freely.
#> D223013, D223023, D223033, D223043, D223063, D223073, D223083, D223103, D223113, D223123, D223133, D223143, D224013, D224023, D224033, D224043, D224053, D224063, D224083, D224093, D224103, D224113
run4T<- runModel(def4T)
#> Error: 'timeFormat' is not an exported object from 'namespace:eatTools'
res4T<- getResults(run4T)
#> Error: object 'run4T' not found
it4T <- itemFromRes(res4T) ### all items except the ones with linking dif with equal item parameters? check
#> Error: object 'res4T' not found
link <- eq[["items"]][["not_specified"]][["Dim1"]][["cleanedLinkItemPars"]][,c("item", "category", "est")]
comp <- merge(link, it4T[,c("item", "category", "est", "offset")], by=c("item", "category"), suffixes = c("_ref", "_foc"), all=TRUE)
#> Error: object 'it4T' not found
equal<- na.omit(comp[,c("est_ref", "offset")])
stopifnot(all(equal[,1] == equal[,2])) ### all item parameters without linking dif should be equal
lDif <- subset(comp, !is.na(est_foc)) ### all items with specific focus paraeter must be included in linking DIF exclusion list
stopifnot(all(paste(subset(lDif, !is.na(est_ref))[,"item"], subset(lDif, !is.na(est_ref))[,"category"], sep="_") %in% eq$items[["not_specified"]][["Dim1"]][["info"]][,"itemExcluded"]))
#> Error: all(paste(subset(lDif, !is.na(est_ref))[, "item"], subset(lDif, .... is not TRUE
# transform to Bista metric
eq4 <- equat1pl(results = res4T) ### reference population mean and SD
#> Error: object 'res4T' not found
refP <- data.frame(domain = c("dimnorm", "dimpilot"), m = 0.0389, sd = 1.07108, stringsAsFactors = FALSE)
cuts <- list ( dimnorm = list(values = 390+0:3*75), dimpilot = list(values = 390+0:3*75))
tf4 <- transformToBista(equatingList=eq4, refPop=refP, cuts=cuts, vera = FALSE)
#> Error in generateOrCheckRefPop(equatingList = equatingList, refPop = refPop, mods = mods, dims = dims, isRunM = isRunM, id = id, weights = weights, defaultM = defaultM, defaultSD = defaultSD): Following 1 dimension(s) not included in 'refPop': 'Dim1'.
################################################################################
### Example 10: GPCM, similar like example 9, but now using mirt ###
################################################################################
# load package Hmisc if necesary
if(!"Hmisc" %in% .packages()) {library(Hmisc)}
#>
#> Attaching package: ‘Hmisc’
#> The following object is masked from ‘package:survey’:
#>
#> deff
#> The following objects are masked from ‘package:base’:
#>
#> format.pval, units
# load partial credit long format data
data(reading)
# transform into wide format
datW <- reshape2::dcast(reading[which(reading[,"type"] != "iglu"),],idstud+sex+language+country~item, value.var = "valueSum")
# only some items are partial credit, which ones?
pc.it<- reading[which(reading[,"valueSum"] > 1),"item"] |> unique()
# combine the two lowest categories to avoid categories with too few observations
datW[,"D205143"] <- car::recode(datW[,"D205143"], "1=0; 2=1; 3=2; 4=3; 5=4")
# restricted generalized partial credit model ... we consider the female subgroup to be the reference population
# we assume that task D223 and D224 are created for students with special educational need and therefore have
# different slope (gpcm). Non-SPF items should have slope = 1
dFema<- datW[which(datW[,"sex"] == "female"),] ### data for females
items<- colnames(dFema)[-c(1:4)]
items<- data.frame(item = items, task = substr(items,1,4), slopeGrp = as.numeric(substr(items,1,4) %in% c("D223", "D224")), stringsAsFactors = FALSE) |> dplyr::mutate(irtmod = NA)
# add model information to 'item'
items[intersect(intersect(which(items[,"slopeGrp"] == 0), which(items[,"task"] %nin% c("D223", "D224"))), which(items[,"item"] %nin% pc.it)),"irtmod"] <- "Rasch"
items[intersect(intersect(which(items[,"slopeGrp"] == 1), which(items[,"task"] %in% c("D223", "D224"))), which(items[,"item"] %nin% pc.it)),"irtmod"] <- "2PL"
# this is partial credit which is specified in 'mirt' with 'Rasch' statement as the slope equals 1 like in dichotomous Rasch models
items[intersect(intersect(which(items[,"slopeGrp"] == 0), which(items[,"task"] %nin% c("D223", "D224"))), which(items[,"item"] %in% pc.it)),"irtmod"] <- "Rasch"
# generalized partial credit for unidimensional models
items[intersect(which(items[,"slopeGrp"] == 1), which(items[,"item"] %in% pc.it)),"irtmod"] <- "gpcm"
# define arbitrary dimensions
items[,"dim"] <- car::recode(substr(items[,"item"], 1,2),"'D0'='norm'; 'D2'='pilot'")
items<- data.frame ( items, model.matrix(~dim-1, data = items), stringsAsFactors = FALSE)
# procedure similar to conquest/tam
def1 <- defineModel(dat=eatTools::na_omit_selection(dFema,varsToOmitIfNA="language"), items = items[,"item"],
id=1,irtmodel = items[,c("item", "irtmod")], software="mirt")
#> 7 variable(s) are not strictly dichotomous with 0/1 ... Expect a rating scale model or partial credit model.
#> Items(s) 'D025033', 'D201113', 'D204013', 'D205143', 'D224053', 'D225113': 0, 1, 2, 3, 4
#> Items(s) 'D223143': 0, 1, 2, 3, 4, 5
#> 12 subject(s) solved each item: P0081 (12 correct), P0869 (13 correct), P1047 (29 correct) ...
#> Dataset is completely linked.
#> Specifying 'method' and 'nodes' not yet implemented for 'mirt'.
#> Q matrix specifies 1 dimension(s).
skel1<- runModel(def1, onlySkeleton = TRUE) ### check the model "skeleton"
run1 <- runModel(def1) ### run the model finally
#> Modify skeleton ...
#>
#>
#> Calculating information matrix...
res1 <- getResults(run1)
#> Getting WLEs calling fscores(method="WLE") from getMirtWles: 11.4 secs
#> Getting PVs calling fscores from getMirtPVs: 0.2 secs
it1 <- itemFromRes(res1)
# males are focus group: initial free estimation of item parameters
def2 <- defineModel(dat=eatTools::na_omit_selection(datW[which(datW[,"sex"] == "male"),],varsToOmitIfNA="language"),
items = -c(1:4), id=1, irtmodel = items[,c("item", "irtmod")],software="mirt")
#> 7 variable(s) are not strictly dichotomous with 0/1 ... Expect a rating scale model or partial credit model.
#> Items(s) 'D025033', 'D201113', 'D204013', 'D205143', 'D224053', 'D225113': 0, 1, 2, 3, 4
#> Items(s) 'D223143': 0, 1, 2, 3, 4, 5
#> 4 subject(s) do not solve any item:
#> P0007 (12 false), P0107 (12 false), P0837 (13 false) ...
#> 8 subject(s) solved each item: P0968 (11 correct), P0839 (13 correct), P1233 (24 correct) ...
#> Dataset is completely linked.
#> Specifying 'method' and 'nodes' not yet implemented for 'mirt'.
#> Q matrix specifies 1 dimension(s).
run2 <- runModel(def2)
#> Modify skeleton ...
#>
#>
#> Calculating information matrix...
res2 <- getResults(run2)
#> Getting WLEs calling fscores(method="WLE") from getMirtWles: 10.5 secs
it2 <- itemFromRes(res2)
# link males to females, using only 1pl items ... males perform worse. 10 items with linking dif identified
eq <- equat1pl(results = res2, prmNorm = subset(it1,estSlope ==1), item = "item", cat="category", value = "est", difBound=.64, iterativ = TRUE)
#> Found 1 model(s).
#> Equating is executed for each dimension in each model separately.
#>
#> ====================================================================================================
#>
#> Model No. 1
#> Model name: not_specified
#> Number of dimension(s): 1
#> Name(s) of dimension(s): Dim1
#> Number of linking items: 84
#> Linking method: Mean.Mean
#>
#> Dimension 'Dim1': 6 of 84 items with linking |DIF| > 0.64 identified.
#> Iteration 1: Exclude item 'D205143_Cat1'.
#> Iteration 2: Exclude item 'D034053_Cat1'.
#> Iteration 3: Exclude item 'D204043_Cat1'.
#> Iteration 4: Exclude item 'D025033_Cat1'.
#> Iteration 5: Exclude item 'D204033_Cat1'.
#> Iteration 6: Exclude item 'D221093_Cat1'.
#>
#> method iter itemExcluded DIF.excluded linking.constant linkerror
#> 1 iterativ 0 -0.263 0.044
#> 2 iterativ 1 D205143_Cat1 1.841 -0.241 0.039
#> 3 iterativ 2 D034053_Cat1 0.907 -0.230 0.037
#> 4 iterativ 3 D204043_Cat1 0.916 -0.219 0.036
#> 5 iterativ 4 D025033_Cat1 0.776 -0.209 0.035
#> 6 iterativ 5 D204033_Cat1 -0.68 -0.218 0.035
#> 7 iterativ 6 D221093_Cat1 -0.687 -0.227 0.034
#>
# re-calibrate males with anchoring: use the DIF-cleaned set of anchor parameters for calibrating males on the reference scale
# variant 1: use the original (1pl) item parameters for females and exclude linking dif items (it1_cleaned)
weg <- eq[["items"]][["not_specified"]][["Dim1"]][["info"]][-1,"itemExcluded"]
weg <- eatTools::whereAre(weg, paste(it1[,"item"], it1[,"category"], sep="_"))
#> Found 6 elements.
it1C <- subset(it1[-weg,], estSlope == 1)
def3A<- defineModel(dat=eatTools::na_omit_selection(datW[which(datW[,"sex"] == "male"),],varsToOmitIfNA="language"), items = -c(1:4), id=1, irtmodel = items[,c("item", "irtmod")],
anchor = it1C, itemCol = "item", valueCol = "est", catCol = "category", software="mirt")
#> 7 variable(s) are not strictly dichotomous with 0/1 ... Expect a rating scale model or partial credit model.
#> Items(s) 'D025033', 'D201113', 'D204013', 'D205143', 'D224053', 'D225113': 0, 1, 2, 3, 4
#> Items(s) 'D223143': 0, 1, 2, 3, 4, 5
#> 4 subject(s) do not solve any item:
#> P0007 (12 false), P0107 (12 false), P0837 (13 false) ...
#> 8 subject(s) solved each item: P0968 (11 correct), P0839 (13 correct), P1233 (24 correct) ...
#> Dataset is completely linked.
#> Specifying 'method' and 'nodes' not yet implemented for 'mirt'.
#> 78 common items found in 'anchor' list and data frame.
#> 28 of 106 item(s) of merging variable 'item' from data set 'item response data' not included in data set 'anchor list'.
#> Q matrix specifies 1 dimension(s).
run3A<- runModel(def3A)
#> Modify skeleton ...
#>
#>
#> Calculating information matrix...
res3A<- getResults(run3A)
#> Getting WLEs calling fscores(method="WLE") from getMirtWles: 10.6 secs
it3A <- itemFromRes(res3A) ### all items except the ones with linking dif with equal item parameters? check
comp <- merge(subset(it1, estSlope==1)[,c("item", "category", "est")], subset(it3A, !is.na(offset))[,c("item", "category", "est", "offset")], by=c("item", "category"), suffixes = c("_ref", "_foc"))
stopifnot(all(round(comp[,"est_ref"],5) == round(comp[,"offset"],5))) ### all item parameters without linking dif should be equal
# all items with specific focus parameter must be included in linking DIF exclusion list
stopifnot(all((paste(comp[,"item"], comp[,"category"],sep="_") %in% eq$items[["not_specified"]][["Dim1"]][["info"]][-1,"itemExcluded"]) == FALSE))
# variant 2: use the item parameters for males (linking dif items excluded), transformed to the metric of females
def3B<- defineModel(dat=eatTools::na_omit_selection(datW[which(datW[,"sex"] == "male"),],varsToOmitIfNA="language"), items = -c(1:4), id=1, irtmodel = items[,c("item", "irtmod")],
anchor = eq[["items"]][["not_specified"]][["Dim1"]][["cleanedLinkItemPars"]],
itemCol = "item", valueCol = "est", catCol = "category", software="mirt")
#> 7 variable(s) are not strictly dichotomous with 0/1 ... Expect a rating scale model or partial credit model.
#> Items(s) 'D025033', 'D201113', 'D204013', 'D205143', 'D224053', 'D225113': 0, 1, 2, 3, 4
#> Items(s) 'D223143': 0, 1, 2, 3, 4, 5
#> 4 subject(s) do not solve any item:
#> P0007 (12 false), P0107 (12 false), P0837 (13 false) ...
#> 8 subject(s) solved each item: P0968 (11 correct), P0839 (13 correct), P1233 (24 correct) ...
#> Dataset is completely linked.
#> Specifying 'method' and 'nodes' not yet implemented for 'mirt'.
#> 78 common items found in 'anchor' list and data frame.
#> 28 of 106 item(s) of merging variable 'item' from data set 'item response data' not included in data set 'anchor list'.
#> Q matrix specifies 1 dimension(s).
run3B<- runModel(def3B)
#> Modify skeleton ...
#>
#>
#> Calculating information matrix...
res3B<- getResults(run3B)
#> Getting WLEs calling fscores(method="WLE") from getMirtWles: 10.8 secs
it3B <- itemFromRes(res3B) ### all items except the ones with linking dif with equal item parameters? check
link <- eq[["items"]][["not_specified"]][["Dim1"]][["cleanedLinkItemPars"]][,c("item", "category", "est", "estSlope")]
stopifnot(all(link[,"estSlope"] == 1))
comp <- merge(link, it3B[,c("item", "category", "est", "offset")], by=c("item", "category"), suffixes = c("_ref", "_foc"))
stopifnot(all(round(comp[,"est_ref"],5) == round(comp[,"offset"],5))) ### all item parameters without linking dif should be equal
# all items with specific focus parameter must be included in linking DIF exclusion list
stopifnot(all((paste(comp[,"item"], comp[,"category"],sep="_") %in% eq$items[["not_specified"]][["Dim1"]][["info"]][-1,"itemExcluded"]) == FALSE))
# transform to Bista metric
eq4 <- equat1pl(results = res3B) ### reference population mean and SD
#> Found 1 model(s).
#> Equating is executed for each dimension in each model separately.
#> No norm parameter defined ('prmNorm' is missing). Treat current sample as drawn from the reference population.
refP <- data.frame(domain = "Dim1", m = 0.0389, sd = 1.07108, stringsAsFactors = FALSE)
cuts <- list ( Dim1 = list(values = 390+0:3*75))
tf4 <- transformToBista(equatingList=eq4, refPop=refP, cuts=cuts, vera = FALSE)
#> Warning: Cannot identify student identifier variable (possibly because 'resultsObj' was created by an older version of 'eatModel'). student id variable will be defaulted to 'idstud'.
#> The 'refPop' data.frame does not include information about reference population mean/SD on Bista metric. Values will be defaulted to mean = 500 and SD = 100.
#> Warning: Skip check whether all competence levels are occupied (due to bayesian plausible values imputation).
################################################################################
### Example 11: differential item functioning in partial credit models ###
################################################################################
# load partial credit long format data
data(reading)
# create a small wide format exemplary data set, only booklet 08
dw <- reshape2::dcast(subset(reading, bookletID == "TH08"), idstud+sex~item, value.var = "valueSum") |>
dplyr::mutate(sex = car::recode(sex, "'male'=0; 'female'=1", as.factor=FALSE)) |>
eatTools::na_omit_selection("sex")
# define the model: the specification resembles exemple 9 in tam.mml help page
defT<- defineModel(dat = dw, items = -c(1:2), DIF.var="sex", id = "idstud", irtmodel = "PCM",software="tam")
#> 4 variable(s) are not strictly dichotomous with 0/1 ... Expect a rating scale model or partial credit model.
#> Items(s) 'D025033', 'D204013', 'D225113': 0, 1, 2, 3, 4
#> Items(s) 'D205143': 0, 1, 2, 3, 4, 5
#> Dataset is completely linked.
#> 'gauss' has been chosen for estimation method. Number of nodes was not explicitly specified. Set nodes to 20.
#> Q matrix specifies 1 dimension(s).
runT<- runModel(defT)
resT<- getResults(runT)
#> Warning: NaNs produced
#> Warning: NaNs produced
#> Warning: NaNs produced
#> Warning: NaNs produced
#> Warning: NaNs produced
#> Warning: NaNs produced
#> Warning: NaNs produced
#> Warning: NaNs produced
#> Warning: NaNs produced
#> Warning: NaNs produced
#> Warning: NaNs produced
#> Warning: NaNs produced
#> Warning: NaNs produced
#> Warning: NaNs produced
#> Warning: NaNs produced
#> Warning: NaNs produced
#> Warning: NaNs produced
#> Warning: NaNs produced
#> Warning: NaNs produced
#> Warning: NaNs produced
#> Warning: NaNs produced
#> Warning: NaNs produced
#> Warning: NaNs produced
#> Warning: NaNs produced
#> Warning: NaNs produced
#> Warning: NaNs produced
#> Warning: NaNs produced
#> Warning: NaNs produced
#> Warning: NaNs produced
#> Warning: NaNs produced
#> Warning: NaNs produced
#> Warning: NaNs produced
#> Warning: NaNs produced
#> Warning: NaNs produced
#> Warning: NaNs produced
#> Warning: NaNs produced
#> Warning: NaNs produced
#> Warning: NaNs produced
#> Warning: NaNs produced
#> Warning: NaNs produced
#> Warning: NaNs produced
#> Warning: NaNs produced
#> Warning: NaNs produced
#> Warning: NaNs produced
#> Warning: NaNs produced
#> Warning: NaNs produced
#> Warning: NaNs produced
#> Warning: NaNs produced
#> Warning: NaNs produced
#> Warning: NaNs produced
#> Warning: NaNs produced
#> Warning: NaNs produced
#> Warning: NaNs produced
#> Warning: NaNs produced
#> Warning: NaNs produced
#> Warning: NaNs produced
#> Warning: NaNs produced
#> Warning: NaNs produced
#> Warning: NaNs produced
#> Warning: NaNs produced
#> Warning: NaNs produced
#> Warning: NaNs produced
#> Warning: NaNs produced
#> Warning: NaNs produced
#> Warning: NaNs produced
#> Warning: NaNs produced
#> Warning: NaNs produced
#> Warning: NaNs produced
#> Warning: NaNs produced
#> Warning: NaNs produced
#> Warning: NaNs produced
#> Warning: NaNs produced
#> Warning: NaNs produced
#> Warning: NaNs produced
#> Warning: NaNs produced
#> Warning: NaNs produced
#> Warning: NaNs produced
#> Warning: NaNs produced
#> Warning: NaNs produced
#> Warning: NaNs produced
#> Warning: NaNs produced
#> Warning: NaNs produced
#> Warning: NaNs produced
#> Warning: NaNs produced
#> Warning: NaNs produced
#> Warning: NaNs produced
#> Warning: NaNs produced
#> Warning: NaNs produced
#> Warning: NaNs produced
#> Warning: NaNs produced
#> Warning: NaNs produced
#> Warning: NaNs produced
#> Warning: NaNs produced
#> Warning: NaNs produced
#> Warning: NaNs produced
#> Warning: NaNs produced
#> Warning: NaNs produced
#> Warning: NaNs produced
#> Warning: NaNs produced
#> Warning: NaNs produced
#> Warning: NaNs produced
#> Warning: NaNs produced
#> Warning: NaNs produced
#> Warning: NaNs produced
#> Warning: NaNs produced
#> Warning: NaNs produced
#> Warning: NaNs produced
#> Warning: NaNs produced
#> Warning: NaNs produced
#> Warning: NaNs produced
#> Warning: NaNs produced
#> Warning: NaNs produced
#> Warning: NaNs produced
#> Warning: NaNs produced
#> Warning: NaNs produced
#> Warning: NaNs produced
#> Warning: NaNs produced
#> Warning: NaNs produced
#> Warning: NaNs produced
#> Warning: NaNs produced
#> Warning: NaNs produced
#> Warning: NaNs produced
#> Warning: NaNs produced
#> Warning: NaNs produced
#> Warning: NaNs produced
#> Warning: NaNs produced
#> Warning: NaNs produced
#> Warning: NaNs produced
#> Warning: NaNs produced
#> Warning: NaNs produced
#> Warning: NaNs produced
#> Warning: NaNs produced
#> Warning: NaNs produced
#> Warning: NaNs produced
#> Warning: NaNs produced
#> Warning: NaNs produced
#> Warning: NaNs produced
#> Warning: NaNs produced
#> Warning: NaNs produced
#> Warning: NaNs produced
#> Warning: NaNs produced
#> Warning: NaNs produced
#> Warning: NaNs produced
#> Warning: NaNs produced
#> Warning: NaNs produced
#> Warning: NaNs produced
#> Warning: NaNs produced
#> Warning: NaNs produced
#> Warning: NaNs produced
#> Warning: NaNs produced
#> Warning: NaNs produced
#> Warning: NaNs produced
#> Warning: NaNs produced
#> Warning: NaNs produced
#> Warning: NaNs produced
#> Warning: NaNs produced
#> Warning: NaNs produced
#> Warning: NaNs produced
#> Warning: NaNs produced
#> Warning: NaNs produced
#> Warning: NaNs produced
#> Warning: NaNs produced
#> Warning: NaNs produced
#> Warning: NaNs produced
#> Warning: NaNs produced
#> Warning: NaNs produced
#> Warning: NaNs produced
#> Warning: NaNs produced
#> Warning: NaNs produced
#> Warning: NaNs produced
#> Warning: NaNs produced
#> Warning: NaNs produced
#> Warning: NaNs produced
#> Warning: NaNs produced
#> Warning: NaNs produced
#> Warning: NaNs produced
#> Warning: NaNs produced
#> Warning: NaNs produced
#> Warning: NaNs produced
#> Warning: NaNs produced
#> Warning: NaNs produced
#> Warning: NaNs produced
#> Warning: NaNs produced
#> Warning: NaNs produced
#> Warning: NaNs produced
#> Warning: NaNs produced
#> Warning: NaNs produced
#> Warning: NaNs produced
#> Warning: NaNs produced
#> Warning: NaNs produced
#> Warning: NaNs produced
#> Warning: NaNs produced
#> Warning: NaNs produced
#> Warning: NaNs produced
#> Warning: NaNs produced
#> Warning: NaNs produced
#> Warning: NaNs produced
#> Warning: NaNs produced
#> Warning: NaNs produced
#> Warning: NaNs produced
#> Warning: NaNs produced
#> Warning: NaNs produced
#> Warning: NaNs produced
#> Warning: NaNs produced
#> Warning: NaNs produced
#> Warning: NaNs produced
#> Warning: NaNs produced
#> Warning: NaNs produced
#> Warning: NaNs produced
#> Warning: NaNs produced
#> Warning: NaNs produced
#> Warning: NaNs produced
#> Warning: NaNs produced
#> Warning: NaNs produced
#> Warning: NaNs produced
#> Warning: NaNs produced
#> Warning: NaNs produced
#> Warning: NaNs produced
#> Warning: NaNs produced
#> Warning: NaNs produced
#> Warning: NaNs produced
#> Warning: NaNs produced
#> Warning: NaNs produced
#> Warning: NaNs produced
#> Warning: NaNs produced
#> Warning: NaNs produced
#> Warning: NaNs produced
#> Warning: NaNs produced
#> Warning: NaNs produced
#> Warning: NaNs produced
#> Warning: NaNs produced
#> Warning: NaNs produced
#> Warning: NaNs produced
#> Warning: NaNs produced
#> Warning: NaNs produced
#> Warning: NaNs produced
#> Warning: NaNs produced
#> Warning: NaNs produced
#> Warning: NaNs produced
#> Warning: NaNs produced
#> Warning: NaNs produced
#> Warning: NaNs produced
#> Warning: NaNs produced
#> Warning: NaNs produced
#> Warning: NaNs produced
#> Warning: NaNs produced
#> Warning: NaNs produced
#> Warning: NaNs produced
#> Warning: NaNs produced
#> Warning: NaNs produced
#> Warning: NaNs produced
#> Warning: NaNs produced
#> Warning: NaNs produced
#> Warning: NaNs produced
#> Warning: NaNs produced
#> Warning: NaNs produced
#> Warning: NaNs produced
#> Warning: NaNs produced
#> Warning: NaNs produced
#> Warning: NaNs produced
#> Warning: NaNs produced
#> Warning: NaNs produced
#> Warning: NaNs produced
#> Warning: NaNs produced
#> Warning: NaNs produced
#> Warning: NaNs produced
#> Warning: NaNs produced
#> Warning: NaNs produced
#> Warning: NaNs produced
#> Warning: NaNs produced
#> Warning: NaNs produced
#> Warning: NaNs produced
#> Warning: NaNs produced
#> Warning: NaNs produced
#> Warning: NaNs produced
#> Warning: NaNs produced
#> Warning: NaNs produced
#> Warning: NaNs produced
#> Warning: NaNs produced
#> Warning: NaNs produced
#> Warning: NaNs produced
#> Warning: NaNs produced
#> Warning: NaNs produced
#> Warning: NaNs produced
#> Warning: NaNs produced
#> Warning: NaNs produced
#> Warning: NaNs produced
#> Warning: NaNs produced
#> Warning: NaNs produced
#> Warning: NaNs produced
#> Warning: NaNs produced
#> Warning: NaNs produced
#> Warning: NaNs produced
#> Warning: NaNs produced
#> Warning: NaNs produced
#> Warning: NaNs produced
#> Warning: NaNs produced
#> Warning: NaNs produced
#> Warning: NaNs produced
#> Warning: NaNs produced
#> Warning: NaNs produced
#> Warning: NaNs produced
#> Getting standard errors with the tam.se function: 2.7 secs
#> Error in car::recode(dat[, "var1"], recSt):
#> in recode term: 'D225123:step3' = 'D:step_2251233'
#> message: Error in parse(text = range[[1]][1]) :
#> <text>:1:2: unexpected INCOMPLETE_STRING
#> 1: 'D225123
#> ^
# the same model in conquest (recommended)
defC<- defineModel(dat = dw, items = -c(1:2), model.statement = "item+item*step - sex + item*sex", DIF.var="sex", id = "idstud", analysis.name = "dif_pcm", dir=tempdir())
#> Cannot find conquest 2007 executable file. Please choose manually.
#> Error in file.choose(): file choice cancelled
runC<- runModel(defC, wait=FALSE)
#> Error: object 'defC' not found
resC<- getResults(runC)
#> Error: object 'runC' not found
it <- itemFromRes(resC)
#> Error: object 'resC' not found
shw <- get.shw(file.path(tempdir(), "dif_pcm.shw"), dif.term = "item*sex")
#> Warning: cannot open file '/tmp/RtmpByQecb/dif_pcm.shw': No such file or directory
#> Error in file(file, "r"): cannot open the connection
################################################################################
### Example 12: Norm study within the realm of partial credit models ###
################################################################################
# The following example mimics the steps the required to norm the educational standards
# when the underlying model is a partial credit model.
# load partial credit long format data. This large scale assessment data surveyed in
# an incomplete block design is representative of a domain covered by the new standards.
# We assume thta the sample was drawn from the norm population (reference population)
data(reading)
# transform the data to the wide format
datW <- reshape2::dcast(reading[which(reading[,"type"] != "iglu"),],idstud+sex+language+country~item, value.var = "valueSum")
# combine the two lowest categories to avoid categories with too few observations
datW[,"D205143"] <- car::recode(datW[,"D205143"], "1=0; 2=1; 3=2; 4=3; 5=4")
# partial credit model. We draw 15 plausible values to achieve greater precision
# in estimating the distribution of individuals.
def1 <- defineModel(dat=datW, items = -c(1:4), id=1, irtmodel = "PCM", software="tam", nodes = 21, n.plausible=15)
#> 7 variable(s) are not strictly dichotomous with 0/1 ... Expect a rating scale model or partial credit model.
#> Items(s) 'D025033', 'D201113', 'D204013', 'D205143', 'D224053', 'D225113': 0, 1, 2, 3, 4
#> Items(s) 'D223143': 0, 1, 2, 3, 4, 5
#> 13 subject(s) do not solve any item:
#> P0007 (12 false), P0837 (13 false), P1569 (49 false) ...
#> 20 subject(s) solved each item: P0968 (11 correct), P0843 (13 correct), P1047 (29 correct) ...
#> Dataset is completely linked.
#> Q matrix specifies 1 dimension(s).
run1 <- runModel(def1)
# use ntheta = 40000 for higher precision
res1 <- getResults(run1, ntheta = 40000, theta.model = FALSE)
#> Getting standard errors with the tam.se function: 1.1 secs
#> Getting infit parameters calling tam.fit from getTamInfit: 0.3 secs
#> Getting WLEs calling tam.wle from getTamWles: 0.6 secs
#> |***************|
#> |---------------|
#> Getting PVs calling tam.pv from getTamPVs: 19.1 secs
#> Getting Q3 statistic calling tam.modelfit from getTamQ3: 1.8 secs
# estimate mean and standard deviation of the reference population from the plausible values
# use the long format by toWideFormat = FALSE. Load the eatRep package for mean/SD computation
library(eatRep)
pv1 <- pvFromRes(res1, toWideFormat = FALSE)
mSD <- repMean(datL = pv1, ID="idstud", imp="imp", dependent="value")
#> 1 analyse(s) overall according to: 'group.splits = 0'.
#> Assume unnested structure with 15 imputations.
#>
mSDR <- report2(mSD, round = FALSE)[["plain"]]; dim(mSDR)
#> [1] 3 10
# use mean/SD to transform the item and person parameters. Since the cur scores have not yet
# been established - as they are still being defined in the standard-setting process - we are
# assigning arbitrary values for now. Since the sample can be considered as being drawn from
# the reference population, no equating procedure is necessary. The function is therefore
# simply passed through. The mean and standard deviation of the educational standards metric
# are set to 500 and 100, respectively.
eq <- equat1pl(res1)
#> Found 1 model(s).
#> Equating is executed for each dimension in each model separately.
#> No norm parameter defined ('prmNorm' is missing). Treat current sample as drawn from the reference population.
trnsf<- transformToBista(equatingList=eq, refPop= data.frame(domain="Dim1",
m = mSDR[which(mSDR[,"parameter"] == "mean"),"est"],
sd = mSDR[which(mSDR[,"parameter"] == "sd"),"est"]),
cuts = list(Dim1 = list(values = c(400,600))), vera=FALSE)
#> The 'refPop' data.frame does not include information about reference population mean/SD on Bista metric. Values will be defaulted to mean = 500 and SD = 100.
#> Warning: Skip check whether all competence levels are occupied (due to bayesian plausible values imputation).
res1 <- getResults(run1)
#> Getting standard errors with the tam.se function: 0.7 secs
#> Getting infit parameters calling tam.fit from getTamInfit: 0.2 secs
#> Getting WLEs calling tam.wle from getTamWles: 0.3 secs
#> |*****|
#> |-----|
#> Getting PVs calling tam.pv from getTamPVs: 0.3 secs
#> Getting Q3 statistic calling tam.modelfit from getTamQ3: 0.9 secs
it1 <- itemFromRes(res1)
# males are focus group: initial free estimation of item parameters
def2 <- defineModel(dat=datW[which(datW[,"sex"] == "male"),], items = -c(1:4), id=1, irtmodel = "PCM",software="tam")
#> 7 variable(s) are not strictly dichotomous with 0/1 ... Expect a rating scale model or partial credit model.
#> Items(s) 'D025033', 'D201113', 'D204013', 'D205143', 'D224053', 'D225113': 0, 1, 2, 3, 4
#> Items(s) 'D223143': 0, 1, 2, 3, 4, 5
#> 4 subject(s) do not solve any item:
#> P0007 (12 false), P0107 (12 false), P0837 (13 false) ...
#> 8 subject(s) solved each item: P0968 (11 correct), P0839 (13 correct), P1233 (24 correct) ...
#> Dataset is completely linked.
#> 'gauss' has been chosen for estimation method. Number of nodes was not explicitly specified. Set nodes to 20.
#> Q matrix specifies 1 dimension(s).
run2 <- runModel(def2)
res2 <- getResults(run2)
#> Getting standard errors with the tam.se function: 0.7 secs
#> Getting infit parameters calling tam.fit from getTamInfit: 0.2 secs
#> Getting WLEs calling tam.wle from getTamWles: 0.3 secs
#> |*****|
#> |-----|
#> Getting PVs calling tam.pv from getTamPVs: 0.3 secs
#> Getting Q3 statistic calling tam.modelfit from getTamQ3: 0.9 secs
it2 <- itemFromRes(res2)
# link males to females ... males perform worse
# 10 items with linking dif identified
eq <- equat1pl(results = res2, prmNorm = it1, item = "item", cat="category", value = "est", difBound=.64, iterativ = TRUE)
#> Found 1 model(s).
#> Equating is executed for each dimension in each model separately.
#>
#> ====================================================================================================
#>
#> Model No. 1
#> Model name: not_specified
#> Number of dimension(s): 1
#> Name(s) of dimension(s): Dim1
#> Number of linking items: 128
#> Linking method: Mean.Mean
#>
#> Dimension 'Dim1': 10 of 128 items with linking |DIF| > 0.64 identified.
#> Iteration 1: Exclude item 'D205143_Cat1'.
#> Iteration 2: Exclude item 'D223143_Cat1'.
#> Iteration 3: Exclude item 'D204043_Cat1'.
#> Iteration 4: Exclude item 'D025033_Cat1'.
#> Iteration 5: Exclude item 'D201113_Cat1'.
#> Iteration 6: Exclude item 'D034053_Cat1'.
#> Iteration 7: Exclude item 'D224023_Cat1'.
#> Iteration 8: Exclude item 'D205143_Cat3'.
#> Iteration 9: Exclude item 'D221093_Cat1'.
#> Iteration 10: Exclude item 'D204033_Cat1'.
#> Iteration 11: Exclude item 'D224053_Cat3'.
#>
#> method iter itemExcluded DIF.excluded linking.constant linkerror
#> 1 iterativ 0 -0.221 0.034
#> 2 iterativ 1 D205143_Cat1 1.231 -0.211 0.033
#> 3 iterativ 2 D223143_Cat1 -1.055 -0.220 0.032
#> 4 iterativ 3 D204043_Cat1 1 -0.212 0.031
#> 5 iterativ 4 D025033_Cat1 0.996 -0.204 0.030
#> 6 iterativ 5 D201113_Cat1 0.898 -0.197 0.030
#> 7 iterativ 6 D034053_Cat1 0.849 -0.190 0.029
#> 8 iterativ 7 D224023_Cat1 0.807 -0.183 0.028
#> 9 iterativ 8 D205143_Cat3 0.795 -0.177 0.028
#> 10 iterativ 9 D221093_Cat1 -0.696 -0.183 0.028
#> 11 iterativ 10 D204033_Cat1 -0.668 -0.188 0.027
#> 12 iterativ 11 D224053_Cat3 0.658 -0.183 0.027
#>
# re-calibrate males with anchoring: use the DIF-cleaned set of anchor parameters for calibrating males on the reference scale
# variant 1: use the original item parameters for females and exclude linking dif items (it1_cleaned)
weg <- eq[["items"]][["not_specified"]][["Dim1"]][["info"]][-1,"itemExcluded"]
weg <- eatTools::whereAre(weg, paste(it1[,"item"], it1[,"category"], sep="_"))
#> Found 11 elements.
it1C <- it1[-weg,]
def3A<- defineModel(dat=datW[which(datW[,"sex"] == "male"),], items = -c(1:4), id=1, irtmodel = "PCM",
anchor = it1C, itemCol = "item", valueCol = "est", catCol = "category", software="tam")
#> 7 variable(s) are not strictly dichotomous with 0/1 ... Expect a rating scale model or partial credit model.
#> Items(s) 'D025033', 'D201113', 'D204013', 'D205143', 'D224053', 'D225113': 0, 1, 2, 3, 4
#> Items(s) 'D223143': 0, 1, 2, 3, 4, 5
#> 4 subject(s) do not solve any item:
#> P0007 (12 false), P0107 (12 false), P0837 (13 false) ...
#> 8 subject(s) solved each item: P0968 (11 correct), P0839 (13 correct), P1233 (24 correct) ...
#> Dataset is completely linked.
#> 'gauss' has been chosen for estimation method. Number of nodes was not explicitly specified. Set nodes to 20.
#> 101 common items found in 'anchor' list and data frame.
#> 5 of 106 item(s) of merging variable 'item' from data set 'item response data' not included in data set 'anchor list'.
#> Merging levels are not unique in data set 'anchor list'.
#> Q matrix specifies 1 dimension(s).
run3A<- runModel(def3A)
res3A<- getResults(run3A)
#> Getting standard errors with the tam.se function: 0.7 secs
#> Getting infit parameters calling tam.fit from getTamInfit: 0.2 secs
#> Getting WLEs calling tam.wle from getTamWles: 0.3 secs
#> |*****|
#> |-----|
#> Getting PVs calling tam.pv from getTamPVs: 0.3 secs
#> Getting Q3 statistic calling tam.modelfit from getTamQ3: 0.9 secs
it3A <- itemFromRes(res3A) ### all items except the ones with linking dif with equal item parameters? check
comp <- merge(it1[,c("item", "category", "est")], it3A[,c("item", "category", "est", "offset")], by=c("item", "category"), suffixes = c("_ref", "_foc"))
equal<- na.omit(comp[,c("est_ref", "offset")])
stopifnot(all(equal[,1] == equal[,2])) ### all item parameters without linking dif should be equal
lDif <- subset(comp, !is.na(est_foc)) ### all items with specific focus parameter must be included in linking DIF exclusion list
stopifnot(all(paste(lDif[,"item"], lDif[,"category"], sep="_") %in% eq$items[["not_specified"]][["Dim1"]][["info"]][,"itemExcluded"]))
# variant 2: use the item parameters for males (linking dif items excluded), transformed to the metric of females
def3B<- defineModel(dat=datW[which(datW[,"sex"] == "male"),], items = -c(1:4), id=1, irtmodel = "PCM",
anchor = eq[["items"]][["not_specified"]][["Dim1"]][["cleanedLinkItemPars"]],
itemCol = "item", valueCol = "est", catCol = "category", software="tam")
#> 7 variable(s) are not strictly dichotomous with 0/1 ... Expect a rating scale model or partial credit model.
#> Items(s) 'D025033', 'D201113', 'D204013', 'D205143', 'D224053', 'D225113': 0, 1, 2, 3, 4
#> Items(s) 'D223143': 0, 1, 2, 3, 4, 5
#> 4 subject(s) do not solve any item:
#> P0007 (12 false), P0107 (12 false), P0837 (13 false) ...
#> 8 subject(s) solved each item: P0968 (11 correct), P0839 (13 correct), P1233 (24 correct) ...
#> Dataset is completely linked.
#> 'gauss' has been chosen for estimation method. Number of nodes was not explicitly specified. Set nodes to 20.
#> 101 common items found in 'anchor' list and data frame.
#> 5 of 106 item(s) of merging variable 'item' from data set 'item response data' not included in data set 'anchor list'.
#> Merging levels are not unique in data set 'anchor list'.
#> Q matrix specifies 1 dimension(s).
run3B<- runModel(def3B)
res3B<- getResults(run3B)
#> Getting standard errors with the tam.se function: 0.7 secs
#> Getting WLEs calling tam.wle from getTamWles: 0.3 secs
#> |*****|
#> |-----|
#> Getting PVs calling tam.pv from getTamPVs: 0.3 secs
#> Getting Q3 statistic calling tam.modelfit from getTamQ3: 0.9 secs
it3B <- itemFromRes(res3B) ### all items except the ones with linking dif with equal item parameters? check
link <- eq[["items"]][["not_specified"]][["Dim1"]][["cleanedLinkItemPars"]][,c("item", "category", "est")]
comp <- merge(link, it3B[,c("item", "category", "est", "offset")], by=c("item", "category"), suffixes = c("_ref", "_foc"))
equal<- na.omit(comp[,c("est_ref", "offset")])
stopifnot(all(equal[,1] == equal[,2])) ### all item parameters without linking dif should be equal
lDif <- subset(comp, !is.na(est_foc)) ### all items with specific focus paraeter must be included in linking DIF exclusion list
stopifnot(all(paste(lDif[,"item"], lDif[,"category"], sep="_") %in% eq$items[["not_specified"]][["Dim1"]][["info"]][,"itemExcluded"]))
# transform to Bista metric
eq4 <- equat1pl(results = res3B) ### reference population mean and SD
#> Found 1 model(s).
#> Equating is executed for each dimension in each model separately.
#> No norm parameter defined ('prmNorm' is missing). Treat current sample as drawn from the reference population.
refP <- data.frame(domain = "Dim1", m = 0.0389, sd = 1.07108, stringsAsFactors = FALSE)
cuts <- list ( Dim1 = list(values = 390+0:3*75))
tf4 <- transformToBista(equatingList=eq4, refPop=refP, cuts=cuts, vera = FALSE)
#> The 'refPop' data.frame does not include information about reference population mean/SD on Bista metric. Values will be defaulted to mean = 500 and SD = 100.
#> Warning: Skip check whether all competence levels are occupied (due to bayesian plausible values imputation).
################################################################################
### Example 8.1: same like example 8 (anchored 1pl PCM), now using Conquest ###
################################################################################
# load partial credit long format data
data(reading)
# transform into wide format
datW <- reshape2::dcast(reading[which(reading[,"type"] != "iglu"),],idstud+sex+language+country~item, value.var = "valueSum")
# only some items are partial credit, which ones?
pc.it<- reading[which(reading[,"valueSum"] > 1),"item"] |> unique()
# give values of partial credit items
# few observation of 1-category for item D223143 ... may cause convergence trouble
lapply(datW[,pc.it], FUN = table)
#> $D204013
#>
#> 0 1 2 3 4
#> 24 42 101 263 324
#>
#> $D205143
#>
#> 0 1 2 3 4 5
#> 7 4 31 53 171 466
#>
#> $D201113
#>
#> 0 1 2 3 4
#> 15 113 141 170 104
#>
#> $D225113
#>
#> 0 1 2 3 4
#> 35 75 153 207 78
#>
#> $D224053
#>
#> 0 1 2 3 4
#> 15 25 94 172 225
#>
#> $D025033
#>
#> 0 1 2 3 4
#> 13 39 74 178 233
#>
#> $D223143
#>
#> 0 1 2 3 4 5
#> 23 6 41 100 106 254
#>
# combine the two lowest categories to avoid categories with too few observations
datW[,"D205143"] <- car::recode(datW[,"D205143"], "1=0; 2=1; 3=2; 4=3; 5=4")
# partial credit model ... we consider the female subgroup to be the reference population
# (norm population) that defines the scale and reference item parameters
dFema<- datW[which(datW[,"sex"] == "female"),] ### data for females
lapply(dFema[,pc.it], FUN = table) ### very few observations for some categories ... may cause convergence trouble
#> $D204013
#>
#> 0 1 2 3 4
#> 6 15 42 127 165
#>
#> $D205143
#>
#> 0 1 2 3 4
#> 3 17 20 97 253
#>
#> $D201113
#>
#> 0 1 2 3 4
#> 4 62 66 85 56
#>
#> $D225113
#>
#> 0 1 2 3 4
#> 14 43 90 107 43
#>
#> $D224053
#>
#> 0 1 2 3 4
#> 4 7 34 90 119
#>
#> $D025033
#>
#> 0 1 2 3 4
#> 3 19 39 100 120
#>
#> $D223143
#>
#> 0 1 2 3 4 5
#> 12 2 21 56 68 159
#>
def1 <- defineModel(dat=dFema, items = -c(1:4), id=1, model.statement = "item+item*step", nodes = 21, analysis.name = "pcm_conquest_females", dir=tempdir())
#> Cannot find conquest 2007 executable file. Please choose manually.
#> Error in file.choose(): file choice cancelled
run1 <- runModel(def1)
res1 <- getResults(run1)
#> Getting standard errors with the tam.se function: 0.7 secs
#> Getting infit parameters calling tam.fit from getTamInfit: 0.2 secs
#> Getting WLEs calling tam.wle from getTamWles: 0.6 secs
#> |*****|
#> |-----|
#> Getting PVs calling tam.pv from getTamPVs: 0.3 secs
#> Getting Q3 statistic calling tam.modelfit from getTamQ3: 0.9 secs
it1 <- itemFromRes(res1)
# males are focus group: initial free estimation of item parameters
def2 <- defineModel(dat=datW[which(datW[,"sex"] == "male"),], items = -c(1:4), id=1, model.statement = "item+item*step", nodes = 21, analysis.name = "pcm_conquest_males", dir=tempdir())
#> Cannot find conquest 2007 executable file. Please choose manually.
#> Error in file.choose(): file choice cancelled
run2 <- runModel(def2)
res2 <- getResults(run2)
#> Getting standard errors with the tam.se function: 0.7 secs
#> Getting infit parameters calling tam.fit from getTamInfit: 0.2 secs
#> Getting WLEs calling tam.wle from getTamWles: 0.3 secs
#> |*****|
#> |-----|
#> Getting PVs calling tam.pv from getTamPVs: 0.3 secs
#> Getting Q3 statistic calling tam.modelfit from getTamQ3: 0.9 secs
it2 <- itemFromRes(res2)
# link males to females ... males perform worse
# 11 items with linking dif identified
eq <- equat1pl(results = res2, prmNorm = it1, item = "item", cat="category", value = "est", difBound=.64, iterativ = TRUE)
#> Found 1 model(s).
#> Equating is executed for each dimension in each model separately.
#>
#> ====================================================================================================
#>
#> Model No. 1
#> Model name: not_specified
#> Number of dimension(s): 1
#> Name(s) of dimension(s): Dim1
#> Number of linking items: 128
#> Linking method: Mean.Mean
#>
#> Dimension 'Dim1': 10 of 128 items with linking |DIF| > 0.64 identified.
#> Iteration 1: Exclude item 'D205143_Cat1'.
#> Iteration 2: Exclude item 'D223143_Cat1'.
#> Iteration 3: Exclude item 'D204043_Cat1'.
#> Iteration 4: Exclude item 'D025033_Cat1'.
#> Iteration 5: Exclude item 'D201113_Cat1'.
#> Iteration 6: Exclude item 'D034053_Cat1'.
#> Iteration 7: Exclude item 'D224023_Cat1'.
#> Iteration 8: Exclude item 'D205143_Cat3'.
#> Iteration 9: Exclude item 'D221093_Cat1'.
#> Iteration 10: Exclude item 'D204033_Cat1'.
#> Iteration 11: Exclude item 'D224053_Cat3'.
#>
#> method iter itemExcluded DIF.excluded linking.constant linkerror
#> 1 iterativ 0 -0.221 0.034
#> 2 iterativ 1 D205143_Cat1 1.231 -0.211 0.033
#> 3 iterativ 2 D223143_Cat1 -1.055 -0.220 0.032
#> 4 iterativ 3 D204043_Cat1 1 -0.212 0.031
#> 5 iterativ 4 D025033_Cat1 0.996 -0.204 0.030
#> 6 iterativ 5 D201113_Cat1 0.898 -0.197 0.030
#> 7 iterativ 6 D034053_Cat1 0.849 -0.190 0.029
#> 8 iterativ 7 D224023_Cat1 0.807 -0.183 0.028
#> 9 iterativ 8 D205143_Cat3 0.795 -0.177 0.028
#> 10 iterativ 9 D221093_Cat1 -0.696 -0.183 0.028
#> 11 iterativ 10 D204033_Cat1 -0.668 -0.188 0.027
#> 12 iterativ 11 D224053_Cat3 0.658 -0.183 0.027
#>
# re-calibrate males with anchoring: use the DIF-cleaned set of anchor parameters for calibrating males on the reference scale
# variant 1: use the original item parameters for females and exclude linking dif items (it1_cleaned)
# please note that to date in Conquest only dichotomous items can be used for anchoring
weg <- eq[["items"]][["pcm_conquest_males"]][["Dim1"]][["info"]][-1,"itemExcluded"]
weg <- eatTools::whereAre(weg, paste(it1[,"item"], it1[,"category"], sep="_"))
#> Error in eatTools::whereAre(weg, paste(it1[, "item"], it1[, "category"], sep = "_")): Assertion on 'a' failed: Must be of type 'vector', not 'NULL'.
it1C <- it1[-weg,]
#> Error in -weg: invalid argument to unary operator
pcmIt<- unique(subset(it1C, category == "Cat2")[,"item"]) ### additionally exclude pcm items
it1C <- it1C[-eatTools::whereAre(pcmIt, it1C[,"item"]),]
#> Found 23 elements.
def3A<- defineModel(dat=datW[which(datW[,"sex"] == "male"),], items = -c(1:4), id=1,
model.statement = "item+item*step", nodes = 21, analysis.name = "pcm_conquest_males_anchored1", dir=tempdir(),
anchor = it1C[,c("item", "est")])
#> Cannot find conquest 2007 executable file. Please choose manually.
#> Error in file.choose(): file choice cancelled
run3A<- runModel(def3A)
res3A<- getResults(run3A)
#> Getting standard errors with the tam.se function: 0.7 secs
#> Getting WLEs calling tam.wle from getTamWles: 0.3 secs
#> |*****|
#> |-----|
#> Getting PVs calling tam.pv from getTamPVs: 0.3 secs
#> Getting Q3 statistic calling tam.modelfit from getTamQ3: 0.9 secs
it3A <- itemFromRes(res3A) ### all dichotomous items except the ones with linking dif with equal item parameters? check
comp <- merge(it1[,c("item", "category", "est")], it3A[,c("item", "category", "est", "offset")], by=c("item", "category"), suffixes = c("_ref", "_foc"))
equal<- na.omit(comp[,c("est_ref", "offset")])
stopifnot(all(equal[,1] == equal[,2])) ### all item parameters without linking dif should be equal
### TO DO: all items with specific focus parameter must be included in linking DIF exclusion list or must be pcm items
# variant 2: use the item parameters for males (linking dif items excluded), transformed to the metric of females
ank3B<- eq[["items"]][["pcm_conquest_males"]][["Dim1"]][["cleanedLinkItemPars"]]
pcmIt<- unique(subset(ank3B, category == "Cat2")[,"item"]) ### additionally exclude pcm items
#> Error: object 'category' not found
ank3B<- ank3B[-eatTools::whereAre(pcmIt, ank3B[,"item"]),]
#> Error in eatTools::whereAre(pcmIt, ank3B[, "item"]): Assertion on 'b' failed: Must be of type 'vector', not 'NULL'.
def3B<- defineModel(dat=datW[which(datW[,"sex"] == "male"),], items = -c(1:4), id=1,
model.statement = "item+item*step", analysis.name = "pcm_conquest_males_anchored2",
dir=tempdir(), anchor = ank3B[,c("item", "est")])
#> Cannot find conquest 2007 executable file. Please choose manually.
#> Error in file.choose(): file choice cancelled
run3B<- runModel(def3B)
res3B<- getResults(run3B)
#> Getting standard errors with the tam.se function: 0.7 secs
#> Getting WLEs calling tam.wle from getTamWles: 0.3 secs
#> |*****|
#> |-----|
#> Getting PVs calling tam.pv from getTamPVs: 0.3 secs
#> Getting Q3 statistic calling tam.modelfit from getTamQ3: 0.9 secs
it3B <- itemFromRes(res3B) ### all items except the ones with linking dif with equal item parameters? check
link <- eq[["items"]][["pcm_conquest_males"]][["Dim1"]][["cleanedLinkItemPars"]][,c("item", "category", "est")]
comp <- merge(link, it3B[,c("item", "category", "est", "offset")], by=c("item", "category"), suffixes = c("_ref", "_foc"))
#> Error in fix.by(by.x, x): 'by' must specify uniquely valid columns
equal<- na.omit(comp[,c("est_ref", "offset")])
stopifnot(all(equal[,1] == equal[,2])) ### all item parameters without linking dif should be equal
### TO DO: all items with specific focus parameter must be included in linking DIF exclusion list or must be pcm items
# transform to Bista metric
eq4 <- equat1pl(results = res3B) ### reference population mean and SD
#> Found 1 model(s).
#> Equating is executed for each dimension in each model separately.
#> No norm parameter defined ('prmNorm' is missing). Treat current sample as drawn from the reference population.
refP <- data.frame(domain = "Dim1", m = 0.0389, sd = 1.07108, stringsAsFactors = FALSE)
cuts <- list ( Dim1 = list(values = 390+0:3*75))
tf4 <- transformToBista(equatingList=eq4, refPop=refP, cuts=cuts, vera = FALSE)
#> The 'refPop' data.frame does not include information about reference population mean/SD on Bista metric. Values will be defaulted to mean = 500 and SD = 100.
#> Warning: Skip check whether all competence levels are occupied (due to bayesian plausible values imputation).
################################################################################
### Example 9: (restricted) generalized partial credit model with anchoring ###
################################################################################
# load partial credit long format data
data(reading)
# transform into wide format
datW <- reshape2::dcast(reading[which(reading[,"type"] != "iglu"),],idstud+sex+language+country~item, value.var = "valueSum")
# combine the two lowest categories to avoid categories with too few observations
datW[,"D205143"] <- car::recode(datW[,"D205143"], "1=0; 2=1; 3=2; 4=3; 5=4")
# restricted generalized partial credit model ... we consider the female subgroup to be the reference population
# we assume that task D223 and D224 are created for students with special educational need and therefore have
# different slope. Non-SPF items should have slope = 1
dFema<- datW[which(datW[,"sex"] == "female"),] ### data for females
items<- colnames(dFema)[-c(1:4)]
items<- data.frame(item = items, task = substr(items,1,4), slopeGrp = as.numeric(substr(items,1,4) %in% c("D223", "D224")), stringsAsFactors = FALSE) |> dplyr::mutate(slope = car::recode(slopeGrp, "0=1; else=NA"))
items[,"dim"] <- car::recode(substr(items[,"item"], 1,2),"'D0'='norm'; 'D2'='pilot'")
items<- data.frame ( items, model.matrix(~dim-1, data = items), stringsAsFactors = FALSE)
def1T<- defineModel(dat=dFema, items = items[,"item"], id=1, irtmodel = "GPCM.groups", software="tam", fixSlopeMat = na.omit(items[,c("item", "slope")]), est.slopegroups = items[,c("item", "slopeGrp")], nodes = 21, fac.oldxsi =0.6, increment.factor=1.05)
#> 7 variable(s) are not strictly dichotomous with 0/1 ... Expect a rating scale model or partial credit model.
#> Items(s) 'D025033', 'D201113', 'D204013', 'D205143', 'D224053', 'D225113': 0, 1, 2, 3, 4
#> Items(s) 'D223143': 0, 1, 2, 3, 4, 5
#> 2 subject(s) do not solve any item:
#> P1296 (24 false), P1299 (24 false), P1299 (24 false) ...
#> 12 subject(s) solved each item: P0081 (12 correct), P0869 (13 correct), P1047 (29 correct) ...
#> Dataset is completely linked.
#> Q matrix specifies 1 dimension(s).
#> Warning: To date, fixing slopes only works for dichotomous unidimensional or between-item multidimensional models.
#> Following 22 items in dataset without fixed slopes in 'fixSlopeMat'. Slope(s) will be estimated freely.
#> D223013, D223023, D223033, D223043, D223063, D223073, D223083, D223103, D223113, D223123, D223133, D223143, D224013, D224023, D224033, D224043, D224053, D224063, D224083, D224093, D224103, D224113
run1T<- runModel(def1T)
res1T<- getResults(run1T)
#> Getting standard errors with the tam.se function: 1 secs
#> Getting infit parameters calling tam.fit from getTamInfit: 0.2 secs
#> Getting WLEs calling tam.wle from getTamWles: 0.3 secs
#> |*****|
#> |-----|
#> Getting PVs calling tam.pv from getTamPVs: 0.3 secs
#> Getting Q3 statistic calling tam.modelfit from getTamQ3: 0.9 secs
it1T <- itemFromRes(res1T)
# males are focus group: initial free estimation of item parameters
def2T<- defineModel(dat=datW[which(datW[,"sex"] == "male"),], items = -c(1:4), id=1, irtmodel = "GPCM.groups",software="tam", fixSlopeMat = na.omit(items[,c("item", "slope")]), est.slopegroups = items[,c("item", "slopeGrp")], nodes = 21)
#> 7 variable(s) are not strictly dichotomous with 0/1 ... Expect a rating scale model or partial credit model.
#> Items(s) 'D025033', 'D201113', 'D204013', 'D205143', 'D224053', 'D225113': 0, 1, 2, 3, 4
#> Items(s) 'D223143': 0, 1, 2, 3, 4, 5
#> 4 subject(s) do not solve any item:
#> P0007 (12 false), P0107 (12 false), P0837 (13 false) ...
#> 8 subject(s) solved each item: P0968 (11 correct), P0839 (13 correct), P1233 (24 correct) ...
#> Dataset is completely linked.
#> Q matrix specifies 1 dimension(s).
#> Warning: To date, fixing slopes only works for dichotomous unidimensional or between-item multidimensional models.
#> Following 22 items in dataset without fixed slopes in 'fixSlopeMat'. Slope(s) will be estimated freely.
#> D223013, D223023, D223033, D223043, D223063, D223073, D223083, D223103, D223113, D223123, D223133, D223143, D224013, D224023, D224033, D224043, D224053, D224063, D224083, D224093, D224103, D224113
run2T<- runModel(def2T)
res2T<- getResults(run2T)
#> Getting standard errors with the tam.se function: 0.7 secs
#> Getting infit parameters calling tam.fit from getTamInfit: 0.2 secs
#> Getting WLEs calling tam.wle from getTamWles: 0.3 secs
#> |*****|
#> |-----|
#> Getting PVs calling tam.pv from getTamPVs: 0.3 secs
#> Getting Q3 statistic calling tam.modelfit from getTamQ3: 0.9 secs
it2T <- itemFromRes(res2T)
unique(round(it1T[,"estSlope"],3)); unique(round(it2T[,"estSlope"],3)) ### average discrimination (reg vs. spf) differs for males, but not for females
#> [1] 1.000 1.001
#> [1] 1.000 0.858
# use only slope=1 items for 1pl linking
eq <- equat1pl(results = res2T, prmNorm = it1T[which(it1T[,"estSlope"] ==1),], item = "item", cat="category", value = "est", difBound=.64, iterativ = TRUE)
#> Found 1 model(s).
#> Equating is executed for each dimension in each model separately.
#>
#> ====================================================================================================
#>
#> Model No. 1
#> Model name: not_specified
#> Number of dimension(s): 1
#> Name(s) of dimension(s): Dim1
#> Number of linking items: 99
#> Linking method: Mean.Mean
#>
#> Dimension 'Dim1': 8 of 99 items with linking |DIF| > 0.64 identified.
#> Iteration 1: Exclude item 'D205143_Cat1'.
#> Iteration 2: Exclude item 'D025033_Cat1'.
#> Iteration 3: Exclude item 'D204043_Cat1'.
#> Iteration 4: Exclude item 'D201113_Cat1'.
#> Iteration 5: Exclude item 'D034053_Cat1'.
#> Iteration 6: Exclude item 'D221093_Cat1'.
#> Iteration 7: Exclude item 'D205143_Cat3'.
#> Iteration 8: Exclude item 'D204033_Cat1'.
#>
#> method iter itemExcluded DIF.excluded linking.constant linkerror
#> 1 iterativ 0 -0.239 0.039
#> 2 iterativ 1 D205143_Cat1 1.3 -0.226 0.037
#> 3 iterativ 2 D025033_Cat1 1.05 -0.215 0.036
#> 4 iterativ 3 D204043_Cat1 1.01 -0.205 0.034
#> 5 iterativ 4 D201113_Cat1 0.874 -0.196 0.033
#> 6 iterativ 5 D034053_Cat1 0.851 -0.187 0.033
#> 7 iterativ 6 D221093_Cat1 -0.687 -0.194 0.032
#> 8 iterativ 7 D205143_Cat3 0.676 -0.187 0.032
#> 9 iterativ 8 D204033_Cat1 -0.666 -0.194 0.031
#>
# variant 1
# use the DIF-cleaned set of original anchor parameters for calibrating males on the reference scale
weg <- eq[["items"]][["not_specified"]][["Dim1"]][["info"]][-1,"itemExcluded"]
weg <- eatTools::whereAre(weg, paste(it1T[,"item"], it1T[,"category"], sep="_"))
#> Found 8 elements.
it1C <- subset(it1T[-weg,], estSlope == 1)
def3T<- defineModel(eatTools::na_omit_selection(dat=datW[which(datW[,"sex"] == "male"),],"language"),
items = -c(1:4), id=1, irtmodel = "GPCM.groups", anchor = it1C, HG.var = c("language", "country"),
itemCol = "item", valueCol = "est", catCol = "category", fixSlopeMat = na.omit(items[,c("item", "slope")]),
qMatrix = items[,c("item", "dimnorm", "dimpilot")], est.slopegroups = items[,c("item", "slopeGrp")], nodes = 21, software="tam")
#> 7 variable(s) are not strictly dichotomous with 0/1 ... Expect a rating scale model or partial credit model.
#> Items(s) 'D025033', 'D201113', 'D204013', 'D205143', 'D224053', 'D225113': 0, 1, 2, 3, 4
#> Items(s) 'D223143': 0, 1, 2, 3, 4, 5
#> Background variable(s) 'language', 'country' of class
#> 'factor', 'factor' will be converted to indicator variables.
#> Variable 'language' was converted to 2 indicator(s) with name(s) 'languagenativeAndOther', 'languageother'.
#> Variable 'country' was converted to 8 indicator(s) with name(s) 'countryCaladan', 'countryDisneyland', 'countryGreenland', 'countryMordor', 'countryRomulus', 'countryTatooine', 'countrytheShire', 'countryVulcan'.
#> 4 subject(s) do not solve any item:
#> P0007 (12 false), P0107 (12 false), P0837 (13 false) ...
#> 8 subject(s) solved each item: P0968 (11 correct), P0839 (13 correct), P1233 (24 correct) ...
#> Dataset is completely linked.
#> 80 common items found in 'anchor' list and data frame.
#> 26 of 106 item(s) of merging variable 'item' from data set 'item response data' not included in data set 'anchor list'.
#> Merging levels are not unique in data set 'anchor list'.
#> Q matrix specifies 2 dimension(s).
#> Warning: To date, fixing slopes only works for dichotomous unidimensional or between-item multidimensional models.
#> Following 22 items in dataset without fixed slopes in 'fixSlopeMat'. Slope(s) will be estimated freely.
#> D223013, D223023, D223033, D223043, D223063, D223073, D223083, D223103, D223113, D223123, D223133, D223143, D224013, D224023, D224033, D224043, D224053, D224063, D224083, D224093, D224103, D224113
run3T<- runModel(def3T)
#> Error: 'timeFormat' is not an exported object from 'namespace:eatTools'
res3T<- getResults(run3T)
#> Error: object 'run3T' not found
it3T <- itemFromRes(res3T) ### all items except the ones with linking dif with equal item parameters? check
#> Error: object 'res3T' not found
comp <- merge(subset(it1T[,c("item", "category", "est", "estSlope")],estSlope == 1), it3T[,c("item", "category", "est", "offset")], by=c("item", "category"), all=TRUE, suffixes = c("_ref", "_foc"))
#> Error: object 'it3T' not found
equal<- na.omit(comp[,c("est_ref", "offset")])
stopifnot(all(equal[,1] == equal[,2])) ### all item parameters without linking dif should be equal
lDif <- subset(comp, !is.na(est_foc))
stopifnot(all(paste(subset(lDif, !is.na(est_ref))[,"item"], subset(lDif, !is.na(est_ref))[,"category"], sep="_") %in% eq$items[["not_specified"]][["Dim1"]][["info"]][,"itemExcluded"]))
#> Error: all(paste(subset(lDif, !is.na(est_ref))[, "item"], subset(lDif, .... is not TRUE
# variant 2: use the 1pl item parameters for males (linking dif items excluded), transformed to the metric of females
def4T<- defineModel(eatTools::na_omit_selection(dat=datW[which(datW[,"sex"] == "male"),],"language"),
items = -c(1:4), id=1, irtmodel = "GPCM.groups", anchor = eq[["items"]][["not_specified"]][["Dim1"]][["cleanedLinkItemPars"]],
HG.var = c("language", "country"), itemCol = "item", valueCol = "est", catCol = "category", fixSlopeMat = na.omit(items[,c("item", "slope")]),
qMatrix = items[,c("item", "dimnorm", "dimpilot")], est.slopegroups = items[,c("item", "slopeGrp")], nodes = 21, software="tam")
#> 7 variable(s) are not strictly dichotomous with 0/1 ... Expect a rating scale model or partial credit model.
#> Items(s) 'D025033', 'D201113', 'D204013', 'D205143', 'D224053', 'D225113': 0, 1, 2, 3, 4
#> Items(s) 'D223143': 0, 1, 2, 3, 4, 5
#> Background variable(s) 'language', 'country' of class
#> 'factor', 'factor' will be converted to indicator variables.
#> Variable 'language' was converted to 2 indicator(s) with name(s) 'languagenativeAndOther', 'languageother'.
#> Variable 'country' was converted to 8 indicator(s) with name(s) 'countryCaladan', 'countryDisneyland', 'countryGreenland', 'countryMordor', 'countryRomulus', 'countryTatooine', 'countrytheShire', 'countryVulcan'.
#> 4 subject(s) do not solve any item:
#> P0007 (12 false), P0107 (12 false), P0837 (13 false) ...
#> 8 subject(s) solved each item: P0968 (11 correct), P0839 (13 correct), P1233 (24 correct) ...
#> Dataset is completely linked.
#> 80 common items found in 'anchor' list and data frame.
#> 26 of 106 item(s) of merging variable 'item' from data set 'item response data' not included in data set 'anchor list'.
#> Merging levels are not unique in data set 'anchor list'.
#> Q matrix specifies 2 dimension(s).
#> Warning: To date, fixing slopes only works for dichotomous unidimensional or between-item multidimensional models.
#> Following 22 items in dataset without fixed slopes in 'fixSlopeMat'. Slope(s) will be estimated freely.
#> D223013, D223023, D223033, D223043, D223063, D223073, D223083, D223103, D223113, D223123, D223133, D223143, D224013, D224023, D224033, D224043, D224053, D224063, D224083, D224093, D224103, D224113
run4T<- runModel(def4T)
#> Error: 'timeFormat' is not an exported object from 'namespace:eatTools'
res4T<- getResults(run4T)
#> Error: object 'run4T' not found
it4T <- itemFromRes(res4T) ### all items except the ones with linking dif with equal item parameters? check
#> Error: object 'res4T' not found
link <- eq[["items"]][["not_specified"]][["Dim1"]][["cleanedLinkItemPars"]][,c("item", "category", "est")]
comp <- merge(link, it4T[,c("item", "category", "est", "offset")], by=c("item", "category"), suffixes = c("_ref", "_foc"), all=TRUE)
#> Error: object 'it4T' not found
equal<- na.omit(comp[,c("est_ref", "offset")])
stopifnot(all(equal[,1] == equal[,2])) ### all item parameters without linking dif should be equal
lDif <- subset(comp, !is.na(est_foc)) ### all items with specific focus paraeter must be included in linking DIF exclusion list
stopifnot(all(paste(subset(lDif, !is.na(est_ref))[,"item"], subset(lDif, !is.na(est_ref))[,"category"], sep="_") %in% eq$items[["not_specified"]][["Dim1"]][["info"]][,"itemExcluded"]))
#> Error: all(paste(subset(lDif, !is.na(est_ref))[, "item"], subset(lDif, .... is not TRUE
# transform to Bista metric
eq4 <- equat1pl(results = res4T) ### reference population mean and SD
#> Error: object 'res4T' not found
refP <- data.frame(domain = c("dimnorm", "dimpilot"), m = 0.0389, sd = 1.07108, stringsAsFactors = FALSE)
cuts <- list ( dimnorm = list(values = 390+0:3*75), dimpilot = list(values = 390+0:3*75))
tf4 <- transformToBista(equatingList=eq4, refPop=refP, cuts=cuts, vera = FALSE)
#> Error in generateOrCheckRefPop(equatingList = equatingList, refPop = refPop, mods = mods, dims = dims, isRunM = isRunM, id = id, weights = weights, defaultM = defaultM, defaultSD = defaultSD): Following 1 dimension(s) not included in 'refPop': 'Dim1'.
################################################################################
### Example 10: GPCM, similar like example 9, but now using mirt ###
################################################################################
# load package Hmisc if necesary
if(!"Hmisc" %in% .packages()) {library(Hmisc)}
#>
#> Attaching package: ‘Hmisc’
#> The following object is masked from ‘package:survey’:
#>
#> deff
#> The following objects are masked from ‘package:base’:
#>
#> format.pval, units
# load partial credit long format data
data(reading)
# transform into wide format
datW <- reshape2::dcast(reading[which(reading[,"type"] != "iglu"),],idstud+sex+language+country~item, value.var = "valueSum")
# only some items are partial credit, which ones?
pc.it<- reading[which(reading[,"valueSum"] > 1),"item"] |> unique()
# combine the two lowest categories to avoid categories with too few observations
datW[,"D205143"] <- car::recode(datW[,"D205143"], "1=0; 2=1; 3=2; 4=3; 5=4")
# restricted generalized partial credit model ... we consider the female subgroup to be the reference population
# we assume that task D223 and D224 are created for students with special educational need and therefore have
# different slope (gpcm). Non-SPF items should have slope = 1
dFema<- datW[which(datW[,"sex"] == "female"),] ### data for females
items<- colnames(dFema)[-c(1:4)]
items<- data.frame(item = items, task = substr(items,1,4), slopeGrp = as.numeric(substr(items,1,4) %in% c("D223", "D224")), stringsAsFactors = FALSE) |> dplyr::mutate(irtmod = NA)
# add model information to 'item'
items[intersect(intersect(which(items[,"slopeGrp"] == 0), which(items[,"task"] %nin% c("D223", "D224"))), which(items[,"item"] %nin% pc.it)),"irtmod"] <- "Rasch"
items[intersect(intersect(which(items[,"slopeGrp"] == 1), which(items[,"task"] %in% c("D223", "D224"))), which(items[,"item"] %nin% pc.it)),"irtmod"] <- "2PL"
# this is partial credit which is specified in 'mirt' with 'Rasch' statement as the slope equals 1 like in dichotomous Rasch models
items[intersect(intersect(which(items[,"slopeGrp"] == 0), which(items[,"task"] %nin% c("D223", "D224"))), which(items[,"item"] %in% pc.it)),"irtmod"] <- "Rasch"
# generalized partial credit for unidimensional models
items[intersect(which(items[,"slopeGrp"] == 1), which(items[,"item"] %in% pc.it)),"irtmod"] <- "gpcm"
# define arbitrary dimensions
items[,"dim"] <- car::recode(substr(items[,"item"], 1,2),"'D0'='norm'; 'D2'='pilot'")
items<- data.frame ( items, model.matrix(~dim-1, data = items), stringsAsFactors = FALSE)
# procedure similar to conquest/tam
def1 <- defineModel(dat=eatTools::na_omit_selection(dFema,varsToOmitIfNA="language"), items = items[,"item"],
id=1,irtmodel = items[,c("item", "irtmod")], software="mirt")
#> 7 variable(s) are not strictly dichotomous with 0/1 ... Expect a rating scale model or partial credit model.
#> Items(s) 'D025033', 'D201113', 'D204013', 'D205143', 'D224053', 'D225113': 0, 1, 2, 3, 4
#> Items(s) 'D223143': 0, 1, 2, 3, 4, 5
#> 12 subject(s) solved each item: P0081 (12 correct), P0869 (13 correct), P1047 (29 correct) ...
#> Dataset is completely linked.
#> Specifying 'method' and 'nodes' not yet implemented for 'mirt'.
#> Q matrix specifies 1 dimension(s).
skel1<- runModel(def1, onlySkeleton = TRUE) ### check the model "skeleton"
run1 <- runModel(def1) ### run the model finally
#> Modify skeleton ...
#>
#>
#> Calculating information matrix...
res1 <- getResults(run1)
#> Getting WLEs calling fscores(method="WLE") from getMirtWles: 11.4 secs
#> Getting PVs calling fscores from getMirtPVs: 0.2 secs
it1 <- itemFromRes(res1)
# males are focus group: initial free estimation of item parameters
def2 <- defineModel(dat=eatTools::na_omit_selection(datW[which(datW[,"sex"] == "male"),],varsToOmitIfNA="language"),
items = -c(1:4), id=1, irtmodel = items[,c("item", "irtmod")],software="mirt")
#> 7 variable(s) are not strictly dichotomous with 0/1 ... Expect a rating scale model or partial credit model.
#> Items(s) 'D025033', 'D201113', 'D204013', 'D205143', 'D224053', 'D225113': 0, 1, 2, 3, 4
#> Items(s) 'D223143': 0, 1, 2, 3, 4, 5
#> 4 subject(s) do not solve any item:
#> P0007 (12 false), P0107 (12 false), P0837 (13 false) ...
#> 8 subject(s) solved each item: P0968 (11 correct), P0839 (13 correct), P1233 (24 correct) ...
#> Dataset is completely linked.
#> Specifying 'method' and 'nodes' not yet implemented for 'mirt'.
#> Q matrix specifies 1 dimension(s).
run2 <- runModel(def2)
#> Modify skeleton ...
#>
#>
#> Calculating information matrix...
res2 <- getResults(run2)
#> Getting WLEs calling fscores(method="WLE") from getMirtWles: 10.5 secs
it2 <- itemFromRes(res2)
# link males to females, using only 1pl items ... males perform worse. 10 items with linking dif identified
eq <- equat1pl(results = res2, prmNorm = subset(it1,estSlope ==1), item = "item", cat="category", value = "est", difBound=.64, iterativ = TRUE)
#> Found 1 model(s).
#> Equating is executed for each dimension in each model separately.
#>
#> ====================================================================================================
#>
#> Model No. 1
#> Model name: not_specified
#> Number of dimension(s): 1
#> Name(s) of dimension(s): Dim1
#> Number of linking items: 84
#> Linking method: Mean.Mean
#>
#> Dimension 'Dim1': 6 of 84 items with linking |DIF| > 0.64 identified.
#> Iteration 1: Exclude item 'D205143_Cat1'.
#> Iteration 2: Exclude item 'D034053_Cat1'.
#> Iteration 3: Exclude item 'D204043_Cat1'.
#> Iteration 4: Exclude item 'D025033_Cat1'.
#> Iteration 5: Exclude item 'D204033_Cat1'.
#> Iteration 6: Exclude item 'D221093_Cat1'.
#>
#> method iter itemExcluded DIF.excluded linking.constant linkerror
#> 1 iterativ 0 -0.263 0.044
#> 2 iterativ 1 D205143_Cat1 1.841 -0.241 0.039
#> 3 iterativ 2 D034053_Cat1 0.907 -0.230 0.037
#> 4 iterativ 3 D204043_Cat1 0.916 -0.219 0.036
#> 5 iterativ 4 D025033_Cat1 0.776 -0.209 0.035
#> 6 iterativ 5 D204033_Cat1 -0.68 -0.218 0.035
#> 7 iterativ 6 D221093_Cat1 -0.687 -0.227 0.034
#>
# re-calibrate males with anchoring: use the DIF-cleaned set of anchor parameters for calibrating males on the reference scale
# variant 1: use the original (1pl) item parameters for females and exclude linking dif items (it1_cleaned)
weg <- eq[["items"]][["not_specified"]][["Dim1"]][["info"]][-1,"itemExcluded"]
weg <- eatTools::whereAre(weg, paste(it1[,"item"], it1[,"category"], sep="_"))
#> Found 6 elements.
it1C <- subset(it1[-weg,], estSlope == 1)
def3A<- defineModel(dat=eatTools::na_omit_selection(datW[which(datW[,"sex"] == "male"),],varsToOmitIfNA="language"), items = -c(1:4), id=1, irtmodel = items[,c("item", "irtmod")],
anchor = it1C, itemCol = "item", valueCol = "est", catCol = "category", software="mirt")
#> 7 variable(s) are not strictly dichotomous with 0/1 ... Expect a rating scale model or partial credit model.
#> Items(s) 'D025033', 'D201113', 'D204013', 'D205143', 'D224053', 'D225113': 0, 1, 2, 3, 4
#> Items(s) 'D223143': 0, 1, 2, 3, 4, 5
#> 4 subject(s) do not solve any item:
#> P0007 (12 false), P0107 (12 false), P0837 (13 false) ...
#> 8 subject(s) solved each item: P0968 (11 correct), P0839 (13 correct), P1233 (24 correct) ...
#> Dataset is completely linked.
#> Specifying 'method' and 'nodes' not yet implemented for 'mirt'.
#> 78 common items found in 'anchor' list and data frame.
#> 28 of 106 item(s) of merging variable 'item' from data set 'item response data' not included in data set 'anchor list'.
#> Q matrix specifies 1 dimension(s).
run3A<- runModel(def3A)
#> Modify skeleton ...
#>
#>
#> Calculating information matrix...
res3A<- getResults(run3A)
#> Getting WLEs calling fscores(method="WLE") from getMirtWles: 10.6 secs
it3A <- itemFromRes(res3A) ### all items except the ones with linking dif with equal item parameters? check
comp <- merge(subset(it1, estSlope==1)[,c("item", "category", "est")], subset(it3A, !is.na(offset))[,c("item", "category", "est", "offset")], by=c("item", "category"), suffixes = c("_ref", "_foc"))
stopifnot(all(round(comp[,"est_ref"],5) == round(comp[,"offset"],5))) ### all item parameters without linking dif should be equal
# all items with specific focus parameter must be included in linking DIF exclusion list
stopifnot(all((paste(comp[,"item"], comp[,"category"],sep="_") %in% eq$items[["not_specified"]][["Dim1"]][["info"]][-1,"itemExcluded"]) == FALSE))
# variant 2: use the item parameters for males (linking dif items excluded), transformed to the metric of females
def3B<- defineModel(dat=eatTools::na_omit_selection(datW[which(datW[,"sex"] == "male"),],varsToOmitIfNA="language"), items = -c(1:4), id=1, irtmodel = items[,c("item", "irtmod")],
anchor = eq[["items"]][["not_specified"]][["Dim1"]][["cleanedLinkItemPars"]],
itemCol = "item", valueCol = "est", catCol = "category", software="mirt")
#> 7 variable(s) are not strictly dichotomous with 0/1 ... Expect a rating scale model or partial credit model.
#> Items(s) 'D025033', 'D201113', 'D204013', 'D205143', 'D224053', 'D225113': 0, 1, 2, 3, 4
#> Items(s) 'D223143': 0, 1, 2, 3, 4, 5
#> 4 subject(s) do not solve any item:
#> P0007 (12 false), P0107 (12 false), P0837 (13 false) ...
#> 8 subject(s) solved each item: P0968 (11 correct), P0839 (13 correct), P1233 (24 correct) ...
#> Dataset is completely linked.
#> Specifying 'method' and 'nodes' not yet implemented for 'mirt'.
#> 78 common items found in 'anchor' list and data frame.
#> 28 of 106 item(s) of merging variable 'item' from data set 'item response data' not included in data set 'anchor list'.
#> Q matrix specifies 1 dimension(s).
run3B<- runModel(def3B)
#> Modify skeleton ...
#>
#>
#> Calculating information matrix...
res3B<- getResults(run3B)
#> Getting WLEs calling fscores(method="WLE") from getMirtWles: 10.8 secs
it3B <- itemFromRes(res3B) ### all items except the ones with linking dif with equal item parameters? check
link <- eq[["items"]][["not_specified"]][["Dim1"]][["cleanedLinkItemPars"]][,c("item", "category", "est", "estSlope")]
stopifnot(all(link[,"estSlope"] == 1))
comp <- merge(link, it3B[,c("item", "category", "est", "offset")], by=c("item", "category"), suffixes = c("_ref", "_foc"))
stopifnot(all(round(comp[,"est_ref"],5) == round(comp[,"offset"],5))) ### all item parameters without linking dif should be equal
# all items with specific focus parameter must be included in linking DIF exclusion list
stopifnot(all((paste(comp[,"item"], comp[,"category"],sep="_") %in% eq$items[["not_specified"]][["Dim1"]][["info"]][-1,"itemExcluded"]) == FALSE))
# transform to Bista metric
eq4 <- equat1pl(results = res3B) ### reference population mean and SD
#> Found 1 model(s).
#> Equating is executed for each dimension in each model separately.
#> No norm parameter defined ('prmNorm' is missing). Treat current sample as drawn from the reference population.
refP <- data.frame(domain = "Dim1", m = 0.0389, sd = 1.07108, stringsAsFactors = FALSE)
cuts <- list ( Dim1 = list(values = 390+0:3*75))
tf4 <- transformToBista(equatingList=eq4, refPop=refP, cuts=cuts, vera = FALSE)
#> Warning: Cannot identify student identifier variable (possibly because 'resultsObj' was created by an older version of 'eatModel'). student id variable will be defaulted to 'idstud'.
#> The 'refPop' data.frame does not include information about reference population mean/SD on Bista metric. Values will be defaulted to mean = 500 and SD = 100.
#> Warning: Skip check whether all competence levels are occupied (due to bayesian plausible values imputation).
################################################################################
### Example 11: differential item functioning in partial credit models ###
################################################################################
# load partial credit long format data
data(reading)
# create a small wide format exemplary data set, only booklet 08
dw <- reshape2::dcast(subset(reading, bookletID == "TH08"), idstud+sex~item, value.var = "valueSum") |>
dplyr::mutate(sex = car::recode(sex, "'male'=0; 'female'=1", as.factor=FALSE)) |>
eatTools::na_omit_selection("sex")
# define the model: the specification resembles exemple 9 in tam.mml help page
defT<- defineModel(dat = dw, items = -c(1:2), DIF.var="sex", id = "idstud", irtmodel = "PCM",software="tam")
#> 4 variable(s) are not strictly dichotomous with 0/1 ... Expect a rating scale model or partial credit model.
#> Items(s) 'D025033', 'D204013', 'D225113': 0, 1, 2, 3, 4
#> Items(s) 'D205143': 0, 1, 2, 3, 4, 5
#> Dataset is completely linked.
#> 'gauss' has been chosen for estimation method. Number of nodes was not explicitly specified. Set nodes to 20.
#> Q matrix specifies 1 dimension(s).
runT<- runModel(defT)
resT<- getResults(runT)
#> Warning: NaNs produced
#> Warning: NaNs produced
#> Warning: NaNs produced
#> Warning: NaNs produced
#> Warning: NaNs produced
#> Warning: NaNs produced
#> Warning: NaNs produced
#> Warning: NaNs produced
#> Warning: NaNs produced
#> Warning: NaNs produced
#> Warning: NaNs produced
#> Warning: NaNs produced
#> Warning: NaNs produced
#> Warning: NaNs produced
#> Warning: NaNs produced
#> Warning: NaNs produced
#> Warning: NaNs produced
#> Warning: NaNs produced
#> Warning: NaNs produced
#> Warning: NaNs produced
#> Warning: NaNs produced
#> Warning: NaNs produced
#> Warning: NaNs produced
#> Warning: NaNs produced
#> Warning: NaNs produced
#> Warning: NaNs produced
#> Warning: NaNs produced
#> Warning: NaNs produced
#> Warning: NaNs produced
#> Warning: NaNs produced
#> Warning: NaNs produced
#> Warning: NaNs produced
#> Warning: NaNs produced
#> Warning: NaNs produced
#> Warning: NaNs produced
#> Warning: NaNs produced
#> Warning: NaNs produced
#> Warning: NaNs produced
#> Warning: NaNs produced
#> Warning: NaNs produced
#> Warning: NaNs produced
#> Warning: NaNs produced
#> Warning: NaNs produced
#> Warning: NaNs produced
#> Warning: NaNs produced
#> Warning: NaNs produced
#> Warning: NaNs produced
#> Warning: NaNs produced
#> Warning: NaNs produced
#> Warning: NaNs produced
#> Warning: NaNs produced
#> Warning: NaNs produced
#> Warning: NaNs produced
#> Warning: NaNs produced
#> Warning: NaNs produced
#> Warning: NaNs produced
#> Warning: NaNs produced
#> Warning: NaNs produced
#> Warning: NaNs produced
#> Warning: NaNs produced
#> Warning: NaNs produced
#> Warning: NaNs produced
#> Warning: NaNs produced
#> Warning: NaNs produced
#> Warning: NaNs produced
#> Warning: NaNs produced
#> Warning: NaNs produced
#> Warning: NaNs produced
#> Warning: NaNs produced
#> Warning: NaNs produced
#> Warning: NaNs produced
#> Warning: NaNs produced
#> Warning: NaNs produced
#> Warning: NaNs produced
#> Warning: NaNs produced
#> Warning: NaNs produced
#> Warning: NaNs produced
#> Warning: NaNs produced
#> Warning: NaNs produced
#> Warning: NaNs produced
#> Warning: NaNs produced
#> Warning: NaNs produced
#> Warning: NaNs produced
#> Warning: NaNs produced
#> Warning: NaNs produced
#> Warning: NaNs produced
#> Warning: NaNs produced
#> Warning: NaNs produced
#> Warning: NaNs produced
#> Warning: NaNs produced
#> Warning: NaNs produced
#> Warning: NaNs produced
#> Warning: NaNs produced
#> Warning: NaNs produced
#> Warning: NaNs produced
#> Warning: NaNs produced
#> Warning: NaNs produced
#> Warning: NaNs produced
#> Warning: NaNs produced
#> Warning: NaNs produced
#> Warning: NaNs produced
#> Warning: NaNs produced
#> Warning: NaNs produced
#> Warning: NaNs produced
#> Warning: NaNs produced
#> Warning: NaNs produced
#> Warning: NaNs produced
#> Warning: NaNs produced
#> Warning: NaNs produced
#> Warning: NaNs produced
#> Warning: NaNs produced
#> Warning: NaNs produced
#> Warning: NaNs produced
#> Warning: NaNs produced
#> Warning: NaNs produced
#> Warning: NaNs produced
#> Warning: NaNs produced
#> Warning: NaNs produced
#> Warning: NaNs produced
#> Warning: NaNs produced
#> Warning: NaNs produced
#> Warning: NaNs produced
#> Warning: NaNs produced
#> Warning: NaNs produced
#> Warning: NaNs produced
#> Warning: NaNs produced
#> Warning: NaNs produced
#> Warning: NaNs produced
#> Warning: NaNs produced
#> Warning: NaNs produced
#> Warning: NaNs produced
#> Warning: NaNs produced
#> Warning: NaNs produced
#> Warning: NaNs produced
#> Warning: NaNs produced
#> Warning: NaNs produced
#> Warning: NaNs produced
#> Warning: NaNs produced
#> Warning: NaNs produced
#> Warning: NaNs produced
#> Warning: NaNs produced
#> Warning: NaNs produced
#> Warning: NaNs produced
#> Warning: NaNs produced
#> Warning: NaNs produced
#> Warning: NaNs produced
#> Warning: NaNs produced
#> Warning: NaNs produced
#> Warning: NaNs produced
#> Warning: NaNs produced
#> Warning: NaNs produced
#> Warning: NaNs produced
#> Warning: NaNs produced
#> Warning: NaNs produced
#> Warning: NaNs produced
#> Warning: NaNs produced
#> Warning: NaNs produced
#> Warning: NaNs produced
#> Warning: NaNs produced
#> Warning: NaNs produced
#> Warning: NaNs produced
#> Warning: NaNs produced
#> Warning: NaNs produced
#> Warning: NaNs produced
#> Warning: NaNs produced
#> Warning: NaNs produced
#> Warning: NaNs produced
#> Warning: NaNs produced
#> Warning: NaNs produced
#> Warning: NaNs produced
#> Warning: NaNs produced
#> Warning: NaNs produced
#> Warning: NaNs produced
#> Warning: NaNs produced
#> Warning: NaNs produced
#> Warning: NaNs produced
#> Warning: NaNs produced
#> Warning: NaNs produced
#> Warning: NaNs produced
#> Warning: NaNs produced
#> Warning: NaNs produced
#> Warning: NaNs produced
#> Warning: NaNs produced
#> Warning: NaNs produced
#> Warning: NaNs produced
#> Warning: NaNs produced
#> Warning: NaNs produced
#> Warning: NaNs produced
#> Warning: NaNs produced
#> Warning: NaNs produced
#> Warning: NaNs produced
#> Warning: NaNs produced
#> Warning: NaNs produced
#> Warning: NaNs produced
#> Warning: NaNs produced
#> Warning: NaNs produced
#> Warning: NaNs produced
#> Warning: NaNs produced
#> Warning: NaNs produced
#> Warning: NaNs produced
#> Warning: NaNs produced
#> Warning: NaNs produced
#> Warning: NaNs produced
#> Warning: NaNs produced
#> Warning: NaNs produced
#> Warning: NaNs produced
#> Warning: NaNs produced
#> Warning: NaNs produced
#> Warning: NaNs produced
#> Warning: NaNs produced
#> Warning: NaNs produced
#> Warning: NaNs produced
#> Warning: NaNs produced
#> Warning: NaNs produced
#> Warning: NaNs produced
#> Warning: NaNs produced
#> Warning: NaNs produced
#> Warning: NaNs produced
#> Warning: NaNs produced
#> Warning: NaNs produced
#> Warning: NaNs produced
#> Warning: NaNs produced
#> Warning: NaNs produced
#> Warning: NaNs produced
#> Warning: NaNs produced
#> Warning: NaNs produced
#> Warning: NaNs produced
#> Warning: NaNs produced
#> Warning: NaNs produced
#> Warning: NaNs produced
#> Warning: NaNs produced
#> Warning: NaNs produced
#> Warning: NaNs produced
#> Warning: NaNs produced
#> Warning: NaNs produced
#> Warning: NaNs produced
#> Warning: NaNs produced
#> Warning: NaNs produced
#> Warning: NaNs produced
#> Warning: NaNs produced
#> Warning: NaNs produced
#> Warning: NaNs produced
#> Warning: NaNs produced
#> Warning: NaNs produced
#> Warning: NaNs produced
#> Warning: NaNs produced
#> Warning: NaNs produced
#> Warning: NaNs produced
#> Warning: NaNs produced
#> Warning: NaNs produced
#> Warning: NaNs produced
#> Warning: NaNs produced
#> Warning: NaNs produced
#> Warning: NaNs produced
#> Warning: NaNs produced
#> Warning: NaNs produced
#> Warning: NaNs produced
#> Warning: NaNs produced
#> Warning: NaNs produced
#> Warning: NaNs produced
#> Warning: NaNs produced
#> Warning: NaNs produced
#> Warning: NaNs produced
#> Warning: NaNs produced
#> Warning: NaNs produced
#> Warning: NaNs produced
#> Warning: NaNs produced
#> Warning: NaNs produced
#> Warning: NaNs produced
#> Warning: NaNs produced
#> Warning: NaNs produced
#> Warning: NaNs produced
#> Warning: NaNs produced
#> Warning: NaNs produced
#> Warning: NaNs produced
#> Warning: NaNs produced
#> Warning: NaNs produced
#> Warning: NaNs produced
#> Warning: NaNs produced
#> Warning: NaNs produced
#> Warning: NaNs produced
#> Warning: NaNs produced
#> Warning: NaNs produced
#> Warning: NaNs produced
#> Warning: NaNs produced
#> Warning: NaNs produced
#> Warning: NaNs produced
#> Warning: NaNs produced
#> Warning: NaNs produced
#> Warning: NaNs produced
#> Warning: NaNs produced
#> Warning: NaNs produced
#> Warning: NaNs produced
#> Warning: NaNs produced
#> Warning: NaNs produced
#> Warning: NaNs produced
#> Warning: NaNs produced
#> Warning: NaNs produced
#> Warning: NaNs produced
#> Warning: NaNs produced
#> Warning: NaNs produced
#> Warning: NaNs produced
#> Getting standard errors with the tam.se function: 2.7 secs
#> Error in car::recode(dat[, "var1"], recSt):
#> in recode term: 'D225123:step3' = 'D:step_2251233'
#> message: Error in parse(text = range[[1]][1]) :
#> <text>:1:2: unexpected INCOMPLETE_STRING
#> 1: 'D225123
#> ^
# the same model in conquest (recommended)
defC<- defineModel(dat = dw, items = -c(1:2), model.statement = "item+item*step - sex + item*sex", DIF.var="sex", id = "idstud", analysis.name = "dif_pcm", dir=tempdir())
#> Cannot find conquest 2007 executable file. Please choose manually.
#> Error in file.choose(): file choice cancelled
runC<- runModel(defC, wait=FALSE)
#> Error: object 'defC' not found
resC<- getResults(runC)
#> Error: object 'runC' not found
it <- itemFromRes(resC)
#> Error: object 'resC' not found
shw <- get.shw(file.path(tempdir(), "dif_pcm.shw"), dif.term = "item*sex")
#> Warning: cannot open file '/tmp/RtmpByQecb/dif_pcm.shw': No such file or directory
#> Error in file(file, "r"): cannot open the connection
################################################################################
### Example 12: Norm study within the realm of partial credit models ###
################################################################################
# The following example mimics the steps the required to norm the educational standards
# when the underlying model is a partial credit model.
# load partial credit long format data. This large scale assessment data surveyed in
# an incomplete block design is representative of a domain covered by the new standards.
# We assume thta the sample was drawn from the norm population (reference population)
data(reading)
# transform the data to the wide format
datW <- reshape2::dcast(reading[which(reading[,"type"] != "iglu"),],idstud+sex+language+country~item, value.var = "valueSum")
# combine the two lowest categories to avoid categories with too few observations
datW[,"D205143"] <- car::recode(datW[,"D205143"], "1=0; 2=1; 3=2; 4=3; 5=4")
# partial credit model. We draw 15 plausible values to achieve greater precision
# in estimating the distribution of individuals.
def1 <- defineModel(dat=datW, items = -c(1:4), id=1, irtmodel = "PCM", software="tam", nodes = 21, n.plausible=15)
#> 7 variable(s) are not strictly dichotomous with 0/1 ... Expect a rating scale model or partial credit model.
#> Items(s) 'D025033', 'D201113', 'D204013', 'D205143', 'D224053', 'D225113': 0, 1, 2, 3, 4
#> Items(s) 'D223143': 0, 1, 2, 3, 4, 5
#> 13 subject(s) do not solve any item:
#> P0007 (12 false), P0837 (13 false), P1569 (49 false) ...
#> 20 subject(s) solved each item: P0968 (11 correct), P0843 (13 correct), P1047 (29 correct) ...
#> Dataset is completely linked.
#> Q matrix specifies 1 dimension(s).
run1 <- runModel(def1)
# use ntheta = 40000 for higher precision
res1 <- getResults(run1, ntheta = 40000, theta.model = FALSE)
#> Getting standard errors with the tam.se function: 1.1 secs
#> Getting infit parameters calling tam.fit from getTamInfit: 0.3 secs
#> Getting WLEs calling tam.wle from getTamWles: 0.6 secs
#> |***************|
#> |---------------|
#> Getting PVs calling tam.pv from getTamPVs: 19.1 secs
#> Getting Q3 statistic calling tam.modelfit from getTamQ3: 1.8 secs
# estimate mean and standard deviation of the reference population from the plausible values
# use the long format by toWideFormat = FALSE. Load the eatRep package for mean/SD computation
library(eatRep)
pv1 <- pvFromRes(res1, toWideFormat = FALSE)
mSD <- repMean(datL = pv1, ID="idstud", imp="imp", dependent="value")
#> 1 analyse(s) overall according to: 'group.splits = 0'.
#> Assume unnested structure with 15 imputations.
#>
mSDR <- report2(mSD, round = FALSE)[["plain"]]; dim(mSDR)
#> [1] 3 10
# use mean/SD to transform the item and person parameters. Since the cur scores have not yet
# been established - as they are still being defined in the standard-setting process - we are
# assigning arbitrary values for now. Since the sample can be considered as being drawn from
# the reference population, no equating procedure is necessary. The function is therefore
# simply passed through. The mean and standard deviation of the educational standards metric
# are set to 500 and 100, respectively.
eq <- equat1pl(res1)
#> Found 1 model(s).
#> Equating is executed for each dimension in each model separately.
#> No norm parameter defined ('prmNorm' is missing). Treat current sample as drawn from the reference population.
trnsf<- transformToBista(equatingList=eq, refPop= data.frame(domain="Dim1",
m = mSDR[which(mSDR[,"parameter"] == "mean"),"est"],
sd = mSDR[which(mSDR[,"parameter"] == "sd"),"est"]),
cuts = list(Dim1 = list(values = c(400,600))), vera=FALSE)
#> The 'refPop' data.frame does not include information about reference population mean/SD on Bista metric. Values will be defaulted to mean = 500 and SD = 100.
#> Warning: Skip check whether all competence levels are occupied (due to bayesian plausible values imputation).